Characterization of Sheath Rot Pathogens from Major Rice-Growing
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Archaea, Bacteria and Termite, Nitrogen Fixation and Sustainable Plants Production
Sun W et al . (2021) Notulae Botanicae Horti Agrobotanici Cluj-Napoca Volume 49, Issue 2, Article number 12172 Notulae Botanicae Horti AcademicPres DOI:10.15835/nbha49212172 Agrobotanici Cluj-Napoca Re view Article Archaea, bacteria and termite, nitrogen fixation and sustainable plants production Wenli SUN 1a , Mohamad H. SHAHRAJABIAN 1a , Qi CHENG 1,2 * 1Chinese Academy of Agricultural Sciences, Biotechnology Research Institute, Beijing 100081, China; [email protected] ; [email protected] 2Hebei Agricultural University, College of Life Sciences, Baoding, Hebei, 071000, China; Global Alliance of HeBAU-CLS&HeQiS for BioAl-Manufacturing, Baoding, Hebei 071000, China; [email protected] (*corresponding author) a,b These authors contributed equally to the work Abstract Certain bacteria and archaea are responsible for biological nitrogen fixation. Metabolic pathways usually are common between archaea and bacteria. Diazotrophs are categorized into two main groups namely: root- nodule bacteria and plant growth-promoting rhizobacteria. Diazotrophs include free living bacteria, such as Azospirillum , Cupriavidus , and some sulfate reducing bacteria, and symbiotic diazotrophs such Rhizobium and Frankia . Three types of nitrogenase are iron and molybdenum (Fe/Mo), iron and vanadium (Fe/V) or iron only (Fe). The Mo-nitrogenase have a higher specific activity which is expressed better when Molybdenum is available. The best hosts for Rhizobium legumiosarum are Pisum , Vicia , Lathyrus and Lens ; Trifolium for Rhizobium trifolii ; Phaseolus vulgaris , Prunus angustifolia for Rhizobium phaseoli ; Medicago, Melilotus and Trigonella for Rhizobium meliloti ; Lupinus and Ornithopus for Lupini, and Glycine max for Rhizobium japonicum . Termites have significant key role in soil ecology, transporting and mixing soil. Termite gut microbes supply the enzymes required to degrade plant polymers, synthesize amino acids, recycle nitrogenous waste and fix atmospheric nitrogen. -

Table S5. the Information of the Bacteria Annotated in the Soil Community at Species Level
Table S5. The information of the bacteria annotated in the soil community at species level No. Phylum Class Order Family Genus Species The number of contigs Abundance(%) 1 Firmicutes Bacilli Bacillales Bacillaceae Bacillus Bacillus cereus 1749 5.145782459 2 Bacteroidetes Cytophagia Cytophagales Hymenobacteraceae Hymenobacter Hymenobacter sedentarius 1538 4.52499338 3 Gemmatimonadetes Gemmatimonadetes Gemmatimonadales Gemmatimonadaceae Gemmatirosa Gemmatirosa kalamazoonesis 1020 3.000970902 4 Proteobacteria Alphaproteobacteria Sphingomonadales Sphingomonadaceae Sphingomonas Sphingomonas indica 797 2.344876284 5 Firmicutes Bacilli Lactobacillales Streptococcaceae Lactococcus Lactococcus piscium 542 1.594633558 6 Actinobacteria Thermoleophilia Solirubrobacterales Conexibacteraceae Conexibacter Conexibacter woesei 471 1.385742446 7 Proteobacteria Alphaproteobacteria Sphingomonadales Sphingomonadaceae Sphingomonas Sphingomonas taxi 430 1.265115184 8 Proteobacteria Alphaproteobacteria Sphingomonadales Sphingomonadaceae Sphingomonas Sphingomonas wittichii 388 1.141545794 9 Proteobacteria Alphaproteobacteria Sphingomonadales Sphingomonadaceae Sphingomonas Sphingomonas sp. FARSPH 298 0.876754244 10 Proteobacteria Alphaproteobacteria Sphingomonadales Sphingomonadaceae Sphingomonas Sorangium cellulosum 260 0.764953367 11 Proteobacteria Deltaproteobacteria Myxococcales Polyangiaceae Sorangium Sphingomonas sp. Cra20 260 0.764953367 12 Proteobacteria Alphaproteobacteria Sphingomonadales Sphingomonadaceae Sphingomonas Sphingomonas panacis 252 0.741416341 -

Understanding the Composition and Role of the Prokaryotic Diversity in the Potato Rhizosphere for Crop Improvement in the Andes
Understanding the composition and role of the prokaryotic diversity in the potato rhizosphere for crop improvement in the Andes Jonas Ghyselinck Dissertation submitted in fulfilment of the requirements for the degree of Doctor (Ph.D.) in Sciences, Biotechnology Promotor - Prof. Dr. Paul De Vos Co-promotor - Dr. Kim Heylen Ghyselinck Jonas – Understanding the composition and role of the prokaryotic diversity in the potato rhizosphere for crop improvement in the Andes Copyright ©2013 Ghyselinck Jonas ISBN-number: 978-94-6197-119-7 No part of this thesis protected by its copyright notice may be reproduced or utilized in any form, or by any means, electronic or mechanical, including photocopying, recording or by any information storage or retrieval system without written permission of the author and promotors. Printed by University Press | www.universitypress.be Ph.D. thesis, Faculty of Sciences, Ghent University, Ghent, Belgium. This Ph.D. work was financially supported by European Community's Seventh Framework Programme FP7/2007-2013 under grant agreement N° 227522 Publicly defended in Ghent, Belgium, May 28th 2013 EXAMINATION COMMITTEE Prof. Dr. Savvas Savvides (chairman) Faculty of Sciences Ghent University, Belgium Prof. Dr. Paul De Vos (promotor) Faculty of Sciences Ghent University, Belgium Dr. Kim Heylen (co-promotor) Faculty of Sciences Ghent University, Belgium Prof. Dr. Anne Willems Faculty of Sciences Ghent University, Belgium Prof. Dr. Peter Dawyndt Faculty of Sciences Ghent University, Belgium Prof. Dr. Stéphane Declerck Faculty of Biological, Agricultural and Environmental Engineering Université catholique de Louvain, Louvain-la-Neuve, Belgium Dr. Angela Sessitsch Department of Health and Environment, Bioresources Unit AIT Austrian Institute of Technology GmbH, Tulln, Austria Dr. -

Saline and Arid Soils: Impact on Bacteria, Plants, and Their Interaction
biology Review Saline and Arid Soils: Impact on Bacteria, Plants, and Their Interaction Elisa Gamalero 1, Elisa Bona 2,* , Valeria Todeschini 2 and Guido Lingua 1 1 Dipartimento di Scienze e Innovazione Tecnologica, Università del Piemonte Orientale, Viale T. Michel 11, 15121 Alessandria, Italy; [email protected] (E.G.); [email protected] (G.L.) 2 Dipartimento di Scienze e Innovazione Tecnologica, Università del Piemonte Orientale, Piazza San Eusebio 5, 13100 Vercelli, Italy; [email protected] * Correspondence: [email protected] Received: 14 April 2020; Accepted: 29 May 2020; Published: 2 June 2020 Abstract: Salinity and drought are the most important abiotic stresses hampering crop growth and yield. It has been estimated that arid areas cover between 41% and 45% of the total Earth area worldwide. At the same time, the world’s population is going to soon reach 9 billion and the survival of this huge amount of people is dependent on agricultural products. Plants growing in saline/arid soil shows low germination rate, short roots, reduced shoot biomass, and serious impairment of photosynthetic efficiency, thus leading to a substantial loss of crop productivity, resulting in significant economic damage. However, plants should not be considered as single entities, but as a superorganism, or a holobiont, resulting from the intimate interactions occurring between the plant and the associated microbiota. Consequently, it is very complex to define how the plant responds to stress on the basis of the interaction with its associated plant growth-promoting bacteria (PGPB). This review provides an overview of the physiological mechanisms involved in plant survival in arid and saline soils and aims at describing the interactions occurring between plants and its bacteriome in such perturbed environments. -

Control of Phytopathogenic Microorganisms with Pseudomonas Sp. and Substances and Compositions Derived Therefrom
(19) TZZ Z_Z_T (11) EP 2 820 140 B1 (12) EUROPEAN PATENT SPECIFICATION (45) Date of publication and mention (51) Int Cl.: of the grant of the patent: A01N 63/02 (2006.01) A01N 37/06 (2006.01) 10.01.2018 Bulletin 2018/02 A01N 37/36 (2006.01) A01N 43/08 (2006.01) C12P 1/04 (2006.01) (21) Application number: 13754767.5 (86) International application number: (22) Date of filing: 27.02.2013 PCT/US2013/028112 (87) International publication number: WO 2013/130680 (06.09.2013 Gazette 2013/36) (54) CONTROL OF PHYTOPATHOGENIC MICROORGANISMS WITH PSEUDOMONAS SP. AND SUBSTANCES AND COMPOSITIONS DERIVED THEREFROM BEKÄMPFUNG VON PHYTOPATHOGENEN MIKROORGANISMEN MIT PSEUDOMONAS SP. SOWIE DARAUS HERGESTELLTE SUBSTANZEN UND ZUSAMMENSETZUNGEN RÉGULATION DE MICRO-ORGANISMES PHYTOPATHOGÈNES PAR PSEUDOMONAS SP. ET DES SUBSTANCES ET DES COMPOSITIONS OBTENUES À PARTIR DE CELLE-CI (84) Designated Contracting States: • O. COUILLEROT ET AL: "Pseudomonas AL AT BE BG CH CY CZ DE DK EE ES FI FR GB fluorescens and closely-related fluorescent GR HR HU IE IS IT LI LT LU LV MC MK MT NL NO pseudomonads as biocontrol agents of PL PT RO RS SE SI SK SM TR soil-borne phytopathogens", LETTERS IN APPLIED MICROBIOLOGY, vol. 48, no. 5, 1 May (30) Priority: 28.02.2012 US 201261604507 P 2009 (2009-05-01), pages 505-512, XP55202836, 30.07.2012 US 201261670624 P ISSN: 0266-8254, DOI: 10.1111/j.1472-765X.2009.02566.x (43) Date of publication of application: • GUANPENG GAO ET AL: "Effect of Biocontrol 07.01.2015 Bulletin 2015/02 Agent Pseudomonas fluorescens 2P24 on Soil Fungal Community in Cucumber Rhizosphere (73) Proprietor: Marrone Bio Innovations, Inc. -

Fungal Flora of Korea
Fungal Flora of Korea Volume 1, Number 2 Ascomycota: Dothideomycetes: Pleosporales: Pleosporaceae Alternaria and Allied Genera 2015 National Institute of Biological Resources Ministry of Environment Fungal Flora of Korea Volume 1, Number 2 Ascomycota: Dothideomycetes: Pleosporales: Pleosporaceae Alternaria and Allied Genera Seung Hun Yu Chungnam National University Fungal Flora of Korea Volume 1, Number 2 Ascomycota: Dothideomycetes: Pleosporales: Pleosporaceae Alternaria and Allied Genera Copyright ⓒ 2015 by the National Institute of Biological Resources Published by the National Institute of Biological Resources Environmental Research Complex, Hwangyeong-ro 42, Seo-gu Incheon, 404-708, Republic of Korea www.nibr.go.kr All rights reserved. No part of this book may be reproduced, stored in a retrieval system, or transmitted, in any form or by any means, electronic, mechanical, photocopying, recording, or otherwise, without the prior permission of the National Institute of Biological Resources. ISBN : 9788968111259-96470 Government Publications Registration Number 11-1480592-000905-01 Printed by Junghaengsa, Inc. in Korea on acid-free paper Publisher : Kim, Sang-Bae Author : Seung Hun Yu Project Staff : Youn-Bong Ku, Ga Youn Cho, Eun-Young Lee Published on March 1, 2015 The Flora and Fauna of Korea logo was designed to represent six major target groups of the project including vertebrates, invertebrates, insects, algae, fungi, and bacteria. The book cover and the logo were designed by Jee-Yeon Koo. Preface The biological resources represent all the composition of organisms and genetic resources which possess the practical and potential values essential for human lives, and occupies a firm position in producing highly value-added products such as new breeds, new materials and new drugs as a means of boosting the national competitiveness. -
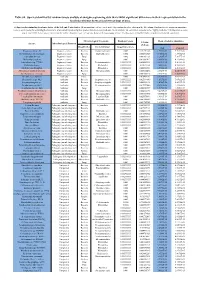
Table S8. Species Identified by Random Forests Analysis of Shotgun Sequencing Data That Exhibit Significant Differences In
Table S8. Species identified by random forests analysis of shotgun sequencing data that exhibit significant differences in their representation in the fecal microbiomes between each two groups of mice. (a) Species discriminating fecal microbiota of the Soil and Control mice. Mean importance of species identified by random forest are shown in the 5th column. Random forests assigns an importance score to each species by estimating the increase in error caused by removing that species from the set of predictors. In our analysis, we considered a species to be “highly predictive” if its importance score was at least 0.001. T-test was performed for the relative abundances of each species between the two groups of mice. P-values were at least 0.05 to be considered statistically significant. Microbiological Taxonomy Random Forests Mean of relative abundance P-Value Species Microbiological Function (T-Test) Classification Bacterial Order Importance Score Soil Control Rhodococcus sp. 2G Engineered strain Bacteria Corynebacteriales 0.002 5.73791E-05 1.9325E-05 9.3737E-06 Herminiimonas arsenitoxidans Engineered strain Bacteria Burkholderiales 0.002 0.005112829 7.1580E-05 1.3995E-05 Aspergillus ibericus Engineered strain Fungi 0.002 0.001061181 9.2368E-05 7.3057E-05 Dichomitus squalens Engineered strain Fungi 0.002 0.018887472 8.0887E-05 4.1254E-05 Acinetobacter sp. TTH0-4 Engineered strain Bacteria Pseudomonadales 0.001333333 0.025523638 2.2311E-05 8.2612E-06 Rhizobium tropici Engineered strain Bacteria Rhizobiales 0.001333333 0.02079554 7.0081E-05 4.2000E-05 Methylocystis bryophila Engineered strain Bacteria Rhizobiales 0.001333333 0.006513543 3.5401E-05 2.2044E-05 Alteromonas naphthalenivorans Engineered strain Bacteria Alteromonadales 0.001 0.000660472 2.0747E-05 4.6463E-05 Saccharomyces cerevisiae Engineered strain Fungi 0.001 0.002980726 3.9901E-05 7.3043E-05 Bacillus phage Belinda Antibiotic Phage 0.002 0.016409765 6.8789E-07 6.0681E-08 Streptomyces sp. -

Characterising Plant Pathogen Communities and Their Environmental Drivers at a National Scale
Lincoln University Digital Thesis Copyright Statement The digital copy of this thesis is protected by the Copyright Act 1994 (New Zealand). This thesis may be consulted by you, provided you comply with the provisions of the Act and the following conditions of use: you will use the copy only for the purposes of research or private study you will recognise the author's right to be identified as the author of the thesis and due acknowledgement will be made to the author where appropriate you will obtain the author's permission before publishing any material from the thesis. Characterising plant pathogen communities and their environmental drivers at a national scale A thesis submitted in partial fulfilment of the requirements for the Degree of Doctor of Philosophy at Lincoln University by Andreas Makiola Lincoln University, New Zealand 2019 General abstract Plant pathogens play a critical role for global food security, conservation of natural ecosystems and future resilience and sustainability of ecosystem services in general. Thus, it is crucial to understand the large-scale processes that shape plant pathogen communities. The recent drop in DNA sequencing costs offers, for the first time, the opportunity to study multiple plant pathogens simultaneously in their naturally occurring environment effectively at large scale. In this thesis, my aims were (1) to employ next-generation sequencing (NGS) based metabarcoding for the detection and identification of plant pathogens at the ecosystem scale in New Zealand, (2) to characterise plant pathogen communities, and (3) to determine the environmental drivers of these communities. First, I investigated the suitability of NGS for the detection, identification and quantification of plant pathogens using rust fungi as a model system. -

Rice Sheath Rot: an Emerging Ubiquitous Destructive Disease Complex
REVIEW published: 11 December 2015 doi: 10.3389/fpls.2015.01066 Rice Sheath Rot: An Emerging Ubiquitous Destructive Disease Complex Vincent de P. Bigirimana1,2, Gia K. H. Hua1, Obedi I. Nyamangyoku2 and Monica Höfte1* 1 Laboratory of Phytopathology, Department of Crop Protection, Faculty of Bioscience Engineering, Ghent University, Ghent, Belgium, 2 Department of Crop Science, School of Agriculture, Rural Development and Agricultural Economics, College of Agriculture, Animal Science and Veterinary Medicine, University of Rwanda, Musanze, Rwanda Around one century ago, a rice disease characterized mainly by rotting of sheaths was reported in Taiwan. The causal agent was identified as Acrocylindrium oryzae, later known as Sarocladium oryzae. Since then it has become clear that various other organisms can cause similar disease symptoms, including Fusarium sp. and fluorescent pseudomonads. These organisms have in common that they produce a range of phytotoxins that induce necrosis in plants. The same agents also cause grain discoloration, chaffiness, and sterility and are all seed-transmitted. Rice sheath rot disease symptoms are found in all rice-growing areas of the world. The disease is Edited by: now getting momentum and is considered as an important emerging rice production Brigitte Mauch-Mani, Université de Neuchâtel, Switzerland threat. The disease can lead to variable yield losses, which can be as high as 85%. This Reviewed by: review aims at improving our understanding of the disease etiology of rice sheath rot Choong-Min Ryu, and mainly deals with the three most reported rice sheath rot pathogens: S. oryzae, Korea Research Institute the Fusarium fujikuroi complex, and Pseudomonas fuscovaginae. Causal agents, of Bioscience and Biotechnology, South Korea pathogenicity determinants, interactions among the various pathogens, epidemiology, Javier Plasencia, geographical distribution, and control options will be discussed. -

Redalyc.Análisis De Pseudomonas Fitopatógenas Usando Métodos
Revista Mexicana de Fitopatología ISSN: 0185-3309 [email protected] Sociedad Mexicana de Fitopatología, A.C. México Slabbinck, Bram; De Baets, Bernard; Dawyndt, Peter; Vos, Paul De Análisis de Pseudomonas Fitopatógenas Usando Métodos Inteligentes de Aprendizaje: Un Enfoque General Sobre Taxonomía y Análisis de Ácidos Grasos Dentro del Género Pseudomonas Revista Mexicana de Fitopatología, vol. 28, núm. 1, 2010, pp. 1-16 Sociedad Mexicana de Fitopatología, A.C. Texcoco, México Disponible en: http://www.redalyc.org/articulo.oa?id=61214206001 Cómo citar el artículo Número completo Sistema de Información Científica Más información del artículo Red de Revistas Científicas de América Latina, el Caribe, España y Portugal Página de la revista en redalyc.org Proyecto académico sin fines de lucro, desarrollado bajo la iniciativa de acceso abierto REVISTA MEXICANA DE FITOPATOLOGÍA/1 Análisis de Pseudomonas Fitopatógenas Usando Métodos Inteligentes de Aprendizaje: Un Enfoque General Sobre Taxonomía y Análisis de Ácidos Grasos Dentro del Género Pseudomonas Analysis of Plant-Pathogenic Pseudomonas Species Using Intelligent Learning Methods: A General Focus on Taxonomy and Fatty Acid Analysis Within the Genus Pseudomonas Bram Slabbinck, Bernard De Baets, Research Group Knowledge-based Systems, Department of Applied Mathematics, Biometrics and Process Control, Ghent University, Coupure links 653, 9000 Ghent, Belgium, Peter Dawyndt, Department of Applied Mathematics and Computer Science, Ghent University, Krijgslaan 281 S9, 9000 Ghent, Belgium y Paul De Vos, Research Group Knowledge-based Systems, Department of Applied Mathematics, Biometrics and Process Control, Ghent University, Coupure links 653, 9000 Ghent, Belgium. (Recibido: Septiembre 20, 2009 Aceptado Enero 12, 2010) Slabbinck, B., De Baets, B., Dawyndt, P., y De Vos, P. -

Applied Mycology / Edited by Mahendra Rai and Paul Dennis Bridge
APPLIED MYCOLOGY This page intentionally left blank APPLIED MYCOLOGY Edited by Mahendra Rai SGB Amravati University, India and Paul Dennis Bridge British Antarctic Survey, UK CABI is a trading name of CAB International CABI Head Office CABI North American Office Nosworthy Way 875 Massachusetts Avenue Wallingford 7th Floor Oxon OX10 8DE Cambridge, MA 02139 UK USA Tel: +44 (0)1491 832111 Tel: +1 617 395 4056 Fax: +44 (0)1491 833508 Fax: +1 617 354 6875 E-mail: [email protected] E-mail: [email protected] Website: www.cabi.org © CAB International 2009. All rights reserved. No part of this publication may be reproduced in any form or by any means, electronically, mechanically, by photocopying, recording or otherwise, without the prior permission of the copyright owners. A catalogue record for this book is available from the British Library, London, UK. Library of Congress Cataloging-in-Publication Data Applied mycology / edited by Mahendra Rai and Paul Dennis Bridge. p. cm. ISBN 978-1-84593-534-4 (alk. paper) 1. Mycology. I. Rai, Mahendra. II. Bridge, P. D. III. Title. QK603.A647 2009 579.5--dc22 2009001597 ISBN-13: 978 1 84593 534 4 Typeset by Columns Design, Reading, UK. Printed and bound in the UK by the MPG Books Group. The paper used for the text pages in this book is FSC certified. The FSC (Forest Stewardship Council) is an international network to promote responsible management of the world’s forests. Contents Contributors vii Preface x Part I: Overview 1 Mycology: a Neglected Megascience 1 David L. Hawksworth Part II: Environment, Agriculture and Forestry 2 Arbuscular Mycorrhizal Fungi and Plant Symbiosis under Stress Conditions: Ecological Implications of Drought, Flooding and Salinity 17 Ileana V. -

Phylogenomic Analyses of the Genus Pseudomonas Lead to the Rearrangement of Several Species and the Definition of New Genera
biology Article Phylogenomic Analyses of the Genus Pseudomonas Lead to the Rearrangement of Several Species and the Definition of New Genera Zaki Saati-Santamaría 1,2,* , Ezequiel Peral-Aranega 1,2 , Encarna Velázquez 1,2,3, Raúl Rivas 1,2,3 and Paula García-Fraile 1,2,3 1 Microbiology and Genetics Department, University of Salamanca, 37007 Salamanca, Spain; [email protected] (E.P.-A.); [email protected] (E.V.); [email protected] (R.R.); [email protected] (P.G.-F.) 2 Institute for Agribiotechnology Research (CIALE), 37185 Salamanca, Spain 3 Associated Research Unit of Plant-Microorganism Interaction, University of Salamanca-IRNASA-CSIC, 37008 Salamanca, Spain * Correspondence: [email protected] Simple Summary: Pseudomonas represents a very important bacterial genus that inhabits many environments and plays either prejudicial or beneficial roles for higher hosts. However, there are many Pseudomonas species which are too divergent to the rest of the genus. This may interfere in the correct development of biological and ecological studies in which Pseudomonas are involved. Thus, we aimed to study the correct taxonomic placement of Pseudomonas species. Based on the study of their genomes and some evolutionary-based methodologies, we suggest the description of three new genera (Denitrificimonas, Parapseudomonas and Neopseudomonas) and many reclassifications of species Citation: Saati-Santamaría, Z.; previously included in Pseudomonas. Peral-Aranega, E.; Velázquez, E.; Rivas, R.; García-Fraile, P. Abstract: Pseudomonas is a large and diverse genus broadly distributed in nature. Its species play Phylogenomic Analyses of the Genus relevant roles in the biology of earth and living beings. Because of its ubiquity, the number of new Pseudomonas Lead to the species is continuously increasing although its taxonomic organization remains quite difficult to Rearrangement of Several Species and unravel.