P53 Deficiency Triggers Dysregulation of Diverse Cellular Processes in Physiological Oxygen
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Deregulated Gene Expression Pathways in Myelodysplastic Syndrome Hematopoietic Stem Cells
Leukemia (2010) 24, 756–764 & 2010 Macmillan Publishers Limited All rights reserved 0887-6924/10 $32.00 www.nature.com/leu ORIGINAL ARTICLE Deregulated gene expression pathways in myelodysplastic syndrome hematopoietic stem cells A Pellagatti1, M Cazzola2, A Giagounidis3, J Perry1, L Malcovati2, MG Della Porta2,MJa¨dersten4, S Killick5, A Verma6, CJ Norbury7, E Hellstro¨m-Lindberg4, JS Wainscoat1 and J Boultwood1 1LRF Molecular Haematology Unit, NDCLS, John Radcliffe Hospital, Oxford, UK; 2Department of Hematology Oncology, University of Pavia Medical School, Fondazione IRCCS Policlinico San Matteo, Pavia, Italy; 3Medizinische Klinik II, St Johannes Hospital, Duisburg, Germany; 4Division of Hematology, Department of Medicine, Karolinska Institutet, Stockholm, Sweden; 5Department of Haematology, Royal Bournemouth Hospital, Bournemouth, UK; 6Albert Einstein College of Medicine, Bronx, NY, USA and 7Sir William Dunn School of Pathology, University of Oxford, Oxford, UK To gain insight into the molecular pathogenesis of the the World Health Organization.6,7 Patients with refractory myelodysplastic syndromes (MDS), we performed global gene anemia (RA) with or without ringed sideroblasts, according to expression profiling and pathway analysis on the hemato- poietic stem cells (HSC) of 183 MDS patients as compared with the the French–American–British classification, were subdivided HSC of 17 healthy controls. The most significantly deregulated based on the presence or absence of multilineage dysplasia. In pathways in MDS include interferon signaling, thrombopoietin addition, patients with RA with excess blasts (RAEB) were signaling and the Wnt pathways. Among the most signifi- subdivided into two categories, RAEB1 and RAEB2, based on the cantly deregulated gene pathways in early MDS are immuno- percentage of bone marrow blasts. -
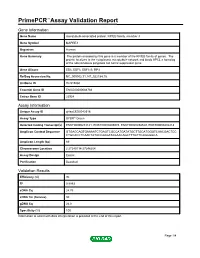
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name microtubule-associated protein, RP/EB family, member 3 Gene Symbol MAPRE3 Organism Human Gene Summary The protein encoded by this gene is a member of the RP/EB family of genes. The protein localizes to the cytoplasmic microtubule network and binds APCL a homolog of the adenomatous polyposis coli tumor suppressor gene. Gene Aliases EB3, EBF3, EBF3-S, RP3 RefSeq Accession No. NC_000002.11, NT_022184.15 UniGene ID Hs.515860 Ensembl Gene ID ENSG00000084764 Entrez Gene ID 22924 Assay Information Unique Assay ID qHsaCED0042518 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000233121, ENST00000405074, ENST00000458529, ENST00000402218 Amplicon Context Sequence GTGACCAGTGAAAATCTGAGTCGCCATGATATGCTTGCATGGGTCAACGACTCC CTGCACCTCAACTATACCAAGATAGAACAGCTTTGTTCAGGGGCA Amplicon Length (bp) 69 Chromosome Location 2:27245114-27246204 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 90 R2 0.9993 cDNA Cq 24.76 cDNA Tm (Celsius) 80 gDNA Cq 26.8 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 1/4 PrimePCR™Assay Validation Report MAPRE3, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 2/4 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Human Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. -

Whole Exome Sequencing in ADHD Trios from Single and Multi-Incident Families Implicates New Candidate Genes and Highlights Polygenic Transmission
European Journal of Human Genetics (2020) 28:1098–1110 https://doi.org/10.1038/s41431-020-0619-7 ARTICLE Whole exome sequencing in ADHD trios from single and multi-incident families implicates new candidate genes and highlights polygenic transmission 1,2 1 1,2 1,3 1 Bashayer R. Al-Mubarak ● Aisha Omar ● Batoul Baz ● Basma Al-Abdulaziz ● Amna I. Magrashi ● 1,2 2 2,4 2,4,5 6 Eman Al-Yemni ● Amjad Jabaan ● Dorota Monies ● Mohamed Abouelhoda ● Dejene Abebe ● 7 1,2 Mohammad Ghaziuddin ● Nada A. Al-Tassan Received: 26 September 2019 / Revised: 26 February 2020 / Accepted: 10 March 2020 / Published online: 1 April 2020 © The Author(s) 2020. This article is published with open access Abstract Several types of genetic alterations occurring at numerous loci have been described in attention deficit hyperactivity disorder (ADHD). However, the role of rare single nucleotide variants (SNVs) remains under investigated. Here, we sought to identify rare SNVs with predicted deleterious effect that may contribute to ADHD risk. We chose to study ADHD families (including multi-incident) from a population with a high rate of consanguinity in which genetic risk factors tend to 1234567890();,: 1234567890();,: accumulate and therefore increasing the chance of detecting risk alleles. We employed whole exome sequencing (WES) to interrogate the entire coding region of 16 trios with ADHD. We also performed enrichment analysis on our final list of genes to identify the overrepresented biological processes. A total of 32 rare variants with predicted damaging effect were identified in 31 genes. At least two variants were detected per proband, most of which were not exclusive to the affected individuals. -

Association of the PSRC1 Rs599839 Variant with Coronary Artery Disease in a Mexican Population
medicina Communication Association of the PSRC1 rs599839 Variant with Coronary Artery Disease in a Mexican Population Martha Eunice Rodríguez-Arellano 1, Jacqueline Solares-Tlapechco 1, Paula Costa-Urrutia 1 , Helios Cárdenas-Hernández 1, Marajael Vallejo-Gómez 1, Julio Granados 2 and Sergio Salas-Padilla 3,* 1 Laboratorio de Medicina Genómica, Hospital Regional Lic. Adolfo López Mateos ISSSTE, Ciudad de Mexico 01030, Mexico; [email protected] (M.E.R.-A.); [email protected] (J.S.-T.); [email protected] (P.C.-U.); [email protected] (H.C.-H.); [email protected] (M.V.-G.) 2 División de Inmunogenética, Departamento de Trasplantes, Instituto Nacional de Ciencias Médicas y Nutrición Salvador Zubirán, Ciudad de Mexico 14080, Mexico; [email protected] 3 Servicio de Cardiología, Hospital Regional Lic. Adolfo López Mateos ISSSTE, Ciudad de Mexico 01030, Mexico * Correspondence: [email protected]; Tel.: +52-555-322-2200 Received: 3 July 2020; Accepted: 12 August 2020; Published: 26 August 2020 Abstract: Background and Objectives: Coronary artery disease (CAD) is a major health problem in México. The identification of modifiable risk factors and genetic biomarkers is crucial for an integrative and personalized CAD risk evaluation. In this work, we aimed to validate in a Mexican population a set of eight selected polymorphisms previously associated with CAD, myocardial infarction (MI), or dyslipidemia. Materials and Methods: A sample of 907 subjects (394 CAD cases and 513 controls) 40–80 years old was genotyped for eight loci: PSRC1 (rs599839), MRAS (rs9818870), BTN2A1 (rs6929846), MTHFD1L (rs6922269), CDKN2B (rs1333049), KIAA1462 (rs3739998), CXCL12 (rs501120), and HNF1A (rs2259816). The association between single nucleotide polymorphisms (SNPs) and CAD was evaluated by logistic regression models. -

Molecular Associations and Clinical Significance of Raps In
ORIGINAL RESEARCH published: 21 June 2021 doi: 10.3389/fmolb.2021.677979 Molecular Associations and Clinical Significance of RAPs in Hepatocellular Carcinoma Sarita Kumari 1†, Mohit Arora 2†, Jay Singh 1, Lokesh K. Kadian 2, Rajni Yadav 3, Shyam S. Chauhan 2* and Anita Chopra 1* 1Laboratory Oncology Unit, Dr. BRA-IRCH, All India Institute of Medical Sciences, New Delhi, India, 2Department of Biochemistry, All India Institute of Medical Sciences, New Delhi, India, 3Department of Pathology, All India Institute of Medical Sciences, New Delhi, India Hepatocellular carcinoma (HCC) is an aggressive gastrointestinal malignancy with a high rate of mortality. Multiple studies have individually recognized members of RAP gene family as critical regulators of tumor progression in several cancers, including hepatocellular carcinoma. These studies suffer numerous limitations including a small sample size and lack of analysis of various clinicopathological and molecular Edited by: Veronica Aran, features. In the current study, we utilized authoritative multi-omics databases to Instituto Estadual do Cérebro Paulo determine the association of RAP gene family expression and detailed molecular and Niemeyer (IECPN), Brazil clinicopathological features in hepatocellular carcinoma (HCC). All five RAP genes Reviewed by: were observed to harbor dysregulated expression in HCC compared to normal liver Pooja Panwalkar, Weill Cornell Medicine, United States tissues. RAP2A exhibited strongest ability to differentiate tumors from the normal Jasminka Omerovic, tissues. RAP2A expression was associated with progressive tumor grade, TP53 and University of Split, Croatia CTNNB1 mutation status. Additionally, RAP2A expression was associated with the *Correspondence: Anita Chopra alteration of its copy numbers and DNA methylation. RAP2A also emerged as an [email protected] independent marker for patient prognosis. -

EB3 (MAPRE3) (NM 012326) Human Tagged ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC204135 EB3 (MAPRE3) (NM_012326) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: EB3 (MAPRE3) (NM_012326) Human Tagged ORF Clone Tag: Myc-DDK Symbol: MAPRE3 Synonyms: EB3; EBF3; EBF3-S; RP3 Vector: pCMV6-Entry (PS100001) E. coli Selection: Kanamycin (25 ug/mL) Cell Selection: Neomycin ORF Nucleotide >RC204135 ORF sequence Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGGCCGTCAATGTGTACTCCACATCTGTGACCAGTGAAAATCTGAGTCGCCATGATATGCTTGCATGGG TCAACGACTCCCTGCACCTCAACTATACCAAGATAGAACAGCTTTGTTCAGGGGCAGCCTACTGCCAGTT CATGGACATGCTCTTCCCCGGCTGTGTGCACTTGAGGAAAGTGAAGTTCCAGGCCAAACTAGAGCATGAA TACATCCACAACTTCAAGGTGCTGCAAGCAGCTTTCAAGAAGATGGGTGTTGACAAAATCATTCCTGTAG AGAAATTAGTGAAAGGAAAATTCCAAGATAATTTTGAGTTTATTCAGTGGTTTAAGAAATTCTTTGACGC AAACTATGATGGAAAGGATTACAACCCTCTGCTGGCGCGGCAGGGCCAGGACGTAGCGCCACCTCCTAAC CCAGGTGATCAGATCTTCAACAAATCCAAGAAACTCATTGGCACAGCAGTTCCACAGAGGACGTCCCCCA CAGGCCCAAAAAACATGCAGACCTCTGGCCGGCTGAGCAATGTGGCCCCCCCCTGCATTCTCCGGAAGAA TCCTCCATCAGCCCGAAATGGCGGCCATGAGACTGATGCCCAAATTCTTGAACTCAACCAACAGCTGGTG GACTTGAAGCTGACAGTGGATGGGCTGGAGAAGGAACGTGACTTCTACTTCAGCAAACTTCGTGACATCG AGCTCATCTGCCAGGAGCATGAAAGTGAAAACAGCCCTGTTATCTCAGGCATCATTGGCATCCTCTATGC CACAGAGGAAGGATTCGCACCCCCTGAGGACGATGAGATTGAAGAGCATCAACAAGAAGACCAGGACGAG TAC ACGCGTACGCGGCCGCTCGAGCAGAAACTCATCTCAGAAGAGGATCTGGCAGCAAATGATATCCTGGATT -

Supporting Information
Supporting Information Pouryahya et al. SI Text Table S1 presents genes with the highest absolute value of Ricci curvature. We expect these genes to have significant contribution to the network’s robustness. Notably, the top two genes are TP53 (tumor protein 53) and YWHAG gene. TP53, also known as p53, it is a well known tumor suppressor gene known as the "guardian of the genome“ given the essential role it plays in genetic stability and prevention of cancer formation (1, 2). Mutations in this gene play a role in all stages of malignant transformation including tumor initiation, promotion, aggressiveness, and metastasis (3). Mutations of this gene are present in more than 50% of human cancers, making it the most common genetic event in human cancer (4, 5). Namely, p53 mutations play roles in leukemia, breast cancer, CNS cancers, and lung cancers, among many others (6–9). The YWHAG gene encodes the 14-3-3 protein gamma, a member of the 14-3-3 family proteins which are involved in many biological processes including signal transduction regulation, cell cycle pro- gression, apoptosis, cell adhesion and migration (10, 11). Notably, increased expression of 14-3-3 family proteins, including protein gamma, have been observed in a number of human cancers including lung and colorectal cancers, among others, suggesting a potential role as tumor oncogenes (12, 13). Furthermore, there is evidence that loss Fig. S1. The histogram of scalar Ricci curvature of 8240 genes. Most of the genes have negative scalar Ricci curvature (75%). TP53 and YWHAG have notably low of p53 function may result in upregulation of 14-3-3γ in lung cancer Ricci curvatures. -

Role and Regulation of the P53-Homolog P73 in the Transformation of Normal Human Fibroblasts
Role and regulation of the p53-homolog p73 in the transformation of normal human fibroblasts Dissertation zur Erlangung des naturwissenschaftlichen Doktorgrades der Bayerischen Julius-Maximilians-Universität Würzburg vorgelegt von Lars Hofmann aus Aschaffenburg Würzburg 2007 Eingereicht am Mitglieder der Promotionskommission: Vorsitzender: Prof. Dr. Dr. Martin J. Müller Gutachter: Prof. Dr. Michael P. Schön Gutachter : Prof. Dr. Georg Krohne Tag des Promotionskolloquiums: Doktorurkunde ausgehändigt am Erklärung Hiermit erkläre ich, dass ich die vorliegende Arbeit selbständig angefertigt und keine anderen als die angegebenen Hilfsmittel und Quellen verwendet habe. Diese Arbeit wurde weder in gleicher noch in ähnlicher Form in einem anderen Prüfungsverfahren vorgelegt. Ich habe früher, außer den mit dem Zulassungsgesuch urkundlichen Graden, keine weiteren akademischen Grade erworben und zu erwerben gesucht. Würzburg, Lars Hofmann Content SUMMARY ................................................................................................................ IV ZUSAMMENFASSUNG ............................................................................................. V 1. INTRODUCTION ................................................................................................. 1 1.1. Molecular basics of cancer .......................................................................................... 1 1.2. Early research on tumorigenesis ................................................................................. 3 1.3. Developing -

In This Table Protein Name, Uniprot Code, Gene Name P-Value
Supplementary Table S1: In this table protein name, uniprot code, gene name p-value and Fold change (FC) for each comparison are shown, for 299 of the 301 significantly regulated proteins found in both comparisons (p-value<0.01, fold change (FC) >+/-0.37) ALS versus control and FTLD-U versus control. Two uncharacterized proteins have been excluded from this list Protein name Uniprot Gene name p value FC FTLD-U p value FC ALS FTLD-U ALS Cytochrome b-c1 complex P14927 UQCRB 1.534E-03 -1.591E+00 6.005E-04 -1.639E+00 subunit 7 NADH dehydrogenase O95182 NDUFA7 4.127E-04 -9.471E-01 3.467E-05 -1.643E+00 [ubiquinone] 1 alpha subcomplex subunit 7 NADH dehydrogenase O43678 NDUFA2 3.230E-04 -9.145E-01 2.113E-04 -1.450E+00 [ubiquinone] 1 alpha subcomplex subunit 2 NADH dehydrogenase O43920 NDUFS5 1.769E-04 -8.829E-01 3.235E-05 -1.007E+00 [ubiquinone] iron-sulfur protein 5 ARF GTPase-activating A0A0C4DGN6 GIT1 1.306E-03 -8.810E-01 1.115E-03 -7.228E-01 protein GIT1 Methylglutaconyl-CoA Q13825 AUH 6.097E-04 -7.666E-01 5.619E-06 -1.178E+00 hydratase, mitochondrial ADP/ATP translocase 1 P12235 SLC25A4 6.068E-03 -6.095E-01 3.595E-04 -1.011E+00 MIC J3QTA6 CHCHD6 1.090E-04 -5.913E-01 2.124E-03 -5.948E-01 MIC J3QTA6 CHCHD6 1.090E-04 -5.913E-01 2.124E-03 -5.948E-01 Protein kinase C and casein Q9BY11 PACSIN1 3.837E-03 -5.863E-01 3.680E-06 -1.824E+00 kinase substrate in neurons protein 1 Tubulin polymerization- O94811 TPPP 6.466E-03 -5.755E-01 6.943E-06 -1.169E+00 promoting protein MIC C9JRZ6 CHCHD3 2.912E-02 -6.187E-01 2.195E-03 -9.781E-01 Mitochondrial 2- -

Autoantibodies As Potential Biomarkers in Breast Cancer
biosensors Review Autoantibodies as Potential Biomarkers in Breast Cancer Jingyi Qiu, Bailey Keyser, Zuan-Tao Lin and Tianfu Wu * ID Department of Biomedical Engineering, University of Houston, 3517 Cullen BLVD, SERC 2008, Houston, TX 77204, USA; [email protected] (J.Q.); [email protected] (B.K.); [email protected] (Z.-T.L.) * Correspondence: [email protected]; Tel.: +1-713-743-0142 Received: 14 June 2018; Accepted: 11 July 2018; Published: 13 July 2018 Abstract: Breast cancer is a major cause of mortality in women; however, technologies for early stage screening and diagnosis (e.g., mammography and other imaging technologies) are not optimal for the accurate detection of cancer. This creates demand for a more effective diagnostic means to replace or be complementary to existing technologies for early discovery of breast cancer. Cancer neoantigens could reflect tumorigenesis, but they are hardly detectable at the early stage. Autoantibodies, however, are biologically amplified and hence may be measurable early on, making them promising biomarkers to discriminate breast cancer from healthy tissue accurately. In this review, we summarized the recent findings of breast cancer specific antigens and autoantibodies, which may be useful in early detection, disease stratification, and monitoring of treatment responses of breast cancer. Keywords: autoantibody; breast cancer; early diagnosis; immunotherapy 1. Introduction Breast cancer is the prevailing cancer among women in developing and developed countries [1]. As such, screening and early diagnosis with respect to risk stratification are critical for prevention and early intervention of the disease, leading to better therapeutic outcomes [2,3]. Breast cancer itself is genetically heterogeneous and expresses a variety of aberrant proteins that, until recently, were un-utilizable. -

Identification of Subgroups Along the Glycolysis-Cholesterol Synthesis
Zhang et al. Human Genomics (2021) 15:53 https://doi.org/10.1186/s40246-021-00350-3 PRIMARY RESEARCH Open Access Identification of subgroups along the glycolysis-cholesterol synthesis axis and the development of an associated prognostic risk model Enchong Zhang1†, Yijing Chen2,3†, Shurui Bao4, Xueying Hou3, Jing Hu3, Oscar Yong Nan Mu5, Yongsheng Song1 and Liping Shan1* Abstract Background: Skin cutaneous melanoma (SKCM) is one of the most highly prevalent and complicated malignancies. Glycolysis and cholesterogenesis pathways both play important roles in cancer metabolic adaptations. The main aims of this study are to subtype SKCM based on glycolytic and cholesterogenic genes and to build a clinical outcome predictive algorithm based on the subtypes. Methods: A dataset with 471 SKCM specimens was downloaded from The Cancer Genome Atlas (TCGA) database. We extracted and clustered genes from the Molecular Signatures Database v7.2 and acquired co-expressed glycolytic and cholesterogenic genes. We then subtyped the SKCM samples and validated the efficacy of subtypes with respect to simple nucleotide variations (SNVs), copy number variation (CNV), patients’ survival statuses, tumor microenvironment, and proliferation scores. We also constructed a risk score model based on metabolic subclassification and verified the model using validating datasets. Finally, we explored potential drugs for high-risk SKCM patients. Results: SKCM patients were divided into four subtype groups: glycolytic, cholesterogenic, mixed, and quiescent subgroups. The glycolytic subtype had the worst prognosis and MGAM SNV extent. Compared with the cholesterogenic subgroup, the glycolytic subgroup had higher rates of DDR2 and TPR CNV and higher proliferation scores and MK167 expression levels, but a lower tumor purity proportion. -

The Rs599839 A>G Variant Disentangles
Preprints (www.preprints.org) | NOT PEER-REVIEWED | Posted: 15 March 2021 doi:10.20944/preprints202103.0400.v1 Article The rs599839 A>G variant disentangles cardiovascular risk and hepatocellular carcinoma in NAFLD patients Meroni Marica PhD1,2, Longo Miriam MSc1,3, Paolini Erika MSc1,4, Alisi Anna PhD5, Miele Luca MD6, De Caro Emilia Rita MSc1, Pisano Giuseppina MD1, Maggioni Marco MD7, Soardo Giorgio MD8, Valenti Luca MD2,9, Fracanzani Anna Ludovica MD1,2, Dongiovanni Paola MSc1. 1 General Medicine and Metabolic Diseases, Fondazione IRCCS Ca’ Granda Ospedale Maggiore Policlinico, Milano, Italy; 2 Department of Pathophysiology and Transplantation, Università degli Studi di Milano, Milano, Italy; 3 Department of Clinical Sciences and Community Health, Università degli Studi di Milano, Italy; 4 Department of Pharmacological and Biomolecular Sciences, Università degli Studi di Milano, Italy; 5 Research Unit of Molecular Genetics of Complex Phenotypes, Bambino Gesù Children Hospital, IRCCS, Rome, Italy; 6 Area Medicina Interna, Gastroenterologia e Oncologia Medica, Fondazione Policlinico Universitario A. Gemelli IRCCS, Rome, Italy; 7 Department of Pathology, Fondazione IRCCS Ca’ Granda Ospedale Maggiore Policlinico, Milano, Italy; 8 Clinic of Internal Medicine-Liver Unit Department of Medical Area (DAME) University School of Medicine, Udine, Italy and Italian Liver Foundation AREA Science Park - Basovizza Campus, Trieste, Italy. 9 Translational Medicine, Department of Transfusion Medicine and Hematology, Fondazione IRCCS Ca’ Granda Ospedale Maggiore Policlinico Milano; * Correspondence: [email protected]. Phone: +39-02-5503-3467. Fax +39-02-5503-4229. Abstract: Background and Aims: Dyslipidemia and cardiovascular diseases (CAD) are comorbidi- Citation: PhD, M.M.; MSc, L.M.; ties of nonalcoholic fatty liver disease (NAFLD), which ranges from steatosis to hepatocellular car- MSc, P.E.; PhD, A.A.; MD, M.L.; cinoma (HCC).