Effect of Valine on Propionate Metabolism in Control and Hyperglycinemic Fibroblasts and in Rat Liver
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
Workshop 1 – Biochemistry (Chem 160)
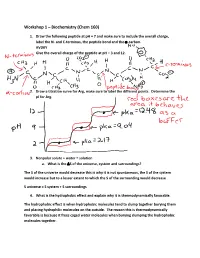
Workshop 1 – Biochemistry (Chem 160) 1. Draw the following peptide at pH = 7 and make sure to include the overall charge, label the N- and C-terminus, the peptide bond and the -carbon. AVDKY Give the overall charge of the peptide at pH = 3 and 12. 2. Draw a titration curve for Arg, make sure to label the different points. Determine the pI for Arg. 3. Nonpolar solute + water = solution a. What is the S of the universe, system and surroundings? The S of the universe would decrease this is why it is not spontaneous, the S of the system would increase but to a lesser extent to which the S of the surrounding would decrease S universe = S system + S surroundings 4. What is the hydrophobic effect and explain why it is thermodynamically favorable. The hydrophobic effect is when hydrophobic molecules tend to clump together burying them and placing hydrophilic molecules on the outside. The reason this is thermodynamically favorable is because it frees caged water molecules when burying clumping the hydrophobic molecules together. 5. Urea dissolves very readily in water, but the solution becomes very cold as the urea dissolves. How is this possible? Urea dissolves in water because when dissolving there is a net increase in entropy of the universe. The heat exchange, getting colder only reflects the enthalpy (H) component of the total energy change. The entropy change is high enough to offset the enthalpy component and to add up to an overall -G 6. A mutation that changes an alanine residue in the interior of a protein to valine is found to lead to a loss of activity. -

Amino Acid Recognition by Aminoacyl-Trna Synthetases
www.nature.com/scientificreports OPEN The structural basis of the genetic code: amino acid recognition by aminoacyl‑tRNA synthetases Florian Kaiser1,2,4*, Sarah Krautwurst3,4, Sebastian Salentin1, V. Joachim Haupt1,2, Christoph Leberecht3, Sebastian Bittrich3, Dirk Labudde3 & Michael Schroeder1 Storage and directed transfer of information is the key requirement for the development of life. Yet any information stored on our genes is useless without its correct interpretation. The genetic code defnes the rule set to decode this information. Aminoacyl-tRNA synthetases are at the heart of this process. We extensively characterize how these enzymes distinguish all natural amino acids based on the computational analysis of crystallographic structure data. The results of this meta-analysis show that the correct read-out of genetic information is a delicate interplay between the composition of the binding site, non-covalent interactions, error correction mechanisms, and steric efects. One of the most profound open questions in biology is how the genetic code was established. While proteins are encoded by nucleic acid blueprints, decoding this information in turn requires proteins. Te emergence of this self-referencing system poses a chicken-or-egg dilemma and its origin is still heavily debated 1,2. Aminoacyl-tRNA synthetases (aaRSs) implement the correct assignment of amino acids to their codons and are thus inherently connected to the emergence of genetic coding. Tese enzymes link tRNA molecules with their amino acid cargo and are consequently vital for protein biosynthesis. Beside the correct recognition of tRNA features3, highly specifc non-covalent interactions in the binding sites of aaRSs are required to correctly detect the designated amino acid4–7 and to prevent errors in biosynthesis5,8. -

Amino Acid Chemistry
Handout 4 Amino Acid and Protein Chemistry ANSC 619 PHYSIOLOGICAL CHEMISTRY OF LIVESTOCK SPECIES Amino Acid Chemistry I. Chemistry of amino acids A. General amino acid structure + HN3- 1. All amino acids are carboxylic acids, i.e., they have a –COOH group at the #1 carbon. 2. All amino acids contain an amino group at the #2 carbon (may amino acids have a second amino group). 3. All amino acids are zwitterions – they contain both positive and negative charges at physiological pH. II. Essential and nonessential amino acids A. Nonessential amino acids: can make the carbon skeleton 1. From glycolysis. 2. From the TCA cycle. B. Nonessential if it can be made from an essential amino acid. 1. Amino acid "sparing". 2. May still be essential under some conditions. C. Essential amino acids 1. Branched chain amino acids (isoleucine, leucine and valine) 2. Lysine 3. Methionine 4. Phenyalanine 5. Threonine 6. Tryptophan 1 Handout 4 Amino Acid and Protein Chemistry D. Essential during rapid growth or for optimal health 1. Arginine 2. Histidine E. Nonessential amino acids 1. Alanine (from pyruvate) 2. Aspartate, asparagine (from oxaloacetate) 3. Cysteine (from serine and methionine) 4. Glutamate, glutamine (from α-ketoglutarate) 5. Glycine (from serine) 6. Proline (from glutamate) 7. Serine (from 3-phosphoglycerate) 8. Tyrosine (from phenylalanine) E. Nonessential and not required for protein synthesis 1. Hydroxyproline (made postranslationally from proline) 2. Hydroxylysine (made postranslationally from lysine) III. Acidic, basic, polar, and hydrophobic amino acids A. Acidic amino acids: amino acids that can donate a hydrogen ion (proton) and thereby decrease pH in an aqueous solution 1. -

Solutions to 7.012 Problem Set 1
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Solutions to 7.012 Problem Set 1 Question 1 Bob, a student taking 7.012, looks at a long-standing puddle outside his dorm window. Curious as to what was growing in the cloudy water, he takes a sample to his TA, Brad Student. He wanted to know whether the organisms in the sample were prokaryotic or eukaryotic. a) Give an example of a prokaryotic and a eukaryotic organism. Prokaryotic: Eukaryotic: All bacteria Yeast, fungi, any animial or plant b) Using a light microscope, how could he tell the difference between a prokaryotic organism and a eukaryotic one? The resolution of the light microscope would allow you to see if the cell had a true nucleus or organelles. A cell with a true nucleus and organelles would be eukaryotic. You could also determine size, but that may not be sufficient to establish whether a cell is prokaryotic or eukaryotic. c) What additional differences exist between prokaryotic and eukaryotic organisms? Any answer from above also fine here. In addition, prokaryotic and eukaryotic organisms differ at the DNA level. Eukaryotes have more complex genomes than prokaryotes do. Question 2 A new startup company hires you to help with their product development. Your task is to find a protein that interacts with a polysaccharide. a) You find a large protein that has a single binding site for the polysaccharide cellulose. Which amino acids might you expect to find in the binding pocket of the protein? What is the strongest type of interaction possible between these amino acids and the cellulose? Cellulose is a polymer of glucose and as such has many free hydroxyl groups. -

Nucleotide Base Coding and Am1ino Acid Replacemients in Proteins* by Emil L
VOL. 48, 1962 BIOCHEMISTRY: E. L. SAIITH 677 18 Britten, R. J., and R. B. Roberts, Science, 131, 32 (1960). '9 Crestfield, A. M., K. C. Smith, and F. WV. Allen, J. Biol. Chem., 216, 185 (1955). 20 Gamow, G., Nature, 173, 318 (1954). 21 Brenner, S., these PROCEEDINGS, 43, 687 (1957). 22 Nirenberg, M. WV., J. H. Matthaei, and 0. WV. Jones, unpublished data. 23 Crick, F. H. C., L. Barnett, S. Brenner, and R. J. Watts-Tobin, Nature, 192, 1227 (1961). 24 Levene, P. A., and R. S. Tipson, J. Biol. Ch-nn., 111, 313 (1935). 25 Gierer, A., and K. W. Mundry, Nature, 182, 1437 (1958). 2' Tsugita, A., and H. Fraenkel-Conrat, J. Mllot. Biol., in press. 27 Tsugita, A., and H. Fraenkel-Conrat, personal communication. 28 Wittmann, H. G., Naturwissenschaften, 48, 729 (1961). 29 Freese, E., in Structure and Function of Genetic Elements, Brookhaven Symposia in Biology, no. 12 (1959), p. 63. NUCLEOTIDE BASE CODING AND AM1INO ACID REPLACEMIENTS IN PROTEINS* BY EMIL L. SMITHt LABORATORY FOR STUDY OF HEREDITARY AND METABOLIC DISORDERS AND THE DEPARTMENTS OF BIOLOGICAL CHEMISTRY AND MEDICINE, UNIVERSITY OF UTAH COLLEGE OF MEDICINE Communicated by Severo Ochoa, February 14, 1962 The problem of which bases of messenger or template RNA' specify the coding of amino acids in proteins has been largely elucidated by the use of synthetic polyri- bonucleotides.2-7 For these triplet nucleotide compositions (Table 1), it is of in- terest to examine some of the presently known cases of amino acid substitutions in polypeptides or proteins of known structure. -

The Role of Reduced Methionine in Mediating the Metabolic Responses to Protein Restriction Using Different Sources of Protein
nutrients Article The Role of Reduced Methionine in Mediating the Metabolic Responses to Protein Restriction Using Different Sources of Protein Han Fang 1 , Kirsten P. Stone 1 , Sujoy Ghosh 2,3 , Laura A. Forney 4 and Thomas W. Gettys 1,* 1 Laboratory of Nutrient Sensing & Adipocyte Signaling, 6400 Perkins Road, Pennington Biomedical Research Center, Baton Rouge, LA 70808, USA; [email protected] (H.F.); [email protected] (K.P.S.) 2 Laboratory of Computational Biology, Pennington Biomedical Research Center, Baton Rouge, LA 70808, USA; [email protected] 3 Program in Cardiovascular and Metabolic Disorders and Center for Computational Biology, Duke-NUS Medical School, Singapore 169857, Singapore 4 Department of Integrative Biology and Pharmacology, University of Texas Health Science Center at Houston, 7000 Fannin St, Houston, TX 77030, USA; [email protected] * Correspondence: [email protected] Abstract: Dietary protein restriction and dietary methionine restriction (MR) produce a comparable series of behavioral, physiological, biochemical, and transcriptional responses. Both dietary regimens produce a similar reduction in intake of sulfur amino acids (e.g., methionine and cystine), and both diets increase expression and release of hepatic FGF21. Given that FGF21 is an essential mediator of the metabolic phenotype produced by both diets, an important unresolved question is whether dietary protein restriction represents de facto methionine restriction. Using diets formulated from either casein or soy protein with matched reductions in sulfur amino acids, we compared the ability Citation: Fang, H.; Stone, K.P.; of the respective diets to recapitulate the metabolic phenotype produced by methionine restriction Ghosh, S.; Forney, L.A.; Gettys, T.W. -

Amino Acid Requirements of the Free-Living Nematode Caenorhabditis Briggsae
AMINO ACID REQUIREMENTS OF THE FREE-LIVING NEMATODE CAENORHABDITIS BRIGGSAE BY J. R. VANFLETEREN Instituut voor Dierkunde, Laboratoria voor Morfologie en Systematiek, RijksuniversiteitGent, Belgium Washed yeast ribosomes promote growth and reproduction of C. briggsae, even when supple- mented to the basal medium at dosages too low to provide the organisms with sufficient amounts of essential amino acids. Hence, a re-investigation of the amino acid requirements of C. briggsae by single and multiple omission of amino acids from the basal medium revealed unambiguously that arginine, histidine, lysine, tryptophan, phenylalanine, methionine, threonine, leucine, isoleucine and valine are not synthetized at levels to permit reproduction; they are called essential amino acids. The requirement for arginine and isoleucinehowever appears to be less clear-cut. On the contrary, evidence is presented that alanine, asparagine, cysteine, glutamate, glutamine, glycine, proline, serine and tryosine can be synthetized at adequate levels; they are called non- essential amino acids. In addition it was shown that multiple omission of the non-essential amino acids is not deleterious. This is believed to be an important step towards the development of a minimum essential medium (MEM) for growth and reproduction of C. briggsae. Sustained growth of the free-living nematode Caenorhabditis brigg.rae can be obtained on a chemically defined medium, supplemented with adequate levels of a proteinaceous growth factor. The most satisfactory, chemically defined medium hitherto reported (Buecher, Hansen & Yarwood, 1966), has been called C. brigg.iae Maintenance Medium (CbMM) and is now commercially available. CbMM is an extremely rich medium, being composed of 53 components, all present at high concentrations. -

Recent Progress in the Microbial Production of Pyruvic Acid
fermentation Review Recent Progress in the Microbial Production of Pyruvic Acid Neda Maleki 1 and Mark A. Eiteman 2,* 1 Department of Food Science, Engineering and Technology, University of Tehran, Karaj 31587-77871, Iran; [email protected] 2 School of Chemical, Materials and Biomedical Engineering, University of Georgia, Athens, GA 30602, USA * Correspondence: [email protected]; Tel.: +1-706-542-0833 Academic Editor: Gunnar Lidén Received: 10 January 2017; Accepted: 6 February 2017; Published: 13 February 2017 Abstract: Pyruvic acid (pyruvate) is a cellular metabolite found at the biochemical junction of glycolysis and the tricarboxylic acid cycle. Pyruvate is used in food, cosmetics, pharmaceutical and agricultural applications. Microbial production of pyruvate from either yeast or bacteria relies on restricting the natural catabolism of pyruvate, while also limiting the accumulation of the numerous potential by-products. In this review we describe research to improve pyruvate formation which has targeted both strain development and process development. Strain development requires an understanding of carbohydrate metabolism and the many competing enzymes which use pyruvate as a substrate, and it often combines classical mutation/isolation approaches with modern metabolic engineering strategies. Process development requires an understanding of operational modes and their differing effects on microbial growth and product formation. Keywords: auxotrophy; Candida glabrata; Escherichia coli; fed-batch; metabolic engineering; pyruvate; pyruvate dehydrogenase 1. Introduction Pyruvic acid (pyruvate at neutral pH) is a three carbon oxo-monocarboxylic acid, also known as 2-oxopropanoic acid, 2-ketopropionic acid or acetylformic acid. Pyruvate is biochemically located at the end of glycolysis and entry into the tricarboxylic acid (TCA) cycle (Figure1). -

The Cardioprotective Effects of the New Crystal Form of Puerarin in Isoproterenol-Induced Myocardial Ischemia Rats Based on Meta
www.nature.com/scientificreports OPEN The cardioprotective efects of the new crystal form of puerarin in isoproterenol‑induced myocardial ischemia rats based on metabolomics Yuzhi Zhou1,2,3, Mengru Li3, Jia Song3, Yongqiang Shi2, Xuemei Qin3, Zhaolin Gao2, Yang Lv1 & Guanhua Du1* Puerarin has shown unique pharmacological efects on myocardial ischemia (MI). Changing the crystal form is an efective approach to improve the cardioprotective efects of puerarin. However, the mechanisms of the new crystal form of puerarin are unclear. In this study, an electrocardiogram, echocardiography, cardiac marker enzymatic activity, oxidative stress indices, and myocardial histology analysis of cardiac tissues were performed to evaluate the cardioprotective efects of the new crystal form of puerarin. Moreover, serum and cardiac tissue metabolomics based on nuclear magnetic resonance (NMR) were used to investigate the potential mechanism of the new crystal form. The results indicated that the new crystal form of puerarin (30 mg/kg) could improve oxidative stress indices, and these improvements were similar to those of the original crystal form of puerarin (120 mg/ kg). The new crystal form of puerarin (30 mg/kg) could efectively improve the activities of cardiac marker enzymes, and the improvement efects were better than those of the original crystal form (120 mg/kg). Moreover, metabolomics analysis showed that amino acid metabolism, oxidative stress and energy metabolism were disturbed after MI and could be improved by puerarin. These results demonstrated -

Radhakrishnan, Iii 4
NATURE OF THE GENETIC BLOCKS IN THE ISOLEUCINE-VALINE MUTANTS OF SALMONELLA R. P. WAGNER AND ARLOA BERGQUIST The Genetics Laboratory of the Department of Zoology, The Uniuersity of Texas, Austin, Texas Received April 26, 1960 HE metabolic pathways leading to the biosynthesis of isoleucine and valine are now sufficiently well understood, as a result of the work of a number of investigators ( STRASSMAN,THOMAS and WEINHOUSE1955; STRASSMAN,THOMAS, LOCKEand WEINHOUSE1956; STRASSMAN,SHATTON, CORSEY and WEINHOUSE 1958; UMBARGER1958a,b; ADELBERG1955; WAGNER,RADHAKRISHNAN and SNELL1958; RADHAKRISHNAN,WAGNER and SNELL1960; and RADHAKRISHNAN and SNELL1960) to make it possible to investigate the nature of the genetic blocks in mutant organisms requiring isoleucine and valine. This communication describes the results of the investigation of a series of mutants of Salmonella typhimurium originally isolated in the laboratory of DR. M. DEMEREC,The Carnegie Institute of Washington, Cold Spring Harbor, New York. It is limited to the purely biochemical aspects of these mutants, but is ,preceded by a com- munication from GLANVILLEand DEMEREC1960, which describes the linkage studies made with these mutants, and correlates the genetic with the biochemical 3ata. The biosynthesis of isoleucine and valine is believed to occur as shown in Figure 1. In the work to be described here only the steps proceeding from the 3-keto acids, a-acetolactic acid and a-aceto-P-hydroxybutyricacid, have been :onsidered in detail. The following abbreviations are used in Figure I and in mbsequent parts of this communication: AHB = a-aceto-a-hydroxybutyric acid; CHa C Ha CHa CHa I 1 I I c*o CHa-C-OH CHfC-OH CHI-C-H 1 I TPNH 1 I c.0 ~ *Valine I COOH COOH I COOH I COOH (PYRUVIC ACID) (ALI I (HKVI I (DHV) I I I I I I I I + Pyruvic Acid Step1 StepII Stepm Step H 4 I I I I cn3 CH, I CH3 I I I I I CHZ C=O I CH~CH~C-OH Threonine-GO I ~CH3CH2-~-OH-C=0I+! I I I I COOH COOH COOH COOH COOH (U-kmtobutyric acid1 (AH01 (HKII (DUI) (KI) FIGURE1 .-The biosynthetic pathway leading to isoleucine and valine. -
Genetic Code Reading Frame Table
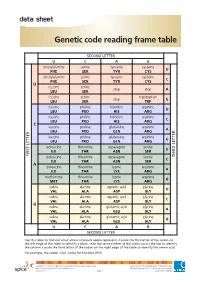
data sheet Genetic code reading frame table SECOND LETTER U C A G phenylalanine serine tyrosine cysteine U PHE SER TYR CYS phenylalanine serine tyrosine cysteine C PHE SER TYR CYS U leucine serine stop stop A LEU SER leucine serine tryptophan stop G LEU SER TRP leucine proline histidine arginine U LEU PRO HIS ARG leucine proline histidine arginine C LEU PRO HIS ARG C leucine proline glutamine arginine A LEU PRO GLN ARG leucine proline glutamine arginine G LEU PRO GLN ARG isoleucine threonine asparagine serine U ILE THR ASN SER FIRST LETTER isoleucine threonine asparagine serine LETTERTHIRD C ILE THR ASN SER A isoleucine threonine lysine arginine A ILE THR LYS ARG methionine threonine lysine arginine G MET THR LYS ARG valine alanine aspartic acid glycine U VAL ALA ASP GLY valine alanine aspartic acid glycine C VAL ALA ASP GLY G valine alanine glutamic acid glycine A VAL ALA GLU GLY valine alanine glutamic acid glycine G VAL ALA GLU GLY U C A G SECOND LETTER Use this table to find out what amino acid each codon represents. Locate the first letter of the codon on the left edge of the table to identify a block, then the second letter of the codon across the top to identify the column. Locate the third letter of the codon on the right edge of the table to identify the amino acid. For example, the codon ‘CAU’ codes for histidine (HIS). ast0812 | Proteins 3: Genetic code reading frame table (data sheet) developed for the Department of Education WA © The University of Western Australia 2012 for conditions of use see spice.wa.edu.au/usage version 1.0 page 1 Licensed for NEALS. -

Amino Acid Degradation
BI/CH 422/622 OUTLINE: OUTLINE: Protein Degradation (Catabolism) Digestion Amino-Acid Degradation Inside of cells Protein turnover Dealing with the carbon Ubiquitin Fates of the 29 Activation-E1 Seven Families Conjugation-E2 nitrogen atoms in 20 1. ADENQ Ligation-E3 AA: Proteosome 2. RPH 9 ammonia oxidase Amino-Acid Degradation 18 transamination Ammonia 2 urea one-carbon metabolism free transamination-mechanism to know THF Urea Cycle – dealing with the nitrogen SAM 5 Steps Carbamoyl-phosphate synthetase 3. GSC Ornithine transcarbamylase PLP uses Arginino-succinate synthetase Arginino-succinase 4. MT – one carbon metabolism Arginase 5. FY – oxidase vs oxygenase Energetics Urea Bi-cycle 6. KW – Urea Cycle – dealing with the nitrogen 7. BCAA – VIL Feeding the Urea Cycle Glucose-Alanine Cycle Convergence with Fatty acid-odd chain Free Ammonia Overview Glutamine Glutamate dehydrogenase Overall energetics Amino Acid A. Concepts 1. ConvergentDegradation 2. ketogenic/glucogenic 3. Reactions seen before The SEVEN (7) Families B. Transaminase (A,D,E) / Deaminase (Q,N) Family C. Related to biosynthesis (R,P,H; C,G,S; M,T) 1.Glu Family a. Introduce oxidases/oxygenases b. Introduce one-carbon metabolism (1C) 2.Pyruvate Family a. PLP reactions 3. a-Ketobutyric Family (M,T) a. 1-C metabolism D. Dedicated 1. Aromatic Family (F,Y) a. oxidases/oxygenases 2. a-Ketoadipic Family (K,W) 3. Branched-chain Family (V,I,L) E. Convergence with Fatty Acids: propionyl-CoA 29 N 1 Amino Acid Degradation • Intermediates of the central metabolic pathway • Some amino acids result in more than one intermediate. • Ketogenic amino acids can be converted to ketone bodies.