Engineering Applications of Artificial Intelligence a Data Mining Framework Based on Boundary-Points for Gene Selection From
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Methamphetamine Induces Alterations in the Long Non-Coding Rnas Expression Profile in the Nucleus Accumbens of the Mouse
Zhu et al. BMC Neuroscience (2015) 16:18 DOI 10.1186/s12868-015-0157-3 RESEARCH ARTICLE Open Access Methamphetamine induces alterations in the long non-coding RNAs expression profile in the nucleus accumbens of the mouse Li Zhu1,2†,JieZhu1,2†,YufengLiu3, Yanjiong Chen4, Yanlin Li1,2,LirenHuang3,SisiChen3,TaoLi1,2,YonghuiDang1,2 andTengChen1,2* Abstract Background: Repeated exposure to addictive drugs elicits long-lasting cellular and molecular changes. It has been reported that the aberrant expression of long non-coding RNAs (lncRNAs) is involved in cocaine and heroin addiction, yet the expression profile of lncRNAs and their potential effects on methamphetamine (METH)-induced locomotor sensitization are largely unknown. Results: Using high-throughput strand-specific complementary DNA sequencing technology (ssRNA-seq), here we examined the alterations in the lncRNAs expression profile in the nucleus accumbens (NAc) of METH-sensitized mice. We found that the expression levels of 6246 known lncRNAs (6215 down-regulated, 31 up-regulated) and 8442 novel lncRNA candidates (8408 down-regulated, 34 up-regulated) were significantly altered in the METH-sensitized mice. Based on characterizations of the genomic contexts of the lncRNAs, we further showed that there were 5139 differentially expressed lncRNAs acted via cis mechanisms, including sense intronic (4295 down-regulated and one up-regulated), overlapping (25 down-regulated and one up-regulated), natural antisense transcripts (NATs, 148 down-regulated and eight up-regulated), long intergenic non-coding RNAs (lincRNAs, 582 down-regulated and five up-regulated), and bidirectional (72 down-regulated and two up-regulated). Moreover, using the program RNAplex, we identified 3994 differentially expressed lncRNAs acted via trans mechanisms. -

Claire Hastie Thesis
Hastie, Claire E. (2011) Discovering common genetic variants for hypertension using an extreme case-control strategy. PhD thesis. http://theses.gla.ac.uk/2423/ Copyright and moral rights for this thesis are retained by the author A copy can be downloaded for personal non-commercial research or study, without prior permission or charge This thesis cannot be reproduced or quoted extensively from without first obtaining permission in writing from the Author The content must not be changed in any way or sold commercially in any format or medium without the formal permission of the Author When referring to this work, full bibliographic details including the author, title, awarding institution and date of the thesis must be given. Glasgow Theses Service http://theses.gla.ac.uk/ [email protected] Discovering Common Genetic Variants for Hypertension Using an Extreme Case-control Strategy Claire E. Hastie, M.Sc. This being a thesis submitted for the degree of Doctor of Philosophy (Ph.D.) in the Faculty of Medicine, University of Glasgow October 2010 BHF Glasgow Cardiovascular Research Centre Institute of Cardiovascular and Medical Sciences College of Medical, Veterinary, and Life Sciences University of Glasgow © C.E. Hastie 2010 Declaration I declare that this thesis has been written entirely by myself and is a record of research performed by myself with the exception of discovery cohort genotyping (Dr Wai K. Lee, Dr Anna Maria Di Blasio, Stewart Laing, and Dr Davide Gentilini), genotyping and association analysis of replication cohorts (undertaken by investigators from each cohort, respectively), and analysis of data from the BRIGHT (Dr Sandosh Padmanabhan), GRECO and HERCULES clinical functional cohorts (undertaken by investigators from each cohort, respectively). -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

Supplementary Material Computational Prediction of SARS
Supplementary_Material Computational prediction of SARS-CoV-2 encoded miRNAs and their putative host targets Sheet_1 List of potential stem-loop structures in SARS-CoV-2 genome as predicted by VMir. Rank Name Start Apex Size Score Window Count (Absolute) Direct Orientation 1 MD13 2801 2864 125 243.8 61 2 MD62 11234 11286 101 211.4 49 4 MD136 27666 27721 104 205.6 119 5 MD108 21131 21184 110 204.7 210 9 MD132 26743 26801 119 188.9 252 19 MD56 9797 9858 128 179.1 59 26 MD139 28196 28233 72 170.4 133 28 MD16 2934 2974 76 169.9 71 43 MD103 20002 20042 80 159.3 403 46 MD6 1489 1531 86 156.7 171 51 MD17 2981 3047 131 152.8 38 87 MD4 651 692 75 140.3 46 95 MD7 1810 1872 121 137.4 58 116 MD140 28217 28252 72 133.8 62 122 MD55 9712 9758 96 132.5 49 135 MD70 13171 13219 93 130.2 131 164 MD95 18782 18820 79 124.7 184 173 MD121 24086 24135 99 123.1 45 176 MD96 19046 19086 75 123.1 179 196 MD19 3197 3236 76 120.4 49 200 MD86 17048 17083 73 119.8 428 223 MD75 14534 14600 137 117 51 228 MD50 8824 8870 94 115.8 79 234 MD129 25598 25642 89 115.6 354 Reverse Orientation 6 MR61 19088 19132 88 197.8 271 10 MR72 23563 23636 148 188.8 286 11 MR11 3775 3844 136 185.1 116 12 MR94 29532 29582 94 184.6 271 15 MR43 14973 15028 109 183.9 226 27 MR14 4160 4206 89 170 241 34 MR35 11734 11792 111 164.2 37 52 MR5 1603 1652 89 152.7 118 53 MR57 18089 18132 101 152.7 139 94 MR8 2804 2864 122 137.4 38 107 MR58 18474 18508 72 134.9 237 117 MR16 4506 4540 72 133.8 311 120 MR34 10010 10048 82 132.7 245 133 MR7 2534 2578 90 130.4 75 146 MR79 24766 24808 75 127.9 59 150 MR65 21528 21576 99 127.4 83 180 MR60 19016 19049 70 122.5 72 187 MR51 16450 16482 75 121 363 190 MR80 25687 25734 96 120.6 75 198 MR64 21507 21544 70 120.3 35 206 MR41 14500 14542 84 119.2 94 218 MR84 26840 26894 108 117.6 94 Sheet_2 List of stable stem-loop structures based on MFE. -

Persistent Transcription-Blocking DNA Lesions Trigger Somatic Growth Attenuation Associated with Longevity
ARTICLES Persistent transcription-blocking DNA lesions trigger somatic growth attenuation associated with longevity George A. Garinis1,2, Lieneke M. Uittenboogaard1, Heike Stachelscheid3,4, Maria Fousteri5, Wilfred van Ijcken6, Timo M. Breit7, Harry van Steeg8, Leon H. F. Mullenders5, Gijsbertus T. J. van der Horst1, Jens C. Brüning4,9, Carien M. Niessen3,9,10, Jan H. J. Hoeijmakers1 and Björn Schumacher1,9,11 The accumulation of stochastic DNA damage throughout an organism’s lifespan is thought to contribute to ageing. Conversely, ageing seems to be phenotypically reproducible and regulated through genetic pathways such as the insulin-like growth factor-1 (IGF-1) and growth hormone (GH) receptors, which are central mediators of the somatic growth axis. Here we report that persistent DNA damage in primary cells from mice elicits changes in global gene expression similar to those occurring in various organs of naturally aged animals. We show that, as in ageing animals, the expression of IGF-1 receptor and GH receptor is attenuated, resulting in cellular resistance to IGF-1. This cell-autonomous attenuation is specifically induced by persistent lesions leading to stalling of RNA polymerase II in proliferating, quiescent and terminally differentiated cells; it is exacerbated and prolonged in cells from progeroid mice and confers resistance to oxidative stress. Our findings suggest that the accumulation of DNA damage in transcribed genes in most if not all tissues contributes to the ageing-associated shift from growth to somatic maintenance that triggers stress resistance and is thought to promote longevity. Ageing represents the progressive functional decline that is exempted levels as a result of pituitary dysfunction (Snell and Ames mice) — have an from evolutionary selection because it largely occurs after reproduc- extended lifespan17–20. -

Lavenex Et Al. Developmental Regulation of Gene Expression and Astrocytic Processes May Explain Selective Hippocampal Vulnerability
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by RERO DOC Digital Library Supplementary Material 1: Lavenex et al. Developmental regulation of gene expression and astrocytic processes may explain selective hippocampal vulnerability Genes differentially expressed in CA1 (Fold change >2), as compared to every other region analyzed (DG, CA3, subiculum, EC) Affymetrix Gene symbol Gene Title Probe Set ID Unigene Entrez Gene Overexpressed (n=8) CACNA1G calcium channel, voltage-dependent, alpha 1G subunit 207869_s_at Hs.591169 8913 CD24 CD24 molecule 209771_x_at Hs.632285 934 CD24 CD24 molecule 216379_x_at Hs.632285 934 DKFZp761H21 hypothetical protein DKFZp761H2121 1553865_a_atHs.134065 171582 NOG Noggin 231798_at Hs.248201 9241 SEZ6L Seizure related 6 homolog (mouse)-like 231650_s_at Hs.194766 23544 SH2D4B SH2 domain containing 4B 1563849_at Hs.147643 387694 SLC35F4 solute carrier family 35, member F4 240799_at Hs.28280 341880 SYNE1 Spectrin repeat containing, nuclear envelope 1 232027_at Hs.12967 23345 MRNA full length insert cDNA clone EUROIMAGE 29222 214896_at Hs.593529 CDNA FLJ42949 fis, clone BRSTN2006583 215126_at Hs.20034 Transcribed locus, moderately similar to NP_055301.1 neuronal thread prote229641_at Hs.49943 Homo sapiens, clone IMAGE:3869276, mRNA 231963_at Hs.26039 Underexpressed (n=108) ACADVL acyl-Coenzyme A dehydrogenase, very long chain 200710_at Hs.437178 37 ACO2 aconitase 2, mitochondrial 200793_s_at Hs.643610 50 ACOT1 /// ACOacyl-CoA thioesterase 2 /// acyl-CoA thioesterase 1 202982_s_at -

BMC Biology Biomed Central
BMC Biology BioMed Central Research article Open Access Classification and nomenclature of all human homeobox genes PeterWHHolland*†1, H Anne F Booth†1 and Elspeth A Bruford2 Address: 1Department of Zoology, University of Oxford, South Parks Road, Oxford, OX1 3PS, UK and 2HUGO Gene Nomenclature Committee, European Bioinformatics Institute (EMBL-EBI), Wellcome Trust Genome Campus, Hinxton, Cambridgeshire, CB10 1SA, UK Email: Peter WH Holland* - [email protected]; H Anne F Booth - [email protected]; Elspeth A Bruford - [email protected] * Corresponding author †Equal contributors Published: 26 October 2007 Received: 30 March 2007 Accepted: 26 October 2007 BMC Biology 2007, 5:47 doi:10.1186/1741-7007-5-47 This article is available from: http://www.biomedcentral.com/1741-7007/5/47 © 2007 Holland et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: The homeobox genes are a large and diverse group of genes, many of which play important roles in the embryonic development of animals. Increasingly, homeobox genes are being compared between genomes in an attempt to understand the evolution of animal development. Despite their importance, the full diversity of human homeobox genes has not previously been described. Results: We have identified all homeobox genes and pseudogenes in the euchromatic regions of the human genome, finding many unannotated, incorrectly annotated, unnamed, misnamed or misclassified genes and pseudogenes. -

Regulatory Architecture of Gene Expression Variation in the 2 Threespine Stickleback, Gasterosteus Aculeatus
bioRxiv preprint doi: https://doi.org/10.1101/055061; this version posted May 24, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 REGULATORY ARCHITECTURE OF GENE EXPRESSION VARIATION IN THE 2 THREESPINE STICKLEBACK, GASTEROSTEUS ACULEATUS. 3 Victoria L. Pritchard 1, Heidi M. Viitaniemi 1, R.J. Scott McCairns 2, Juha Merilä 2, 4 Mikko Nikinmaa 1, Craig R. Primmer 1 & Erica H. Leder 1. 5 1 Division of Genetics and Physiology, Department of Biology, University of Turku, Turku, Finland 6 2 Ecological Genetics Research Unit, Department of Biosciences, University of Helsinki, Helsinki, 7 Finland. 8 9 Abstract 10 Much adaptive evolutionary change is underlain by mutational variation in regions of the genome that 11 regulate gene expression rather than in the coding regions of the genes themselves. An understanding 12 of the role of gene expression variation in facilitating local adaptation will be aided by an 13 understanding of underlying regulatory networks. Here, we characterize the genetic architecture of 14 gene expression variation in the threespine stickleback (Gasterosteus aculeatus), an important model 15 in the study of adaptive evolution. We collected transcriptomic and genomic data from 60 half-sib 16 families using an expression microarray and genotyping-by-sequencing, and located QTL underlying 17 the variation in gene expression (eQTL) in liver tissue using an interval mapping approach. We 18 identified eQTL for several thousand expression traits. Expression was influenced by polymorphism 19 in both cis and trans regulatory regions. Trans eQTL clustered into hotspots. -

(12) Patent Application Publication (10) Pub. No.: US 2009/0269772 A1 Califano Et Al
US 20090269772A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2009/0269772 A1 Califano et al. (43) Pub. Date: Oct. 29, 2009 (54) SYSTEMS AND METHODS FOR Publication Classification IDENTIFYING COMBINATIONS OF (51) Int. Cl. COMPOUNDS OF THERAPEUTIC INTEREST CI2O I/68 (2006.01) CI2O 1/02 (2006.01) (76) Inventors: Andrea Califano, New York, NY G06N 5/02 (2006.01) (US); Riccardo Dalla-Favera, New (52) U.S. Cl. ........... 435/6: 435/29: 706/54; 707/E17.014 York, NY (US); Owen A. (57) ABSTRACT O'Connor, New York, NY (US) Systems, methods, and apparatus for searching for a combi nation of compounds of therapeutic interest are provided. Correspondence Address: Cell-based assays are performed, each cell-based assay JONES DAY exposing a different sample of cells to a different compound 222 EAST 41ST ST in a plurality of compounds. From the cell-based assays, a NEW YORK, NY 10017 (US) Subset of the tested compounds is selected. For each respec tive compound in the Subset, a molecular abundance profile from cells exposed to the respective compound is measured. (21) Appl. No.: 12/432,579 Targets of transcription factors and post-translational modu lators of transcription factor activity are inferred from the (22) Filed: Apr. 29, 2009 molecular abundance profile data using information theoretic measures. This data is used to construct an interaction net Related U.S. Application Data work. Variances in edges in the interaction network are used to determine the drug activity profile of compounds in the (60) Provisional application No. 61/048.875, filed on Apr. -

Nature's Solution to a Hard-Soft Interface Lara Angelika Dorothée Kuntz
Technische Universität München Fakultät für Medizin The enthesis: nature’s solution to a hard-soft interface Lara Angelika Dorothée Kuntz Vollständiger Abdruck der von der Fakultät für Medizin der Technischen Universität München zur Erlangung des akademischen Grades eines Doktors der Naturwissenschaften (Dr. rer. nat.) genehmigten Dissertation. Vorsitzender: Prof. Dr. Gil G. Westmeyer Prüfende der Dissertation: 1. apl. Prof. Dr. Rainer Burgkart 2. Prof. Dr. Andreas Bausch 3. Prof. Dr. Felix Eckstein Die Dissertation wurde am 10.05.2017 bei der Technischen Universität München eingereicht und durch die Fakultät für Medizin am 21.02.2018 angenommen. Prefix This dissertation was part of the biomaterials focus group project “The enthesis: Nature’s solution to a hard-soft junction“ funded by the International Graduate School of Science and Engineering (IGSSE) of the Technical University of Munich. It was a collaborative project between the Technical University of Munich Clinics of Orthopedics/Sports Orthopedics and the Chair of Cellular Biophysics. Herewith I would like to thank everybody who contributed to this thesis directly or indirectly. It was a very lively, interesting time and many have contributed to it being fun and productive. I thank my advisors Professor Rainer Burgkart and Professor Andreas Bausch for supervi- sion and guidance. Thank you, Rainer, for keeping me motivated to dig deep into the Achilles heel with your positive attitude and your passion for the enthesis. Thank you, Andreas, for your scientific advice and for important lessons for life. Thank you both for giving me the freedom to develop project ideas on my own, yet giving me guidance and direction when I felt stuck. -

A Meta-Analysis of the Effects of High-LET Ionizing Radiations in Human Gene Expression
Supplementary Materials A Meta-Analysis of the Effects of High-LET Ionizing Radiations in Human Gene Expression Table S1. Statistically significant DEGs (Adj. p-value < 0.01) derived from meta-analysis for samples irradiated with high doses of HZE particles, collected 6-24 h post-IR not common with any other meta- analysis group. This meta-analysis group consists of 3 DEG lists obtained from DGEA, using a total of 11 control and 11 irradiated samples [Data Series: E-MTAB-5761 and E-MTAB-5754]. Ensembl ID Gene Symbol Gene Description Up-Regulated Genes ↑ (2425) ENSG00000000938 FGR FGR proto-oncogene, Src family tyrosine kinase ENSG00000001036 FUCA2 alpha-L-fucosidase 2 ENSG00000001084 GCLC glutamate-cysteine ligase catalytic subunit ENSG00000001631 KRIT1 KRIT1 ankyrin repeat containing ENSG00000002079 MYH16 myosin heavy chain 16 pseudogene ENSG00000002587 HS3ST1 heparan sulfate-glucosamine 3-sulfotransferase 1 ENSG00000003056 M6PR mannose-6-phosphate receptor, cation dependent ENSG00000004059 ARF5 ADP ribosylation factor 5 ENSG00000004777 ARHGAP33 Rho GTPase activating protein 33 ENSG00000004799 PDK4 pyruvate dehydrogenase kinase 4 ENSG00000004848 ARX aristaless related homeobox ENSG00000005022 SLC25A5 solute carrier family 25 member 5 ENSG00000005108 THSD7A thrombospondin type 1 domain containing 7A ENSG00000005194 CIAPIN1 cytokine induced apoptosis inhibitor 1 ENSG00000005381 MPO myeloperoxidase ENSG00000005486 RHBDD2 rhomboid domain containing 2 ENSG00000005884 ITGA3 integrin subunit alpha 3 ENSG00000006016 CRLF1 cytokine receptor like -
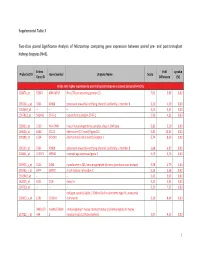
Supplemental Table 3 Two-Class Paired Significance Analysis of Microarrays Comparing Gene Expression Between Paired
Supplemental Table 3 Two‐class paired Significance Analysis of Microarrays comparing gene expression between paired pre‐ and post‐transplant kidneys biopsies (N=8). Entrez Fold q‐value Probe Set ID Gene Symbol Unigene Name Score Gene ID Difference (%) Probe sets higher expressed in post‐transplant biopsies in paired analysis (N=1871) 218870_at 55843 ARHGAP15 Rho GTPase activating protein 15 7,01 3,99 0,00 205304_s_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 6,30 4,50 0,00 1563649_at ‐‐ ‐‐ ‐‐ 6,24 3,51 0,00 1567913_at 541466 CT45‐1 cancer/testis antigen CT45‐1 5,90 4,21 0,00 203932_at 3109 HLA‐DMB major histocompatibility complex, class II, DM beta 5,83 3,20 0,00 204606_at 6366 CCL21 chemokine (C‐C motif) ligand 21 5,82 10,42 0,00 205898_at 1524 CX3CR1 chemokine (C‐X3‐C motif) receptor 1 5,74 8,50 0,00 205303_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 5,68 6,87 0,00 226841_at 219972 MPEG1 macrophage expressed gene 1 5,59 3,76 0,00 203923_s_at 1536 CYBB cytochrome b‐245, beta polypeptide (chronic granulomatous disease) 5,58 4,70 0,00 210135_s_at 6474 SHOX2 short stature homeobox 2 5,53 5,58 0,00 1562642_at ‐‐ ‐‐ ‐‐ 5,42 5,03 0,00 242605_at 1634 DCN decorin 5,23 3,92 0,00 228750_at ‐‐ ‐‐ ‐‐ 5,21 7,22 0,00 collagen, type III, alpha 1 (Ehlers‐Danlos syndrome type IV, autosomal 201852_x_at 1281 COL3A1 dominant) 5,10 8,46 0,00 3493///3 IGHA1///IGHA immunoglobulin heavy constant alpha 1///immunoglobulin heavy 217022_s_at 494 2 constant alpha 2 (A2m marker) 5,07 9,53 0,00 1 202311_s_at