A Structurally Validated Sequence Alignment of All 497 Typical Human Protein Kinase Domains
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Symbol Gene Description ACVR1B Activin a Receptor, Type IB
Table S1. Kinase clones included in human kinase cDNA library for yeast two-hybrid screening Gene Symbol Gene Description ACVR1B activin A receptor, type IB ADCK2 aarF domain containing kinase 2 ADCK4 aarF domain containing kinase 4 AGK multiple substrate lipid kinase;MULK AK1 adenylate kinase 1 AK3 adenylate kinase 3 like 1 AK3L1 adenylate kinase 3 ALDH18A1 aldehyde dehydrogenase 18 family, member A1;ALDH18A1 ALK anaplastic lymphoma kinase (Ki-1) ALPK1 alpha-kinase 1 ALPK2 alpha-kinase 2 AMHR2 anti-Mullerian hormone receptor, type II ARAF v-raf murine sarcoma 3611 viral oncogene homolog 1 ARSG arylsulfatase G;ARSG AURKB aurora kinase B AURKC aurora kinase C BCKDK branched chain alpha-ketoacid dehydrogenase kinase BMPR1A bone morphogenetic protein receptor, type IA BMPR2 bone morphogenetic protein receptor, type II (serine/threonine kinase) BRAF v-raf murine sarcoma viral oncogene homolog B1 BRD3 bromodomain containing 3 BRD4 bromodomain containing 4 BTK Bruton agammaglobulinemia tyrosine kinase BUB1 BUB1 budding uninhibited by benzimidazoles 1 homolog (yeast) BUB1B BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) C9orf98 chromosome 9 open reading frame 98;C9orf98 CABC1 chaperone, ABC1 activity of bc1 complex like (S. pombe) CALM1 calmodulin 1 (phosphorylase kinase, delta) CALM2 calmodulin 2 (phosphorylase kinase, delta) CALM3 calmodulin 3 (phosphorylase kinase, delta) CAMK1 calcium/calmodulin-dependent protein kinase I CAMK2A calcium/calmodulin-dependent protein kinase (CaM kinase) II alpha CAMK2B calcium/calmodulin-dependent -

The Prevalence of MADH4 and BMPR1A Mutations in Juvenile Polyposis and Absence of BMPR2, BMPR1B, and ACVR1 Mutations
484 ORIGINAL ARTICLE J Med Genet: first published as 10.1136/jmg.2004.018598 on 2 July 2004. Downloaded from The prevalence of MADH4 and BMPR1A mutations in juvenile polyposis and absence of BMPR2, BMPR1B, and ACVR1 mutations J R Howe, M G Sayed, A F Ahmed, J Ringold, J Larsen-Haidle, A Merg, F A Mitros, C A Vaccaro, G M Petersen, F M Giardiello, S T Tinley, L A Aaltonen, H T Lynch ............................................................................................................................... J Med Genet 2004;41:484–491. doi: 10.1136/jmg.2004.018598 Background: Juvenile polyposis (JP) is an autosomal dominant syndrome predisposing to colorectal and gastric cancer. We have identified mutations in two genes causing JP, MADH4 and bone morphogenetic protein receptor 1A (BMPR1A): both are involved in bone morphogenetic protein (BMP) mediated signalling and are members of the TGF-b superfamily. This study determined the prevalence of mutations See end of article for in MADH4 and BMPR1A, as well as three other BMP/activin pathway candidate genes in a large number authors’ affiliations ....................... of JP patients. Methods: DNA was extracted from the blood of JP patients and used for PCR amplification of each exon of Correspondence to: these five genes, using primers flanking each intron–exon boundary. Mutations were determined by Dr J R Howe, Department of Surgery, 4644 JCP, comparison to wild type sequences using sequence analysis software. A total of 77 JP cases were University of Iowa College sequenced for mutations in the MADH4, BMPR1A, BMPR1B, BMPR2, and/or ACVR1 (activin A receptor) of Medicine, 200 Hawkins genes. The latter three genes were analysed when MADH4 and BMPR1A sequencing found no mutations. -

Casein Kinase 1 Isoforms in Degenerative Disorders
CASEIN KINASE 1 ISOFORMS IN DEGENERATIVE DISORDERS DISSERTATION Presented in Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy in the Graduate School of The Ohio State University By Theresa Joseph Kannanayakal, M.Sc., M.S. * * * * * The Ohio State University 2004 Dissertation Committee: Approved by Professor Jeff A. Kuret, Adviser Professor John D. Oberdick Professor Dale D. Vandre Adviser Professor Mike X. Zhu Biophysics Graduate Program ABSTRACT Casein Kinase 1 (CK1) enzyme is one of the largest family of Serine/Threonine protein kinases. CK1 has a wide distribution spanning many eukaryotic families. In cells, its kinase activity has been found in various sub-cellular compartments enabling it to phosphorylate many proteins involved in cellular maintenance and disease pathogenesis. Tau is one such substrate whose hyperphosphorylation results in degeneration of neurons in Alzheimer’s disease (AD). AD is a slow neuroprogessive disorder histopathologically characterized by Granulovacuolar degeneration bodies (GVBs) and intraneuronal accumulation of tau in Neurofibrillary Tangles (NFTs). The level of CK1 isoforms, CK1α, CK1δ and CK1ε has been shown to be elevated in AD. Previous studies of the correlation of CK1δ with lesions had demonstrated its importance in tau hyperphosphorylation. Hence we investigated distribution of CK1α and CK1ε with the lesions to understand if they would play role in tau hyperphosphorylation similar to CK1δ. The kinase results were also compared with lesion correlation studies of peptidyl cis/trans prolyl isomerase (Pin1) and caspase-3. Our results showed that among the enzymes investigated, CK1 isoforms have the greatest extent of colocalization with the lesions. We have also investigated the distribution of CK1α with different stages of NFTs that follow AD progression. -

Map2k1 and Map2k2 Genes Contribute to the Normal Development of Syncytiotrophoblasts During Placentation
RESEARCH ARTICLE 1363 Development 136, 1363-1374 (2009) doi:10.1242/dev.031872 Map2k1 and Map2k2 genes contribute to the normal development of syncytiotrophoblasts during placentation Valérie Nadeau*, Stéphanie Guillemette*, Louis-François Bélanger, Olivier Jacob, Sophie Roy and Jean Charron† The mammalian genome contains two ERK/MAP kinase kinase genes, Map2k1 and Map2k2, which encode dual-specificity kinases responsible for ERK/MAP kinase activation. In the mouse, loss of Map2k1 function causes embryonic lethality, whereas Map2k2 mutants survive with a normal lifespan, suggesting that Map2k1 masks the phenotype due to the Map2k2 mutation. To uncover the specific function of MAP2K2 and the threshold requirement of MAP2K proteins during embryo formation, we have successively ablated the Map2k gene functions. We report here that Map2k2 haploinsufficiency affects the normal development of placenta in the absence of one Map2k1 allele. Most Map2k1+/–Map2k2+/– embryos die during gestation because of placenta defects restricted to extra-embryonic tissues. The impaired viability of Map2k1+/–Map2k2+/– embryos can be rescued when the Map2k1 deletion is restricted to the embryonic tissues. The severity of the placenta phenotype is dependent on the number of Map2k mutant alleles, the deletion of the Map2k1 allele being more deleterious. Moreover, the deletion of one or both Map2k2 alleles in the context of one null Map2k1 allele leads to the formation of multinucleated trophoblast giant (MTG) cells. Genetic experiments indicate that these structures are derived from Gcm1-expressing syncytiotrophoblasts (SynT), which are affected in their ability to form the uniform SynT layer II lining the maternal sinuses. Thus, even though Map2k1 plays a predominant role, these results enlighten the function of Map2k2 in placenta development. -

The Genetic Landscape of Clinical Resistance to RAF Inhibition in Metastatic Melanoma
CD-13-0617_PAP.indd Page OF1 19/11/13 9:36 PM user-f028 /Books-Arts/JOURNAL-Cancer%20Discovery/01-JAN-Issue-2014/PAP Published OnlineFirst November 21, 2013; DOI: 10.1158/2159-8290.CD-13-0617 RESEARCH ARTICLE The Genetic Landscape of Clinical Resistance to RAF Inhibition in Metastatic Melanoma Eliezer M. Van Allen 1 , 3 , Nikhil Wagle 1 , 3 , Antje Sucker 5 , 6 , Daniel J. Treacy 1 , Cory M. Johannessen 3 , Eva M. Goetz 1 , Chelsea S. Place 1 , 3 , Amaro Taylor-Weiner 3 , Steven Whittaker 3 , Gregory V. Kryukov 3 , Eran Hodis 1 , 3,4 , Mara Rosenberg 3 , Aaron McKenna 3 , 15 , Kristian Cibulskis 3 , Deborah Farlow 3 , Lisa Zimmer 5 , 6 , Uwe Hillen 5 , 6 , Ralf Gutzmer 8 , Simone M. Goldinger 16 , Selma Ugurel 9 , Helen J. Gogas 17 , Friederike Egberts 10 , Carola Berking 6 , 11 , Uwe Trefzer 6 , 12 , Carmen Loquai 6 , 13 , Benjamin Weide 6 , 14 , Jessica C. Hassel 6 , 7 , Stacey B. Gabriel 3 , Scott L. Carter 3 , Gad Getz 2 , 3 , Levi A. Garraway 1 , 3 , and Dirk Schadendorf 5 , 6 on behalf of the Dermatologic Cooperative Oncology Group of Germany (DeCOG) Downloaded from cancerdiscovery.aacrjournals.org on September 25, 2021. © 2013 American Association for Cancer Research. CD-13-0617_PAP.indd Page OF2 19/11/13 9:36 PM user-f028 /Books-Arts/JOURNAL-Cancer%20Discovery/01-JAN-Issue-2014/PAP Published OnlineFirst November 21, 2013; DOI: 10.1158/2159-8290.CD-13-0617 ABSTRACT Most patients with BRAF V600 -mutant metastatic melanoma develop resistance to selective RAF kinase inhibitors. The spectrum of clinical genetic resistance mechanisms to RAF inhibitors and options for salvage therapy are incompletely understood. -

RAF Protein-Serine/Threonine Kinases: Structure and Regulation
Biochemical and Biophysical Research Communications 399 (2010) 313–317 Contents lists available at ScienceDirect Biochemical and Biophysical Research Communications journal homepage: www.elsevier.com/locate/ybbrc Mini Review RAF protein-serine/threonine kinases: Structure and regulation Robert Roskoski Jr. * Blue Ridge Institute for Medical Research, 3754 Brevard Road, Suite 116, Box 19, Horse Shoe, NC 28742, USA article info abstract Article history: A-RAF, B-RAF, and C-RAF are a family of three protein-serine/threonine kinases that participate in the Received 12 July 2010 RAS-RAF-MEK-ERK signal transduction cascade. This cascade participates in the regulation of a large vari- Available online 30 July 2010 ety of processes including apoptosis, cell cycle progression, differentiation, proliferation, and transforma- tion to the cancerous state. RAS mutations occur in 15–30% of all human cancers, and B-RAF mutations Keywords: occur in 30–60% of melanomas, 30–50% of thyroid cancers, and 5–20% of colorectal cancers. Activation 14-3-3 of the RAF kinases requires their interaction with RAS-GTP along with dephosphorylation and also phos- ERK phorylation by SRC family protein-tyrosine kinases and other protein-serine/threonine kinases. The for- GDC-0879 mation of unique side-to-side RAF dimers is required for full kinase activity. RAF kinase inhibitors are MEK Melanoma effective in blocking MEK1/2 and ERK1/2 activation in cells containing the oncogenic B-RAF Val600Glu PLX4032 activating mutation. RAF kinase inhibitors lead to the paradoxical increase in RAF kinase activity in cells PLX4720 containing wild-type B-RAF and wild-type or activated mutant RAS. -

Genome-Wide DNA Methylation Analysis of KRAS Mutant Cell Lines Ben Yi Tew1,5, Joel K
www.nature.com/scientificreports OPEN Genome-wide DNA methylation analysis of KRAS mutant cell lines Ben Yi Tew1,5, Joel K. Durand2,5, Kirsten L. Bryant2, Tikvah K. Hayes2, Sen Peng3, Nhan L. Tran4, Gerald C. Gooden1, David N. Buckley1, Channing J. Der2, Albert S. Baldwin2 ✉ & Bodour Salhia1 ✉ Oncogenic RAS mutations are associated with DNA methylation changes that alter gene expression to drive cancer. Recent studies suggest that DNA methylation changes may be stochastic in nature, while other groups propose distinct signaling pathways responsible for aberrant methylation. Better understanding of DNA methylation events associated with oncogenic KRAS expression could enhance therapeutic approaches. Here we analyzed the basal CpG methylation of 11 KRAS-mutant and dependent pancreatic cancer cell lines and observed strikingly similar methylation patterns. KRAS knockdown resulted in unique methylation changes with limited overlap between each cell line. In KRAS-mutant Pa16C pancreatic cancer cells, while KRAS knockdown resulted in over 8,000 diferentially methylated (DM) CpGs, treatment with the ERK1/2-selective inhibitor SCH772984 showed less than 40 DM CpGs, suggesting that ERK is not a broadly active driver of KRAS-associated DNA methylation. KRAS G12V overexpression in an isogenic lung model reveals >50,600 DM CpGs compared to non-transformed controls. In lung and pancreatic cells, gene ontology analyses of DM promoters show an enrichment for genes involved in diferentiation and development. Taken all together, KRAS-mediated DNA methylation are stochastic and independent of canonical downstream efector signaling. These epigenetically altered genes associated with KRAS expression could represent potential therapeutic targets in KRAS-driven cancer. Activating KRAS mutations can be found in nearly 25 percent of all cancers1. -

And Rasopathies-Associated SHP2 and BRAF Mutations
Functional Characterization of Cancer- and RASopathies-associated SHP2 and BRAF Mutations Dissertation zur Erlangung des akademischen Grades doctor rerum naturalium (Dr. rer. nat.) im Fach Biologie/ Molekularbiologie eingereicht an der Lebenswissenschaftlichen Fakult¨at der Humboldt–Universit¨at zu Berlin von M. Sc. Paula Andrea Medina-P´erez Pr¨asident der Humboldt–Universit¨at zu Berlin Prof. Dr. Jan–Hendrik Olbertz Dekan der Lebenswissenschaftlichen Fakult¨at Prof. Dr. Richard Lucius Gutachter: 1. Prof. Dr. Reinhold Sch¨afer 2. Prof. Dr. Hanspeter Herzel 3. Prof. Dr. Holger S¨ultmann Tag der m¨undlichen Pr¨ufung: 12.03.2015 Abstract Deregulation of the Ras/MAPK signaling is implicated in a wide variety of human diseases, including developmental disorders and cancer. In the last years, a group of developmental disorders, characterized by an overlapping phenotype in patients, was clustered under the term RASopathies. These disorders result from germline mutations in genes encoding key components of the Ras/MAPK signaling cascade. Although the incidence of solid tumors in patients suffering from these disorders is rather low, reports on different forms of leukemia have considerably increased. In this work, a group of mutations in the genes SHP2/PTPN11 and BRAF, both key regulators of the MAPK signaling pathway and implicated in RASopathies and cancer, were selected for expression in well-established cell systems for a comprehensive molecular and phenotypic characterization using high-throughput approaches and functional assays. Synthetic cDNA sequences carrying the SHP2 mutations T42A, E76D, I282V (Noonan syndrome-associated), E76G, E76K, E139D (Noonan- and leukemia-associated), T468M (LEOPARD syndrome -associated) and the BRAF mutations Q257R, S467A, L485F and K499E (cardio-facio-cutaneous syndrome-associated) were shuttled into the modified lentivi- ral vector pCDH-EF1-IRES-GFP. -

Analysis of Protein Kinase Domain and Tyrosine Kinase Or Serine/Threonine Kinase Signatures Involved in Lung Cancer
Analysis of Protein Kinase Domain and Tyrosine Kinase Or serine/Threonine Kinase Signatures Involved In Lung Cancer Lung cancer results when normal check and balance system of cell division is disrupted and ultimately the cells divide and proliferate in an uncontrollable manner forming a mass of cells in our body, known as tumor. Frequent mutations in Protein Kinase Domain alter the process of phosphorylation which results in abnormality in regulations of cell apoptosis and differentiation. Tyrosine Protein kinases and Serine/Threonine Protein Kinases are the two s t broad classes of protein kinases in accordance to their substrate specificity. The study of n i r Tyrosine protein kinase and serine Kinase coding regions have the importance of sequence P and structure determinants of cancer-causing mutations from mutation-dependent activation e r P process. In the present study, we analyzed huge amounts of data extracted from various biological databases and NCBI. Out of the 534 proteins that may play a role in lung cancer, 71 proteins were selected that are likely to be actively involved in lung cancer. These proteins were evaluated by employing Multiple Sequence Alignment and a Phylogenetic tree was constructed using Neighbor-Joining Algorithm. From the constructed phylogenetic tree, protein kinase domain and motif study was performed. The results of this study revealed that the presence of Protein Kinase Domain and Tyrosine or Serine/Threonine Kinase signatures in some of the proteins are mutated, which play a dominant role in the pathogenesis of Lung Cancer and these may be addressed with the help of inhibitors to develop an efficient anticancer drugs. -
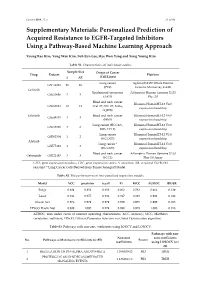
Personalized Prediction of Acquired Resistance to EGFR-Targeted Inhibitors Using a Pathway-Based Machine Learning Approach
Cancers 2019, 11, x S1 of S9 Supplementary Materials: Personalized Prediction of Acquired Resistance to EGFR-Targeted Inhibitors Using a Pathway-Based Machine Learning Approach Young Rae Kim, Yong Wan Kim, Suh Eun Lee, Hye Won Yang and Sung Young Kim Table S1. Characteristics of individual studies. Sample Size Origin of Cancer Drug Dataset Platform S AR (Cell Lines) Lung cancer Agilent-014850 Whole Human GSE34228 26 26 (PC9) Genome Microarray 4x44K Gefitinib Epidermoid carcinoma Affymetrix Human Genome U133 GSE10696 3 3 (A431) Plus 2.0 Head and neck cancer Illumina HumanHT-12 V4.0 GSE62061 12 12 (Cal-27, SSC-25, FaDu, expression beadchip SQ20B) Erlotinib Head and neck cancer Illumina HumanHT-12 V4.0 GSE49135 3 3 (HN5) expression beadchip Lung cancer (HCC827, Illumina HumanHT-12 V3.0 GSE38310 3 6 ER3, T15-2) expression beadchip Lung cancer Illumina HumanHT-12 V3.0 GSE62504 1 2 (HCC827) expression beadchip Afatinib Lung cancer * Illumina HumanHT-12 V4.0 GSE75468 1 3 (HCC827) expression beadchip Head and neck cancer Affymetrix Human Genome U133 Cetuximab GSE21483 3 3 (SCC1) Plus 2.0 Array GEO, gene expression omnibus; GSE, gene expression series; S, sensitive; AR, acquired EGFR-TKI resistant; * Lung Cancer Cells Derived from Tumor Xenograft Model. Table S2. The performances of four penalized regression models. Model ACC precision recall F1 MCC AUROC BRIER Ridge 0.889 0.852 0.958 0.902 0.782 0.964 0.129 Lasso 0.944 0.957 0.938 0.947 0.889 0.991 0.042 Elastic Net 0.978 0.979 0.979 0.979 0.955 0.999 0.023 EPSGO Elastic Net 0.989 1.000 0.979 0.989 0.978 1.000 0.018 AUROC, area under curve of receiver operating characteristic; ACC, accuracy; MCC, Matthews correlation coefficient; EPSGO, Efficient Parameter Selection via Global Optimization algorithm. -

Two Novel Disease-Causing Variants in BMPR1B Are Associated with Brachydactyly Type A1
European Journal of Human Genetics (2015) 23, 1640–1645 & 2015 Macmillan Publishers Limited All rights reserved 1018-4813/15 www.nature.com/ejhg ARTICLE Two novel disease-causing variants in BMPR1B are associated with brachydactyly type A1 Lemuel Racacho1,2, Ashley M Byrnes1,3, Heather MacDonald3, Helen J Dranse4, Sarah M Nikkel5,6, Judith Allanson6, Elisabeth Rosser7, T Michael Underhill4 and Dennis E Bulman*,1,2,5 Brachydactyly type A1 is an autosomal dominant disorder primarily characterized by hypoplasia/aplasia of the middle phalanges of digits 2–5. Human and mouse genetic perturbations in the BMP-SMAD signaling pathway have been associated with many brachymesophalangies, including BDA1, as causative mutations in IHH and GDF5 have been previously identified. GDF5 interacts directly as the preferred ligand for the BMP type-1 receptor BMPR1B and is important for both chondrogenesis and digit formation. We report pathogenic variants in BMPR1B that are associated with complex BDA1. A c.975A4C (p. (Lys325Asn)) was identified in the first patient displaying absent middle phalanges and shortened distal phalanges of the toes in addition to the significant shortening of middle phalanges in digits 2, 3 and 5 of the hands. The second patient displayed a combination of brachydactyly and arachnodactyly. The sequencing of BMPR1B in this individual revealed a novel c.447-1G4A at a canonical acceptor splice site of exon 8, which is predicted to create a novel acceptor site, thus leading to a translational reading frameshift. Both mutations are most likely to act in a dominant-negative manner, similar to the effects observed in BMPR1B mutations that cause BDA2. -

Characterization of the Small Molecule Kinase Inhibitor SU11248 (Sunitinib/ SUTENT in Vitro and in Vivo
TECHNISCHE UNIVERSITÄT MÜNCHEN Lehrstuhl für Genetik Characterization of the Small Molecule Kinase Inhibitor SU11248 (Sunitinib/ SUTENT in vitro and in vivo - Towards Response Prediction in Cancer Therapy with Kinase Inhibitors Michaela Bairlein Vollständiger Abdruck der von der Fakultät Wissenschaftszentrum Weihenstephan für Ernährung, Landnutzung und Umwelt der Technischen Universität München zur Erlangung des akademischen Grades eines Doktors der Naturwissenschaften genehmigten Dissertation. Vorsitzender: Univ. -Prof. Dr. K. Schneitz Prüfer der Dissertation: 1. Univ.-Prof. Dr. A. Gierl 2. Hon.-Prof. Dr. h.c. A. Ullrich (Eberhard-Karls-Universität Tübingen) 3. Univ.-Prof. A. Schnieke, Ph.D. Die Dissertation wurde am 07.01.2010 bei der Technischen Universität München eingereicht und durch die Fakultät Wissenschaftszentrum Weihenstephan für Ernährung, Landnutzung und Umwelt am 19.04.2010 angenommen. FOR MY PARENTS 1 Contents 2 Summary ................................................................................................................................................................... 5 3 Zusammenfassung .................................................................................................................................................... 6 4 Introduction .............................................................................................................................................................. 8 4.1 Cancer ..............................................................................................................................................................