Map2k1 and Map2k2 Genes Contribute to the Normal Development of Syncytiotrophoblasts During Placentation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Symbol Gene Description ACVR1B Activin a Receptor, Type IB
Table S1. Kinase clones included in human kinase cDNA library for yeast two-hybrid screening Gene Symbol Gene Description ACVR1B activin A receptor, type IB ADCK2 aarF domain containing kinase 2 ADCK4 aarF domain containing kinase 4 AGK multiple substrate lipid kinase;MULK AK1 adenylate kinase 1 AK3 adenylate kinase 3 like 1 AK3L1 adenylate kinase 3 ALDH18A1 aldehyde dehydrogenase 18 family, member A1;ALDH18A1 ALK anaplastic lymphoma kinase (Ki-1) ALPK1 alpha-kinase 1 ALPK2 alpha-kinase 2 AMHR2 anti-Mullerian hormone receptor, type II ARAF v-raf murine sarcoma 3611 viral oncogene homolog 1 ARSG arylsulfatase G;ARSG AURKB aurora kinase B AURKC aurora kinase C BCKDK branched chain alpha-ketoacid dehydrogenase kinase BMPR1A bone morphogenetic protein receptor, type IA BMPR2 bone morphogenetic protein receptor, type II (serine/threonine kinase) BRAF v-raf murine sarcoma viral oncogene homolog B1 BRD3 bromodomain containing 3 BRD4 bromodomain containing 4 BTK Bruton agammaglobulinemia tyrosine kinase BUB1 BUB1 budding uninhibited by benzimidazoles 1 homolog (yeast) BUB1B BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) C9orf98 chromosome 9 open reading frame 98;C9orf98 CABC1 chaperone, ABC1 activity of bc1 complex like (S. pombe) CALM1 calmodulin 1 (phosphorylase kinase, delta) CALM2 calmodulin 2 (phosphorylase kinase, delta) CALM3 calmodulin 3 (phosphorylase kinase, delta) CAMK1 calcium/calmodulin-dependent protein kinase I CAMK2A calcium/calmodulin-dependent protein kinase (CaM kinase) II alpha CAMK2B calcium/calmodulin-dependent -

Recombinant MAP2K2 Protein
Recombinant MAP2K2 protein Catalog No: 81332, 81632 Quantity: 20, 1000 µg Expressed In: Baculovirus Concentration: 0.3 µg/µl Source: Human Buffer Contents: Recombinant MAP2K2 protein is supplied in 25 mM HEPES-NaOH pH 7.5, 300 mM NaCl, 10% glycerol, 0.04% Triton X-100, and 0.5 mM TCEP. Background: MAP2K2 (Mitogen-Activated Protein Kinase Kinase Kinase 2) is a dual specificity protein kinase that belongs to the MAP kinase kinase family. This kinase plays a critical role in mitogen growth factor signal transduction. It phosphorylates and activates MAPK1/ERK2 and MAPK2/ERK3. The activation of this kinase itself is dependent on the Ser/Thr phosphorylation by MAP kinase kinase kinases. Mutations in this gene cause cardiofaciocutaneous syndrome (CFC syndrome), a disease characterized by heart defects, cognitive disability, and distinctive facial features similar to those found in Noonan syndrome. Protein Details: Recombinant MAP2K2 protein was expressed in baculovirus expression system as the full length protein (accession number NP_109587.1) with a N-terminal FLAG Tag. The molecular weight of the protein is 45.7 kDa. Application Notes: This product was manufactured as described in Protein Details. Where possible, Active Motif has developed functional or activity assays for Recombinant MAP2K2 protein gel recombinant proteins. Additional characterization such as enzyme kinetic activity 10% SDS-PAGE with Coomassie blue assays, inhibitor screening or other biological activity assays may not have been staining MW: 45.7 kDa performed for every product. All available data for a given product is shown on the lot- Purity: >90% specific Technical Data Sheet. Storage and Guarantee: Recombinant proteins in solution are temperature sensitive and must be stored at -80°C to prevent degradation. -

The Genetic Landscape of Clinical Resistance to RAF Inhibition in Metastatic Melanoma
CD-13-0617_PAP.indd Page OF1 19/11/13 9:36 PM user-f028 /Books-Arts/JOURNAL-Cancer%20Discovery/01-JAN-Issue-2014/PAP Published OnlineFirst November 21, 2013; DOI: 10.1158/2159-8290.CD-13-0617 RESEARCH ARTICLE The Genetic Landscape of Clinical Resistance to RAF Inhibition in Metastatic Melanoma Eliezer M. Van Allen 1 , 3 , Nikhil Wagle 1 , 3 , Antje Sucker 5 , 6 , Daniel J. Treacy 1 , Cory M. Johannessen 3 , Eva M. Goetz 1 , Chelsea S. Place 1 , 3 , Amaro Taylor-Weiner 3 , Steven Whittaker 3 , Gregory V. Kryukov 3 , Eran Hodis 1 , 3,4 , Mara Rosenberg 3 , Aaron McKenna 3 , 15 , Kristian Cibulskis 3 , Deborah Farlow 3 , Lisa Zimmer 5 , 6 , Uwe Hillen 5 , 6 , Ralf Gutzmer 8 , Simone M. Goldinger 16 , Selma Ugurel 9 , Helen J. Gogas 17 , Friederike Egberts 10 , Carola Berking 6 , 11 , Uwe Trefzer 6 , 12 , Carmen Loquai 6 , 13 , Benjamin Weide 6 , 14 , Jessica C. Hassel 6 , 7 , Stacey B. Gabriel 3 , Scott L. Carter 3 , Gad Getz 2 , 3 , Levi A. Garraway 1 , 3 , and Dirk Schadendorf 5 , 6 on behalf of the Dermatologic Cooperative Oncology Group of Germany (DeCOG) Downloaded from cancerdiscovery.aacrjournals.org on September 25, 2021. © 2013 American Association for Cancer Research. CD-13-0617_PAP.indd Page OF2 19/11/13 9:36 PM user-f028 /Books-Arts/JOURNAL-Cancer%20Discovery/01-JAN-Issue-2014/PAP Published OnlineFirst November 21, 2013; DOI: 10.1158/2159-8290.CD-13-0617 ABSTRACT Most patients with BRAF V600 -mutant metastatic melanoma develop resistance to selective RAF kinase inhibitors. The spectrum of clinical genetic resistance mechanisms to RAF inhibitors and options for salvage therapy are incompletely understood. -
Type of the Paper (Article
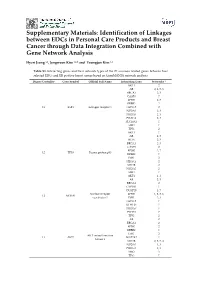
Supplementary Materials: Identification of Linkages between EDCs in Personal Care Products and Breast Cancer through Data Integration Combined with Gene Network Analysis Hyeri Jeong 1,2, Jongwoon Kim 1,2,* and Youngjun Kim 1,2 Table S1. Interacting genes and their network types of the 27 common related genes between four selected EDCs and ER positive breast cancer based on GeneMANIA network analysis. Degree Centrality Gene Symbol Official Full Name Interacting Gene Networks * AKT1 2 AR 2, 3, 5, 6 BRCA1 2, 3 CASP8 7 EP300 2, 3 ERBB2 2 13 ESR1 Estrogen receptor 1 HDAC5 2 NCOA1 2, 3 NCOA7 2, 3 PIK3CA 2, 3 SLC10A1 1 SMO 1 TP53 2 AKT1 1 AR 2, 7 BCL6 2, 3 BRCA1 2, 3 CASP8 2 EP300 2, 7 12 TP53 Tumor protein p53 ERBB2 1 ESR1 2 HDAC5 2 MTOR 2 NCOA1 2 SMO 1 AKT1 1, 3 AR 2, 3 BRCA1 2 CYP1A1 1 DUSP10 1, 7 Nuclear receptor EP300 1, 2, 3, 6 12 NCOA1 coactivator 1 ESR1 2, 3 HDAC5 7 KLHL24 1 NCOA7 7 PTCH1 7 TP53 2 AR 2 BRCA1 2 EP300 2 ERBB2 1 ESR1 2 AKT serine/threonine 11 AKT1 MAP2K2 1 kinase 1 MTOR 2, 3, 5, 4 NCOA1 1, 3 PIK3CA 2, 3 SMO 3 TP53 1 Int. J. Environ. Res. Public Health 2017, 14 S2 of S7 Table S1. Cont. Degree Centrality Gene Symbol Official Full Name Interacting Gene Networks * ABCG1 7 APOB 1, 7 EP300 2, 7 GABRR1 7 HDAC5 2, 3 11 BCL6 B-cell CLL/lymphoma 6 KLHL24 1 PIK3CA 1 PTCH1 7 SLC10A1 1 SMO 2 TP53 2, 3 AKT1 2 BRCA1 2, 3 CASP8 1, 3, 4 EP300 2 ESR1 2, 3, 5, 6 10 AR Androgen receptor NCOA1 2, 3 PIK3CA 3 SLC10A1 1 SMO 1 TP53 2, 4 AKT1 2 AR 2 BCL6 2, 7 BRCA1 2, 7 E1A binding protein 9 EP300 ESR1 2, 3 p300 KLHL24 1 NCOA1 1, 2, 3, 6 PIK3CA 1 TP53 2, 7 ABCG1 1 AKT1 2, 3 AR 3 Phosphatidylinositol- BCL6 1 4,5-bisphosphate 9 PIK3CA EP300 1 3-kinase catalytic ERBB2 3 subunit alpha ESR1 2, 3 KLHL24 1 MTOR 3, 6 AR 1 AKT1 3 BCL6 2 Smoothened, frizzled CYP1A1 1 8 SMO class receptor ESR1 1 ERBB2 1 PTCH1 2, 3 TP53 1 AKT1 2 AR 2, 3 EP300 2, 7 BRCA1, DNA 7 BRCA1 ESR1 2, 3 repair associated MTOR 2 NCOA1 2 TP53 2, 3 AR 2, 3, 4 CYP1A1 1 ESR1 7 7 CASP8 Caspase 8 GABRA6 7 KLHL24 7 MTOR 7 TP53 2 Int. -
Personalized Prediction of Acquired Resistance to EGFR-Targeted Inhibitors Using a Pathway-Based Machine Learning Approach
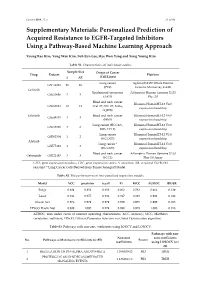
Cancers 2019, 11, x S1 of S9 Supplementary Materials: Personalized Prediction of Acquired Resistance to EGFR-Targeted Inhibitors Using a Pathway-Based Machine Learning Approach Young Rae Kim, Yong Wan Kim, Suh Eun Lee, Hye Won Yang and Sung Young Kim Table S1. Characteristics of individual studies. Sample Size Origin of Cancer Drug Dataset Platform S AR (Cell Lines) Lung cancer Agilent-014850 Whole Human GSE34228 26 26 (PC9) Genome Microarray 4x44K Gefitinib Epidermoid carcinoma Affymetrix Human Genome U133 GSE10696 3 3 (A431) Plus 2.0 Head and neck cancer Illumina HumanHT-12 V4.0 GSE62061 12 12 (Cal-27, SSC-25, FaDu, expression beadchip SQ20B) Erlotinib Head and neck cancer Illumina HumanHT-12 V4.0 GSE49135 3 3 (HN5) expression beadchip Lung cancer (HCC827, Illumina HumanHT-12 V3.0 GSE38310 3 6 ER3, T15-2) expression beadchip Lung cancer Illumina HumanHT-12 V3.0 GSE62504 1 2 (HCC827) expression beadchip Afatinib Lung cancer * Illumina HumanHT-12 V4.0 GSE75468 1 3 (HCC827) expression beadchip Head and neck cancer Affymetrix Human Genome U133 Cetuximab GSE21483 3 3 (SCC1) Plus 2.0 Array GEO, gene expression omnibus; GSE, gene expression series; S, sensitive; AR, acquired EGFR-TKI resistant; * Lung Cancer Cells Derived from Tumor Xenograft Model. Table S2. The performances of four penalized regression models. Model ACC precision recall F1 MCC AUROC BRIER Ridge 0.889 0.852 0.958 0.902 0.782 0.964 0.129 Lasso 0.944 0.957 0.938 0.947 0.889 0.991 0.042 Elastic Net 0.978 0.979 0.979 0.979 0.955 0.999 0.023 EPSGO Elastic Net 0.989 1.000 0.979 0.989 0.978 1.000 0.018 AUROC, area under curve of receiver operating characteristic; ACC, accuracy; MCC, Matthews correlation coefficient; EPSGO, Efficient Parameter Selection via Global Optimization algorithm. -

Phosphorylation Sites in Protein Kinases and Phosphatases Regulated by Formyl Peptide Receptor 2 Signaling
International Journal of Molecular Sciences Review Phosphorylation Sites in Protein Kinases and Phosphatases Regulated by Formyl Peptide Receptor 2 Signaling Maria Carmela Annunziata 1, Melania Parisi 1, Gabriella Esposito 2 , Gabriella Fabbrocini 1, Rosario Ammendola 2 and Fabio Cattaneo 2,* 1 Department of Clinical Medicine and Surgery, School of Medicine, University of Naples Federico II, Via S. Pansini 5, 80131 Naples, Italy; [email protected] (M.C.A.); [email protected] (M.P.); [email protected] (G.F.) 2 Department of Molecular Medicine and Medical Biotechnology, School of Medicine,, University of Naples Federico II, Via S. Pansini 5, 80131 Naples, Italy; [email protected] (G.E.); [email protected] (R.A.) * Correspondence: [email protected]; Fax: +39-081-7464-359 Received: 5 May 2020; Accepted: 25 May 2020; Published: 27 May 2020 Abstract: FPR1, FPR2, and FPR3 are members of Formyl Peptides Receptors (FPRs) family belonging to the GPCR superfamily. FPR2 is a low affinity receptor for formyl peptides and it is considered the most promiscuous member of this family. Intracellular signaling cascades triggered by FPRs include the activation of different protein kinases and phosphatase, as well as tyrosine kinase receptors transactivation. Protein kinases and phosphatases act coordinately and any impairment of their activation or regulation represents one of the most common causes of several human diseases. Several phospho-sites has been identified in protein kinases and phosphatases, whose role may be to expand the repertoire of molecular mechanisms of regulation or may be necessary for fine-tuning of switch properties. We previously performed a phospho-proteomic analysis in FPR2-stimulated cells that revealed, among other things, not yet identified phospho-sites on six protein kinases and one protein phosphatase. -

Characterization of the Small Molecule Kinase Inhibitor SU11248 (Sunitinib/ SUTENT in Vitro and in Vivo
TECHNISCHE UNIVERSITÄT MÜNCHEN Lehrstuhl für Genetik Characterization of the Small Molecule Kinase Inhibitor SU11248 (Sunitinib/ SUTENT in vitro and in vivo - Towards Response Prediction in Cancer Therapy with Kinase Inhibitors Michaela Bairlein Vollständiger Abdruck der von der Fakultät Wissenschaftszentrum Weihenstephan für Ernährung, Landnutzung und Umwelt der Technischen Universität München zur Erlangung des akademischen Grades eines Doktors der Naturwissenschaften genehmigten Dissertation. Vorsitzender: Univ. -Prof. Dr. K. Schneitz Prüfer der Dissertation: 1. Univ.-Prof. Dr. A. Gierl 2. Hon.-Prof. Dr. h.c. A. Ullrich (Eberhard-Karls-Universität Tübingen) 3. Univ.-Prof. A. Schnieke, Ph.D. Die Dissertation wurde am 07.01.2010 bei der Technischen Universität München eingereicht und durch die Fakultät Wissenschaftszentrum Weihenstephan für Ernährung, Landnutzung und Umwelt am 19.04.2010 angenommen. FOR MY PARENTS 1 Contents 2 Summary ................................................................................................................................................................... 5 3 Zusammenfassung .................................................................................................................................................... 6 4 Introduction .............................................................................................................................................................. 8 4.1 Cancer .............................................................................................................................................................. -

Recurrent Somatic MAP2K1 Mutations in Papillary Thyroid Cancer and Colorectal Cancer
ORIGINAL RESEARCH published: 11 May 2021 doi: 10.3389/fonc.2021.670423 Recurrent Somatic MAP2K1 Mutations in Papillary Thyroid Cancer and Colorectal Cancer † † Rong Bu 1 , Abdul K. Siraj 1 , Tariq Masoodi 1, Sandeep Kumar Parvathareddy 1, Kaleem Iqbal 1, Maha Al-Rasheed 1, Wael Haqawi 1, Mark Diaz 1, Ingrid G. Victoria 1, Saud M. Aldughaither 1, Saif S. Al-Sobhi 2, Fouad Al-Dayel 3 and Khawla S. Al-Kuraya 1* 1 Human Cancer Genomic Research, Research Center, King Faisal Specialist Hospital and Research Center, Riyadh, Saudi Arabia, 2 Department of Surgery, King Faisal Specialist Hospital and Research Center, Riyadh, Saudi Arabia, 3 Department of Pathology, King Faisal Specialist Hospital and Research Centre, Riyadh, Saudi Arabia fi Edited by: Mitogen-activated protein kinase kinase 1 (MAP2K1) is a dual speci city protein kinase Obul Reddy Bandapalli, that phosphorylates both threonine and tyrosine residues in ERK. MAP2K1 mutations Hopp Children’s Cancer Center have been identified in several cancers. However, their role in Middle Eastern papillary Heidelberg (KiTZ), Germany thyroid cancer (PTC) and colorectal cancer (CRC) is lacking. In this study, we evaluated Reviewed by: Shaolei Teng, the prevalence of MAP2K1 mutations in a large cohort of Middle Eastern PTC and CRC Howard University, United States using whole-exome and Sanger sequencing technology. In the discovery cohort of 100 Beifang Niu, Chinese Academy of Sciences (CAS), PTC and 100 CRC cases (comprising 50 MAPK mutant and 50 MAPK wildtype cases China each), we found one MAP2K1 mutation each in PTC and CRC, both of which were MAPK *Correspondence: wildtype. -

Activation of Diverse Signalling Pathways by Oncogenic PIK3CA Mutations
ARTICLE Received 14 Feb 2014 | Accepted 12 Aug 2014 | Published 23 Sep 2014 DOI: 10.1038/ncomms5961 Activation of diverse signalling pathways by oncogenic PIK3CA mutations Xinyan Wu1, Santosh Renuse2,3, Nandini A. Sahasrabuddhe2,4, Muhammad Saddiq Zahari1, Raghothama Chaerkady1, Min-Sik Kim1, Raja S. Nirujogi2, Morassa Mohseni1, Praveen Kumar2,4, Rajesh Raju2, Jun Zhong1, Jian Yang5, Johnathan Neiswinger6, Jun-Seop Jeong6, Robert Newman6, Maureen A. Powers7, Babu Lal Somani2, Edward Gabrielson8, Saraswati Sukumar9, Vered Stearns9, Jiang Qian10, Heng Zhu6, Bert Vogelstein5, Ben Ho Park9 & Akhilesh Pandey1,8,9 The PIK3CA gene is frequently mutated in human cancers. Here we carry out a SILAC-based quantitative phosphoproteomic analysis using isogenic knockin cell lines containing ‘driver’ oncogenic mutations of PIK3CA to dissect the signalling mechanisms responsible for oncogenic phenotypes induced by mutant PIK3CA. From 8,075 unique phosphopeptides identified, we observe that aberrant activation of PI3K pathway leads to increased phosphorylation of a surprisingly wide variety of kinases and downstream signalling networks. Here, by integrating phosphoproteomic data with human protein microarray-based AKT1 kinase assays, we discover and validate six novel AKT1 substrates, including cortactin. Through mutagenesis studies, we demonstrate that phosphorylation of cortactin by AKT1 is important for mutant PI3K-enhanced cell migration and invasion. Our study describes a quantitative and global approach for identifying mutation-specific signalling events and for discovering novel signalling molecules as readouts of pathway activation or potential therapeutic targets. 1 McKusick-Nathans Institute of Genetic Medicine and Department of Biological Chemistry, Johns Hopkins University School of Medicine, 733 North Broadway, BRB 527, Baltimore, Maryland 21205, USA. -

Protein-Protein Interactions Among Signaling Pathways May Become New Therapeutic Targets in Liver Cancer (Review)
ONCOLOGY REPORTS 35: 625-638, 2016 Protein-protein interactions among signaling pathways may become new therapeutic targets in liver cancer (Review) XIAO ZHANG1*, YULAN WANG1*, Jiayi WANG1,2 and FENYONG SUN1 1Department of Clinical Laboratory Medicine, Shanghai Tenth People's Hospital of Tongji University, Shanghai 200072; 2Translation Medicine of High Institute, Tongji University, Shanghai 200092, P.R. China Received May 29, 2015; Accepted July 6, 2015 DOI: 10.3892/or.2015.4464 Abstract. Numerous signaling pathways have been shown to be 1. Introduction dysregulated in liver cancer. In addition, some protein-protein interactions are prerequisite for the uncontrolled activation Liver cancer is the sixth most common cancer and the second or inhibition of these signaling pathways. For instance, in most common cause of cancer-associated mortality world- the PI3K/AKT signaling pathway, protein AKT binds with wide (1). Approximately 75% of all primary liver cancer types a number of proteins such as mTOR, FOXO1 and MDM2 to are hepatocellular carcinoma (HCC) that formed from liver play an oncogenic role in liver cancer. The aim of the present cells. Liver cancer can be formed from other structures in review was to focus on a series of important protein-protein the liver such as bile duct, blood vessels and immune cells. interactions that can serve as potential therapeutic targets Secondary liver cancer is a result of metastasis of cancer from in liver cancer among certain important pro-carcinogenic other body sites into the liver. The major cause of primary liver signaling pathways. The strategies of how to investigate and cancer is viral infection with either hepatitis C virus (HCV) analyze the protein-protein interactions are also included in or hepatitis B virus (HBV), which leads to massive inflamma- this review. -

Inhibitors of Cyclin-Dependent Kinases: Types and Their Mechanism of Action
International Journal of Molecular Sciences Review Inhibitors of Cyclin-Dependent Kinases: Types and Their Mechanism of Action Paweł Łukasik 1, Irena Baranowska-Bosiacka 2, Katarzyna Kulczycka 3 and Izabela Gutowska 1,* 1 Department of Medical Chemistry, Pomeranian Medical University in Szczecin, Powstancow Wlkp. 72 Av., 70-111 Szczecin, Poland; [email protected] 2 Department of Biochemistry, Pomeranian Medical University in Szczecin, Powstancow Wlkp. 72 Av., 70-111 Szczecin, Poland; [email protected] 3 Department of Pharmaceutical Chemistry, Pomeranian Medical University in Szczecin, Powstancow Wlkp. 72 Av., 70-111 Szczecin, Poland; [email protected] * Correspondence: [email protected] Abstract: Recent studies on cyclin-dependent kinase (CDK) inhibitors have revealed that small molecule drugs have become very attractive for the treatment of cancer and neurodegenerative disorders. Most CDK inhibitors have been developed to target the ATP binding pocket. However, CDK kinases possess a very similar catalytic domain and three-dimensional structure. These features make it difficult to achieve required selectivity. Therefore, inhibitors which bind outside the ATP binding site present a great interest in the biomedical field, both from the fundamental point of view and for the wide range of their potential applications. This review tries to explain whether the ATP competitive inhibitors are still an option for future research, and highlights alternative approaches to discover more selective and potent small molecule inhibitors. Keywords: cyclin-dependent kinase inhibitors; cancer; cell cycle; CDKs; CDK inhibitors Citation: Łukasik, P.; Baranowska-Bosiacka, I.; Kulczycka, K.; Gutowska, I. Inhibitors of Cyclin-Dependent Kinases: Types 1. Cyclin-Dependent Kinases (CDKs) and Their Mechanism of Action. -

Inhibition of ERK 1/2 Kinases Prevents Tendon Matrix Breakdown Ulrich Blache1,2,3, Stefania L
www.nature.com/scientificreports OPEN Inhibition of ERK 1/2 kinases prevents tendon matrix breakdown Ulrich Blache1,2,3, Stefania L. Wunderli1,2,3, Amro A. Hussien1,2, Tino Stauber1,2, Gabriel Flückiger1,2, Maja Bollhalder1,2, Barbara Niederöst1,2, Sandro F. Fucentese1 & Jess G. Snedeker1,2* Tendon extracellular matrix (ECM) mechanical unloading results in tissue degradation and breakdown, with niche-dependent cellular stress directing proteolytic degradation of tendon. Here, we show that the extracellular-signal regulated kinase (ERK) pathway is central in tendon degradation of load-deprived tissue explants. We show that ERK 1/2 are highly phosphorylated in mechanically unloaded tendon fascicles in a vascular niche-dependent manner. Pharmacological inhibition of ERK 1/2 abolishes the induction of ECM catabolic gene expression (MMPs) and fully prevents loss of mechanical properties. Moreover, ERK 1/2 inhibition in unloaded tendon fascicles suppresses features of pathological tissue remodeling such as collagen type 3 matrix switch and the induction of the pro-fbrotic cytokine interleukin 11. This work demonstrates ERK signaling as a central checkpoint to trigger tendon matrix degradation and remodeling using load-deprived tissue explants. Tendon is a musculoskeletal tissue that transmits muscle force to bone. To accomplish its biomechanical function, tendon tissues adopt a specialized extracellular matrix (ECM) structure1. Te load-bearing tendon compart- ment consists of highly aligned collagen-rich fascicles that are interspersed with tendon stromal cells. Tendon is a mechanosensitive tissue whereby physiological mechanical loading is vital for maintaining tendon archi- tecture and homeostasis2. Mechanical unloading of the tissue, for instance following tendon rupture or more localized micro trauma, leads to proteolytic breakdown of the tissue with severe deterioration of both structural and mechanical properties3–5.