Datasheet Blank Template
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
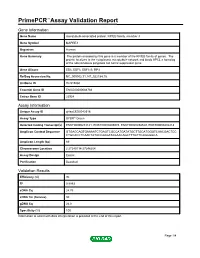
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name microtubule-associated protein, RP/EB family, member 3 Gene Symbol MAPRE3 Organism Human Gene Summary The protein encoded by this gene is a member of the RP/EB family of genes. The protein localizes to the cytoplasmic microtubule network and binds APCL a homolog of the adenomatous polyposis coli tumor suppressor gene. Gene Aliases EB3, EBF3, EBF3-S, RP3 RefSeq Accession No. NC_000002.11, NT_022184.15 UniGene ID Hs.515860 Ensembl Gene ID ENSG00000084764 Entrez Gene ID 22924 Assay Information Unique Assay ID qHsaCED0042518 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000233121, ENST00000405074, ENST00000458529, ENST00000402218 Amplicon Context Sequence GTGACCAGTGAAAATCTGAGTCGCCATGATATGCTTGCATGGGTCAACGACTCC CTGCACCTCAACTATACCAAGATAGAACAGCTTTGTTCAGGGGCA Amplicon Length (bp) 69 Chromosome Location 2:27245114-27246204 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 90 R2 0.9993 cDNA Cq 24.76 cDNA Tm (Celsius) 80 gDNA Cq 26.8 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 1/4 PrimePCR™Assay Validation Report MAPRE3, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 2/4 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Human Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. -

EB3 (MAPRE3) (NM 012326) Human Tagged ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC204135 EB3 (MAPRE3) (NM_012326) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: EB3 (MAPRE3) (NM_012326) Human Tagged ORF Clone Tag: Myc-DDK Symbol: MAPRE3 Synonyms: EB3; EBF3; EBF3-S; RP3 Vector: pCMV6-Entry (PS100001) E. coli Selection: Kanamycin (25 ug/mL) Cell Selection: Neomycin ORF Nucleotide >RC204135 ORF sequence Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGGCCGTCAATGTGTACTCCACATCTGTGACCAGTGAAAATCTGAGTCGCCATGATATGCTTGCATGGG TCAACGACTCCCTGCACCTCAACTATACCAAGATAGAACAGCTTTGTTCAGGGGCAGCCTACTGCCAGTT CATGGACATGCTCTTCCCCGGCTGTGTGCACTTGAGGAAAGTGAAGTTCCAGGCCAAACTAGAGCATGAA TACATCCACAACTTCAAGGTGCTGCAAGCAGCTTTCAAGAAGATGGGTGTTGACAAAATCATTCCTGTAG AGAAATTAGTGAAAGGAAAATTCCAAGATAATTTTGAGTTTATTCAGTGGTTTAAGAAATTCTTTGACGC AAACTATGATGGAAAGGATTACAACCCTCTGCTGGCGCGGCAGGGCCAGGACGTAGCGCCACCTCCTAAC CCAGGTGATCAGATCTTCAACAAATCCAAGAAACTCATTGGCACAGCAGTTCCACAGAGGACGTCCCCCA CAGGCCCAAAAAACATGCAGACCTCTGGCCGGCTGAGCAATGTGGCCCCCCCCTGCATTCTCCGGAAGAA TCCTCCATCAGCCCGAAATGGCGGCCATGAGACTGATGCCCAAATTCTTGAACTCAACCAACAGCTGGTG GACTTGAAGCTGACAGTGGATGGGCTGGAGAAGGAACGTGACTTCTACTTCAGCAAACTTCGTGACATCG AGCTCATCTGCCAGGAGCATGAAAGTGAAAACAGCCCTGTTATCTCAGGCATCATTGGCATCCTCTATGC CACAGAGGAAGGATTCGCACCCCCTGAGGACGATGAGATTGAAGAGCATCAACAAGAAGACCAGGACGAG TAC ACGCGTACGCGGCCGCTCGAGCAGAAACTCATCTCAGAAGAGGATCTGGCAGCAAATGATATCCTGGATT -

Supporting Information
Supporting Information Pouryahya et al. SI Text Table S1 presents genes with the highest absolute value of Ricci curvature. We expect these genes to have significant contribution to the network’s robustness. Notably, the top two genes are TP53 (tumor protein 53) and YWHAG gene. TP53, also known as p53, it is a well known tumor suppressor gene known as the "guardian of the genome“ given the essential role it plays in genetic stability and prevention of cancer formation (1, 2). Mutations in this gene play a role in all stages of malignant transformation including tumor initiation, promotion, aggressiveness, and metastasis (3). Mutations of this gene are present in more than 50% of human cancers, making it the most common genetic event in human cancer (4, 5). Namely, p53 mutations play roles in leukemia, breast cancer, CNS cancers, and lung cancers, among many others (6–9). The YWHAG gene encodes the 14-3-3 protein gamma, a member of the 14-3-3 family proteins which are involved in many biological processes including signal transduction regulation, cell cycle pro- gression, apoptosis, cell adhesion and migration (10, 11). Notably, increased expression of 14-3-3 family proteins, including protein gamma, have been observed in a number of human cancers including lung and colorectal cancers, among others, suggesting a potential role as tumor oncogenes (12, 13). Furthermore, there is evidence that loss Fig. S1. The histogram of scalar Ricci curvature of 8240 genes. Most of the genes have negative scalar Ricci curvature (75%). TP53 and YWHAG have notably low of p53 function may result in upregulation of 14-3-3γ in lung cancer Ricci curvatures. -

In This Table Protein Name, Uniprot Code, Gene Name P-Value
Supplementary Table S1: In this table protein name, uniprot code, gene name p-value and Fold change (FC) for each comparison are shown, for 299 of the 301 significantly regulated proteins found in both comparisons (p-value<0.01, fold change (FC) >+/-0.37) ALS versus control and FTLD-U versus control. Two uncharacterized proteins have been excluded from this list Protein name Uniprot Gene name p value FC FTLD-U p value FC ALS FTLD-U ALS Cytochrome b-c1 complex P14927 UQCRB 1.534E-03 -1.591E+00 6.005E-04 -1.639E+00 subunit 7 NADH dehydrogenase O95182 NDUFA7 4.127E-04 -9.471E-01 3.467E-05 -1.643E+00 [ubiquinone] 1 alpha subcomplex subunit 7 NADH dehydrogenase O43678 NDUFA2 3.230E-04 -9.145E-01 2.113E-04 -1.450E+00 [ubiquinone] 1 alpha subcomplex subunit 2 NADH dehydrogenase O43920 NDUFS5 1.769E-04 -8.829E-01 3.235E-05 -1.007E+00 [ubiquinone] iron-sulfur protein 5 ARF GTPase-activating A0A0C4DGN6 GIT1 1.306E-03 -8.810E-01 1.115E-03 -7.228E-01 protein GIT1 Methylglutaconyl-CoA Q13825 AUH 6.097E-04 -7.666E-01 5.619E-06 -1.178E+00 hydratase, mitochondrial ADP/ATP translocase 1 P12235 SLC25A4 6.068E-03 -6.095E-01 3.595E-04 -1.011E+00 MIC J3QTA6 CHCHD6 1.090E-04 -5.913E-01 2.124E-03 -5.948E-01 MIC J3QTA6 CHCHD6 1.090E-04 -5.913E-01 2.124E-03 -5.948E-01 Protein kinase C and casein Q9BY11 PACSIN1 3.837E-03 -5.863E-01 3.680E-06 -1.824E+00 kinase substrate in neurons protein 1 Tubulin polymerization- O94811 TPPP 6.466E-03 -5.755E-01 6.943E-06 -1.169E+00 promoting protein MIC C9JRZ6 CHCHD3 2.912E-02 -6.187E-01 2.195E-03 -9.781E-01 Mitochondrial 2- -

PRODUCT SPECIFICATION Product Datasheet
Product Datasheet QPrEST PRODUCT SPECIFICATION Product Name QPrEST MARE3 Mass Spectrometry Protein Standard Product Number QPrEST38970 Protein Name Microtubule-associated protein RP/EB family member 3 Uniprot ID Q9UPY8 Gene MAPRE3 Product Description Stable isotope-labeled standard for absolute protein quantification of Microtubule- associated protein RP/EB family member 3. Lys (13C and 15N) and Arg (13C and 15N) metabolically labeled recombinant human protein fragment. Application Absolute protein quantification using mass spectrometry Sequence (excluding YDGKDYNPLLARQGQDVAPPPNPGDQIFNKSKKLIGTAVPQRTSPTGPKN fusion tag) MQTSGRLSNVAPPCILRKNPPSARNGGHETDAQILELNQQLVDLKLTVDG LEKERDFYFSKLR Theoretical MW 30368 Da including N-terminal His6ABP fusion tag Fusion Tag A purification and quantification tag (QTag) consisting of a hexahistidine sequence followed by an Albumin Binding Protein (ABP) domain derived from Streptococcal Protein G. Expression Host Escherichia coli LysA ArgA BL21(DE3) Purification IMAC purification Purity >90% as determined by Bioanalyzer Protein 230 Purity Assay Isotopic Incorporation >99% Concentration >5 μM after reconstitution in 100 μl H20 Concentration Concentration determined by LC-MS/MS using a highly pure amino acid analyzed internal Determination reference (QTag), CV ≤10%. Amount >0.5 nmol per vial, two vials supplied. Formulation Lyophilized in 100 mM Tris-HCl 5% Trehalose, pH 8.0 Instructions for Spin vial before opening. Add 100 μL ultrapure H2O to the vial. Vortex thoroughly and spin Reconstitution down. For further dilution, see Application Protocol. Shipping Shipped at ambient temperature Storage Lyophilized product shall be stored at -20°C. See COA for expiry date. Reconstituted product can be stored at -20°C for up to 4 weeks. Avoid repeated freeze-thaw cycles. Notes For research use only Product of Sweden. -

Predicted Coronavirus Nsp5 Protease Cleavage Sites in The
bioRxiv preprint doi: https://doi.org/10.1101/2021.06.08.447224; this version posted June 8, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Predicted Coronavirus Nsp5 Protease Cleavage Sites in the 2 Human Proteome: A Resource for SARS-CoV-2 Research 3 Benjamin M. Scott1,2*, Vincent Lacasse3, Ditte G. Blom4, Peter D. Tonner5, Nikolaj S. Blom6 4 1 Associate, Biosystems and Biomaterials Division, National Institute of Standards and Technology, 5 Gaithersburg, Maryland, USA. 6 2 Department of Chemistry and Biochemistry, University of Maryland, College Park, Maryland, USA. 7 3 Segal Cancer Proteomics Centre, Lady Davis Institute, Jewish General Hospital, McGill University, Montreal, 8 Quebec, Canada 9 4 Department of Applied Mathematics and Computer Science, Technical University of Denmark, Lyngby, 10 Denmark. 11 5 Statistical Engineering Division, National Institute of Standards and Technology, Gaithersburg, Maryland, USA 12 6 Department of Bioengineering, Kongens Lyngby, Technical University of Denmark 13 14 *Corresponding author, [email protected] 15 16 BMS current affiliation: Concordia University, Centre for Applied Synthetic Biology, Montreal, Quebec, Canada 17 18 19 Abstract 20 Background: The coronavirus nonstructural protein 5 (Nsp5) is a cysteine protease required for 21 processing the viral polyprotein and is therefore crucial for viral replication. Nsp5 from several 22 coronaviruses have also been found to cleave host proteins, disrupting molecular pathways 23 involved in innate immunity. Nsp5 from the recently emerged SARS-CoV-2 virus interacts with 24 and can cleave human proteins, which may be relevant to the pathogenesis of COVID-19. -

Identification of Endogenous Adenomatous Polyposis Coli Interaction Partners and Β-Catenin−Independent Targets by Proteomics
Published OnlineFirst June 3, 2019; DOI: 10.1158/1541-7786.MCR-18-1154 Cancer Genes and Networks Molecular Cancer Research Identification of Endogenous Adenomatous Polyposis Coli Interaction Partners and b-Catenin– Independent Targets by Proteomics Olesja Popow1,2,3,Joao~ A. Paulo2, Michael H. Tatham4, Melanie S. Volk3, Alejandro Rojas-Fernandez5, Nicolas Loyer3, Ian P. Newton3, Jens Januschke3, Kevin M. Haigis1,6, and Inke Nathke€ 3 Abstract Adenomatous Polyposis Coli (APC) is the most frequently tion between endogenous MINK1 and APC and further mutated gene in colorectal cancer. APC negatively regulates confirmed the negative, and b-catenin–independent, regu- the Wnt signaling pathway by promoting the degradation of lation of MINK1 by APC. Increased Mink1/Msn levels were b-catenin, but the extent to which APC exerts Wnt/b-cate- also observed in mouse intestinal tissue and Drosophila nin–independent tumor-suppressive activity is unclear. To follicular cells expressing mutant Apc/APC when compared identify interaction partners and b-catenin–independent with wild-type tissue/cells. Collectively, our results highlight targets of endogenous, full-length APC, we applied label- the extent and importance of Wnt-independent APC func- free and multiplexed tandem mass tag-based mass spec- tions in epithelial biology and disease. trometry. Affinity enrichment-mass spectrometry identified more than 150 previously unidentified APC interaction Implications: The tumor-suppressive function of APC, the partners. Moreover, our global proteomic analysis revealed most frequently mutated gene in colorectal cancer, is mainly that roughly half of the protein expression changes that attributed to its role in b-catenin/Wnt signaling. Our study occur in response to APC loss are independent of b-catenin. -

Chromatin Conformation Links Distal Target Genes to CKD Loci
BASIC RESEARCH www.jasn.org Chromatin Conformation Links Distal Target Genes to CKD Loci Maarten M. Brandt,1 Claartje A. Meddens,2,3 Laura Louzao-Martinez,4 Noortje A.M. van den Dungen,5,6 Nico R. Lansu,2,3,6 Edward E.S. Nieuwenhuis,2 Dirk J. Duncker,1 Marianne C. Verhaar,4 Jaap A. Joles,4 Michal Mokry,2,3,6 and Caroline Cheng1,4 1Experimental Cardiology, Department of Cardiology, Thoraxcenter Erasmus University Medical Center, Rotterdam, The Netherlands; and 2Department of Pediatrics, Wilhelmina Children’s Hospital, 3Regenerative Medicine Center Utrecht, Department of Pediatrics, 4Department of Nephrology and Hypertension, Division of Internal Medicine and Dermatology, 5Department of Cardiology, Division Heart and Lungs, and 6Epigenomics Facility, Department of Cardiology, University Medical Center Utrecht, Utrecht, The Netherlands ABSTRACT Genome-wide association studies (GWASs) have identified many genetic risk factors for CKD. However, linking common variants to genes that are causal for CKD etiology remains challenging. By adapting self-transcribing active regulatory region sequencing, we evaluated the effect of genetic variation on DNA regulatory elements (DREs). Variants in linkage with the CKD-associated single-nucleotide polymorphism rs11959928 were shown to affect DRE function, illustrating that genes regulated by DREs colocalizing with CKD-associated variation can be dysregulated and therefore, considered as CKD candidate genes. To identify target genes of these DREs, we used circular chro- mosome conformation capture (4C) sequencing on glomerular endothelial cells and renal tubular epithelial cells. Our 4C analyses revealed interactions of CKD-associated susceptibility regions with the transcriptional start sites of 304 target genes. Overlap with multiple databases confirmed that many of these target genes are involved in kidney homeostasis. -

The Microtubule Skeleton and the Evolution of Neuronal Complexity in Vertebrates
Biol. Chem. 2019; 400(9): 1163–1179 Nataliya I. Trushina, Armen Y. Mulkidjanian and Roland Brandt* The microtubule skeleton and the evolution of neuronal complexity in vertebrates https://doi.org/10.1515/hsz-2019-0149 Received February 4, 2019; accepted April 17, 2019; previously Introduction published online May 22, 2019 The complexity of the nervous system permits the devel- Abstract: The evolution of a highly developed nervous opment of sophisticated behavioral repertoires, such as system is mirrored by the ability of individual neurons to language, tool use, self-awareness, symbolic thought, develop increased morphological complexity. As microtu- cultural learning and consciousness. The basis for the bules (MTs) are crucially involved in neuronal development, development of such a complexity is a high neuronal het- we tested the hypothesis that the evolution of complexity is erogeneity caused by neuronal diversification on the one driven by an increasing capacity of the MT system for regu- hand and a large interconnectivity between the individual lated molecular interactions as it may be implemented by neurons on the other hand (Muotri and Gage, 2006). In a higher number of molecular players and a greater ability addition, the brain constantly needs to adapt to func- of the individual molecules to interact. We performed bio- tional challenges of various kinds during development informatics analysis on different classes of components of and adulthood by a process called neural plasticity (Zilles, the vertebrate neuronal MT cytoskeleton. We show that the 1992). At a single cell level, neural plasticity involves number of orthologs of tubulin structure proteins, MT-bind- changes in the number or the strength of synaptic con- ing proteins and tubulin-sequestering proteins expanded tacts between individual neurons thereby changing the during vertebrate evolution. -

And Hippo Signaling-Dependent Plasticity During Lineage
RESEARCH ARTICLE Position- and Hippo signaling-dependent plasticity during lineage segregation in the early mouse embryo Eszter Posfai1, Sophie Petropoulos2,3,4, Flavia Regina Oliveira de Barros1, John Paul Schell2,4, Igor Jurisica5,6,7, Rickard Sandberg3,8, Fredrik Lanner2,4, Janet Rossant1,9* 1Program in Developmental and Stem Cell Biology, Hospital for Sick Children, Toronto, Canada; 2Department of Clinical Science, Intervention and Technology, Karolinska Institutet, Stockholm, Sweden; 3Ludwig Institute for Cancer Research, Karolinska Institutet, Stockholm, Sweden; 4Division of Obstetrics and Gynecology, Karolinska Universitetssjukhuset, Stockholm, Sweden; 5Princess Margaret Cancer Centre, University Health Network, University of Toronto, Toronto, Canada; 6Departments of Medical Biophysics and Computer Science, University of Toronto, Toronto, Canada; 7Institute of Neuroimmunology, Slovak Academy of Sciences, Bratislava, Slovakia; 8Department of Cell and Molecular Biology, Karolinska Institutet, Stockholm, Sweden; 9Department of Molecular Genetics, University of Toronto, Toronto, Canada Abstract The segregation of the trophectoderm (TE) from the inner cell mass (ICM) in the mouse blastocyst is determined by position-dependent Hippo signaling. However, the window of responsiveness to Hippo signaling, the exact timing of lineage commitment and the overall relationship between cell commitment and global gene expression changes are still unclear. Single- cell RNA sequencing during lineage segregation revealed that the TE transcriptional profile *For correspondence: janet. stabilizes earlier than the ICM and prior to blastocyst formation. Using quantitative Cdx2-eGFP [email protected] expression as a readout of Hippo signaling activity, we assessed the experimental potential of individual blastomeres based on their level of Cdx2-eGFP expression and correlated potential with Competing interests: The gene expression dynamics. We find that TE specification and commitment coincide and occur at authors declare that no competing interests exist. -

Elabscience.Com ® E-Mail:[email protected] Elabscience Elabscience Biotechnology Inc
Produktinformation Diagnostik & molekulare Diagnostik Laborgeräte & Service Zellkultur & Verbrauchsmaterial Forschungsprodukte & Biochemikalien Weitere Information auf den folgenden Seiten! See the following pages for more information! Lieferung & Zahlungsart Lieferung: frei Haus Bestellung auf Rechnung SZABO-SCANDIC Lieferung: € 10,- HandelsgmbH & Co KG Erstbestellung Vorauskassa Quellenstraße 110, A-1100 Wien T. +43(0)1 489 3961-0 Zuschläge F. +43(0)1 489 3961-7 [email protected] • Mindermengenzuschlag www.szabo-scandic.com • Trockeneiszuschlag • Gefahrgutzuschlag linkedin.com/company/szaboscandic • Expressversand facebook.com/szaboscandic Tel:240-252-7368(USA) Fax:240-252-7376(USA) www.elabscience.com ® E-mail:[email protected] Elabscience Elabscience Biotechnology Inc. MAPRE3 Polyclonal Antibody Catalog No. E-AB-11172 Reactivity H,M,R Storage Store at -20℃. Avoid freeze / thaw cycles. Host Rabbit Applications WB,ELISA ElabscienceIsotype IgG Note: Centrifuge before opening to ensure complete recovery of vial contents. Images Immunogen Information Elabscience Immunogen Recombinant protein of human MAPRE3 Gene Accession BC011557 Swissprot Q9UPY8 Synonyms EB3,RP3,EBF3,EBF3-S Western Blot analysis of Human fetal Product Information brain and Mouse brain tissue using Calculated MW 32kDa MAPRE3 Polyclonal Antibody at Buffer PBS with 0.05% sodium azide and 50% glycerol, dilution of 1:500 PH7.4 Purify Affinity purification Dilution WB 1:500-1:2000 Elabscience Background The protein encoded by this gene is a member of the RP/EB family of genes. The protein localizes to the cytoplasmic microtubule network and binds APCL, a homolog of the adenomatous polyposis coli tumor suppressor gene. Elabscience Elabscience For Research Use Only Focus on your research Thank you for your recent purchase. -

The Genetic Architecture of Structural Left-Right Asymmetry of the Human Brain
bioRxiv preprint doi: https://doi.org/10.1101/2020.06.30.179721; this version posted July 7, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. The genetic architecture of structural left-right asymmetry of the human brain Zhiqiang Sha1, Dick Schijven1, Amaia Carrion-Castillo1, Marc Joliot2, Bernard Mazoyer2, Simon E. Fisher1,3, Fabrice Crivello2, Clyde Francks1,3* 1 Language and Genetics Department, Max Planck Institute for Psycholinguistics, Nijmegen, The Netherlands 2 Groupe d’Imagerie Neurofonctionnelle, Institut des Maladies Neurodégénératives, Centre National de la Recherche Scientifique, Commissariat à l’Energie Atomique, et Université de Bordeaux, Bordeaux, France 3 Donders Institute for Brain, Cognition and Behaviour, Radboud University, Nijmegen, The Netherlands Corresponding author: Clyde Francks, DPhil. Language and Genetics Department, Max Planck Institute for Psycholinguistics, Nijmegen, The Netherlands; E-mail: [email protected] Keywords Magnetic resonance imaging, brain asymmetry, brain laterality, brain structure, multivariate analysis, heritability, genome-wide association scan, population neuroscience, microtubule, imaging genetics, UK Biobank. Manuscript information: 28 text pages, 5 figures, 1 table, 1684 words. (Supplementary Information: 15 text pages, 10 figures, 19 tables.) 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.06.30.179721; this version posted July 7, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Left-right hemispheric asymmetry is an important aspect of healthy brain organization for many functions including language, and can be altered in cognitive and psychiatric disorders1-8.