Structural and Mechanistic Insights Into RAF Kinase Regulation by the KSR/CNK/HYP Complex
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Hidden Targets in RAF Signalling Pathways to Block Oncogenic RAS Signalling
G C A T T A C G G C A T genes Review Hidden Targets in RAF Signalling Pathways to Block Oncogenic RAS Signalling Aoife A. Nolan 1, Nourhan K. Aboud 1, Walter Kolch 1,2,* and David Matallanas 1,* 1 Systems Biology Ireland, School of Medicine, University College Dublin, Belfield, Dublin 4, Ireland; [email protected] (A.A.N.); [email protected] (N.K.A.) 2 Conway Institute of Biomolecular & Biomedical Research, University College Dublin, Belfield, Dublin 4, Ireland * Correspondence: [email protected] (W.K.); [email protected] (D.M.) Abstract: Oncogenic RAS (Rat sarcoma) mutations drive more than half of human cancers, and RAS inhibition is the holy grail of oncology. Thirty years of relentless efforts and harsh disappointments have taught us about the intricacies of oncogenic RAS signalling that allow us to now get a pharma- cological grip on this elusive protein. The inhibition of effector pathways, such as the RAF-MEK-ERK pathway, has largely proven disappointing. Thus far, most of these efforts were aimed at blocking the activation of ERK. Here, we discuss RAF-dependent pathways that are regulated through RAF functions independent of catalytic activity and their potential role as targets to block oncogenic RAS signalling. We focus on the now well documented roles of RAF kinase-independent functions in apoptosis, cell cycle progression and cell migration. Keywords: RAF kinase-independent; RAS; MST2; ASK; PLK; RHO-α; apoptosis; cell cycle; cancer therapy Citation: Nolan, A.A.; Aboud, N.K.; Kolch, W.; Matallanas, D. Hidden Targets in RAF Signalling Pathways to Block Oncogenic RAS Signalling. -

Atlas Antibodies in Breast Cancer Research Table of Contents
ATLAS ANTIBODIES IN BREAST CANCER RESEARCH TABLE OF CONTENTS The Human Protein Atlas, Triple A Polyclonals and PrecisA Monoclonals (4-5) Clinical markers (6) Antibodies used in breast cancer research (7-13) Antibodies against MammaPrint and other gene expression test proteins (14-16) Antibodies identified in the Human Protein Atlas (17-14) Finding cancer biomarkers, as exemplified by RBM3, granulin and anillin (19-22) Co-Development program (23) Contact (24) Page 2 (24) Page 3 (24) The Human Protein Atlas: a map of the Human Proteome The Human Protein Atlas (HPA) is a The Human Protein Atlas consortium cell types. All the IHC images for Swedish-based program initiated in is mainly funded by the Knut and Alice the normal tissue have undergone 2003 with the aim to map all the human Wallenberg Foundation. pathology-based annotation of proteins in cells, tissues and organs expression levels. using integration of various omics The Human Protein Atlas consists of technologies, including antibody- six separate parts, each focusing on References based imaging, mass spectrometry- a particular aspect of the genome- 1. Sjöstedt E, et al. (2020) An atlas of the based proteomics, transcriptomics wide analysis of the human proteins: protein-coding genes in the human, pig, and and systems biology. mouse brain. Science 367(6482) 2. Thul PJ, et al. (2017) A subcellular map of • The Tissue Atlas shows the the human proteome. Science. 356(6340): All the data in the knowledge resource distribution of proteins across all eaal3321 is open access to allow scientists both major tissues and organs in the 3. -

The Prevalence of MADH4 and BMPR1A Mutations in Juvenile Polyposis and Absence of BMPR2, BMPR1B, and ACVR1 Mutations
484 ORIGINAL ARTICLE J Med Genet: first published as 10.1136/jmg.2004.018598 on 2 July 2004. Downloaded from The prevalence of MADH4 and BMPR1A mutations in juvenile polyposis and absence of BMPR2, BMPR1B, and ACVR1 mutations J R Howe, M G Sayed, A F Ahmed, J Ringold, J Larsen-Haidle, A Merg, F A Mitros, C A Vaccaro, G M Petersen, F M Giardiello, S T Tinley, L A Aaltonen, H T Lynch ............................................................................................................................... J Med Genet 2004;41:484–491. doi: 10.1136/jmg.2004.018598 Background: Juvenile polyposis (JP) is an autosomal dominant syndrome predisposing to colorectal and gastric cancer. We have identified mutations in two genes causing JP, MADH4 and bone morphogenetic protein receptor 1A (BMPR1A): both are involved in bone morphogenetic protein (BMP) mediated signalling and are members of the TGF-b superfamily. This study determined the prevalence of mutations See end of article for in MADH4 and BMPR1A, as well as three other BMP/activin pathway candidate genes in a large number authors’ affiliations ....................... of JP patients. Methods: DNA was extracted from the blood of JP patients and used for PCR amplification of each exon of Correspondence to: these five genes, using primers flanking each intron–exon boundary. Mutations were determined by Dr J R Howe, Department of Surgery, 4644 JCP, comparison to wild type sequences using sequence analysis software. A total of 77 JP cases were University of Iowa College sequenced for mutations in the MADH4, BMPR1A, BMPR1B, BMPR2, and/or ACVR1 (activin A receptor) of Medicine, 200 Hawkins genes. The latter three genes were analysed when MADH4 and BMPR1A sequencing found no mutations. -

Casein Kinase 1 Isoforms in Degenerative Disorders
CASEIN KINASE 1 ISOFORMS IN DEGENERATIVE DISORDERS DISSERTATION Presented in Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy in the Graduate School of The Ohio State University By Theresa Joseph Kannanayakal, M.Sc., M.S. * * * * * The Ohio State University 2004 Dissertation Committee: Approved by Professor Jeff A. Kuret, Adviser Professor John D. Oberdick Professor Dale D. Vandre Adviser Professor Mike X. Zhu Biophysics Graduate Program ABSTRACT Casein Kinase 1 (CK1) enzyme is one of the largest family of Serine/Threonine protein kinases. CK1 has a wide distribution spanning many eukaryotic families. In cells, its kinase activity has been found in various sub-cellular compartments enabling it to phosphorylate many proteins involved in cellular maintenance and disease pathogenesis. Tau is one such substrate whose hyperphosphorylation results in degeneration of neurons in Alzheimer’s disease (AD). AD is a slow neuroprogessive disorder histopathologically characterized by Granulovacuolar degeneration bodies (GVBs) and intraneuronal accumulation of tau in Neurofibrillary Tangles (NFTs). The level of CK1 isoforms, CK1α, CK1δ and CK1ε has been shown to be elevated in AD. Previous studies of the correlation of CK1δ with lesions had demonstrated its importance in tau hyperphosphorylation. Hence we investigated distribution of CK1α and CK1ε with the lesions to understand if they would play role in tau hyperphosphorylation similar to CK1δ. The kinase results were also compared with lesion correlation studies of peptidyl cis/trans prolyl isomerase (Pin1) and caspase-3. Our results showed that among the enzymes investigated, CK1 isoforms have the greatest extent of colocalization with the lesions. We have also investigated the distribution of CK1α with different stages of NFTs that follow AD progression. -

Application of a MYC Degradation
SCIENCE SIGNALING | RESEARCH ARTICLE CANCER Copyright © 2019 The Authors, some rights reserved; Application of a MYC degradation screen identifies exclusive licensee American Association sensitivity to CDK9 inhibitors in KRAS-mutant for the Advancement of Science. No claim pancreatic cancer to original U.S. Devon R. Blake1, Angelina V. Vaseva2, Richard G. Hodge2, McKenzie P. Kline3, Thomas S. K. Gilbert1,4, Government Works Vikas Tyagi5, Daowei Huang5, Gabrielle C. Whiten5, Jacob E. Larson5, Xiaodong Wang2,5, Kenneth H. Pearce5, Laura E. Herring1,4, Lee M. Graves1,2,4, Stephen V. Frye2,5, Michael J. Emanuele1,2, Adrienne D. Cox1,2,6, Channing J. Der1,2* Stabilization of the MYC oncoprotein by KRAS signaling critically promotes the growth of pancreatic ductal adeno- carcinoma (PDAC). Thus, understanding how MYC protein stability is regulated may lead to effective therapies. Here, we used a previously developed, flow cytometry–based assay that screened a library of >800 protein kinase inhibitors and identified compounds that promoted either the stability or degradation of MYC in a KRAS-mutant PDAC cell line. We validated compounds that stabilized or destabilized MYC and then focused on one compound, Downloaded from UNC10112785, that induced the substantial loss of MYC protein in both two-dimensional (2D) and 3D cell cultures. We determined that this compound is a potent CDK9 inhibitor with a previously uncharacterized scaffold, caused MYC loss through both transcriptional and posttranslational mechanisms, and suppresses PDAC anchorage- dependent and anchorage-independent growth. We discovered that CDK9 enhanced MYC protein stability 62 through a previously unknown, KRAS-independent mechanism involving direct phosphorylation of MYC at Ser . -

RAF Protein-Serine/Threonine Kinases: Structure and Regulation
Biochemical and Biophysical Research Communications 399 (2010) 313–317 Contents lists available at ScienceDirect Biochemical and Biophysical Research Communications journal homepage: www.elsevier.com/locate/ybbrc Mini Review RAF protein-serine/threonine kinases: Structure and regulation Robert Roskoski Jr. * Blue Ridge Institute for Medical Research, 3754 Brevard Road, Suite 116, Box 19, Horse Shoe, NC 28742, USA article info abstract Article history: A-RAF, B-RAF, and C-RAF are a family of three protein-serine/threonine kinases that participate in the Received 12 July 2010 RAS-RAF-MEK-ERK signal transduction cascade. This cascade participates in the regulation of a large vari- Available online 30 July 2010 ety of processes including apoptosis, cell cycle progression, differentiation, proliferation, and transforma- tion to the cancerous state. RAS mutations occur in 15–30% of all human cancers, and B-RAF mutations Keywords: occur in 30–60% of melanomas, 30–50% of thyroid cancers, and 5–20% of colorectal cancers. Activation 14-3-3 of the RAF kinases requires their interaction with RAS-GTP along with dephosphorylation and also phos- ERK phorylation by SRC family protein-tyrosine kinases and other protein-serine/threonine kinases. The for- GDC-0879 mation of unique side-to-side RAF dimers is required for full kinase activity. RAF kinase inhibitors are MEK Melanoma effective in blocking MEK1/2 and ERK1/2 activation in cells containing the oncogenic B-RAF Val600Glu PLX4032 activating mutation. RAF kinase inhibitors lead to the paradoxical increase in RAF kinase activity in cells PLX4720 containing wild-type B-RAF and wild-type or activated mutant RAS. -
Protein Kinases Phosphorylation/Dephosphorylation Protein Phosphorylation Is One of the Most Important Mechanisms of Cellular Re
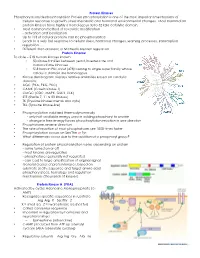
Protein Kinases Phosphorylation/dephosphorylation Protein phosphorylation is one of the most important mechanisms of cellular responses to growth, stress metabolic and hormonal environmental changes. Most mammalian protein kinases have highly a homologous 30 to 32 kDa catalytic domain. • Most common method of reversible modification - activation and localization • Up to 1/3 of cellular proteins can be phosphorylated • Leads to a very fast response to cellular stress, hormonal changes, learning processes, transcription regulation .... • Different than allosteric or Michealis Menten regulation Protein Kinome To date – 518 human kinases known • 50 kinase families between yeast, invertebrate and mammaliane kinomes • 518 human PKs, most (478) belong to single super family whose catalytic domain are homologous. • Kinase dendrogram displays relative similarities based on catalytic domains. • AGC (PKA, PKG, PKC) • CAMK (Casein kinase 1) • CMGC (CDC, MAPK, GSK3, CLK) • STE (Sterile 7, 11 & 20 kinases) • TK (Tryosine kinases memb and cyto) • TKL (Tyrosine kinase-like) • Phosphorylation stabilized thermodynamically - only half available energy used in adding phosphoryl to protein - change in free energy forces phosphorylation reaction in one direction • Phosphatases reverse direction • The rate of reaction of most phosphatases are 1000 times faster • Phosphorylation occurs on Ser/The or Tyr • What differences occur due to the addition of a phosphoryl group? • Regulation of protein phosphorylation varies depending on protein - some turned on or off -

Analysis of Protein Kinase Domain and Tyrosine Kinase Or Serine/Threonine Kinase Signatures Involved in Lung Cancer
Analysis of Protein Kinase Domain and Tyrosine Kinase Or serine/Threonine Kinase Signatures Involved In Lung Cancer Lung cancer results when normal check and balance system of cell division is disrupted and ultimately the cells divide and proliferate in an uncontrollable manner forming a mass of cells in our body, known as tumor. Frequent mutations in Protein Kinase Domain alter the process of phosphorylation which results in abnormality in regulations of cell apoptosis and differentiation. Tyrosine Protein kinases and Serine/Threonine Protein Kinases are the two s t broad classes of protein kinases in accordance to their substrate specificity. The study of n i r Tyrosine protein kinase and serine Kinase coding regions have the importance of sequence P and structure determinants of cancer-causing mutations from mutation-dependent activation e r P process. In the present study, we analyzed huge amounts of data extracted from various biological databases and NCBI. Out of the 534 proteins that may play a role in lung cancer, 71 proteins were selected that are likely to be actively involved in lung cancer. These proteins were evaluated by employing Multiple Sequence Alignment and a Phylogenetic tree was constructed using Neighbor-Joining Algorithm. From the constructed phylogenetic tree, protein kinase domain and motif study was performed. The results of this study revealed that the presence of Protein Kinase Domain and Tyrosine or Serine/Threonine Kinase signatures in some of the proteins are mutated, which play a dominant role in the pathogenesis of Lung Cancer and these may be addressed with the help of inhibitors to develop an efficient anticancer drugs. -

Mutation-Specific RAS Oncogenicity Explains NRAS Codon 61 Selection in Melanoma
Published OnlineFirst September 24, 2014; DOI: 10.1158/2159-8290.CD-14-0729 reseArch Article Mutation-Specific RAS Oncogenicity Explains NRAS Codon 61 Selection in Melanoma Christin E. Burd1,2, Wenjin Liu3,4, Minh V. Huynh5, Meriam A. Waqas1, James E. Gillahan1,2, Kelly S. Clark3,4, Kailing Fu3,4, Brit L. Martin1, William R. Jeck3,4,. George P Souroullas3,4, David B. Darr3,4, Daniel C. Zedek6, Michael J. Miley7, Bruce C. Baguley8, Sharon L. Campbell4,5, and Norman E. Sharpless3,4 Abstrt Ac NRAS mutation at codons 12, 13, or 61 is associated with transformation; yet, in melanoma, such alterations are nearly exclusive to codon 61. Here, we compared the melanoma susceptibility of an NrasQ61R knock-in allele to similarly designed KrasG12D and NrasG12D alleles. With concomitant p16INK4a inactivation, KrasG12D or NrasQ61R expression efficiently promoted melanoma in vivo, whereas NrasG12D did not. In addition, NrasQ61R mutation potently cooperated with Lkb1/Stk11 loss to induce highly metastatic disease. Functional comparisons of NrasQ61R and NrasG12D revealed little difference in the ability of these proteins to engage PI3K or RAF. Instead, NrasQ61R showed enhanced nucleotide binding, decreased intrinsic GTPase activity, and increased stability when compared with NrasG12D. This work identifies a faithful model of human NRAS-mutant melanoma, and suggests that the increased melanomagenecity of NrasQ61R over NrasG12D is due to heightened abundance of the active, GTP-bound form rather than differences in the engagement of downstream effector pathways. SIGNIFICANCE: This work explains the curious predominance in human melanoma of mutations of codon 61 of NRAS over other oncogenic NRAS mutations. -

Supplementary Table 1. in Vitro Side Effect Profiling Study for LDN/OSU-0212320. Neurotransmitter Related Steroids
Supplementary Table 1. In vitro side effect profiling study for LDN/OSU-0212320. Percent Inhibition Receptor 10 µM Neurotransmitter Related Adenosine, Non-selective 7.29% Adrenergic, Alpha 1, Non-selective 24.98% Adrenergic, Alpha 2, Non-selective 27.18% Adrenergic, Beta, Non-selective -20.94% Dopamine Transporter 8.69% Dopamine, D1 (h) 8.48% Dopamine, D2s (h) 4.06% GABA A, Agonist Site -16.15% GABA A, BDZ, alpha 1 site 12.73% GABA-B 13.60% Glutamate, AMPA Site (Ionotropic) 12.06% Glutamate, Kainate Site (Ionotropic) -1.03% Glutamate, NMDA Agonist Site (Ionotropic) 0.12% Glutamate, NMDA, Glycine (Stry-insens Site) 9.84% (Ionotropic) Glycine, Strychnine-sensitive 0.99% Histamine, H1 -5.54% Histamine, H2 16.54% Histamine, H3 4.80% Melatonin, Non-selective -5.54% Muscarinic, M1 (hr) -1.88% Muscarinic, M2 (h) 0.82% Muscarinic, Non-selective, Central 29.04% Muscarinic, Non-selective, Peripheral 0.29% Nicotinic, Neuronal (-BnTx insensitive) 7.85% Norepinephrine Transporter 2.87% Opioid, Non-selective -0.09% Opioid, Orphanin, ORL1 (h) 11.55% Serotonin Transporter -3.02% Serotonin, Non-selective 26.33% Sigma, Non-Selective 10.19% Steroids Estrogen 11.16% 1 Percent Inhibition Receptor 10 µM Testosterone (cytosolic) (h) 12.50% Ion Channels Calcium Channel, Type L (Dihydropyridine Site) 43.18% Calcium Channel, Type N 4.15% Potassium Channel, ATP-Sensitive -4.05% Potassium Channel, Ca2+ Act., VI 17.80% Potassium Channel, I(Kr) (hERG) (h) -6.44% Sodium, Site 2 -0.39% Second Messengers Nitric Oxide, NOS (Neuronal-Binding) -17.09% Prostaglandins Leukotriene, -

Combined Inhibition of MEK and Plk1 Has Synergistic Anti-Tumor Activity in NRAS Mutant Melanoma
Combined inhibition of MEK and Plk1 has synergistic anti-tumor activity in NRAS mutant melanoma The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Posch, C., B. Cholewa, I. Vujic, M. Sanlorenzo, J. Ma, S. Kim, S. Kleffel, et al. 2015. “Combined inhibition of MEK and Plk1 has synergistic anti-tumor activity in NRAS mutant melanoma.” The Journal of investigative dermatology 135 (10): 2475-2483. doi:10.1038/jid.2015.198. http://dx.doi.org/10.1038/jid.2015.198. Published Version doi:10.1038/jid.2015.198 Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:26860102 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA HHS Public Access Author manuscript Author Manuscript Author ManuscriptJ Invest Author ManuscriptDermatol. Author Author Manuscript manuscript; available in PMC 2016 April 01. Published in final edited form as: J Invest Dermatol. 2015 October ; 135(10): 2475–2483. doi:10.1038/jid.2015.198. Combined inhibition of MEK and Plk1 has synergistic anti-tumor activity in NRAS mutant melanoma C Posch#1,2,3,*, BD Cholewa#4, I Vujic1,3, M Sanlorenzo1,5, J Ma1, ST Kim1, S Kleffel2, T Schatton2, K Rappersberger3, R Gutteridge4, N Ahmad4, and S Ortiz/Urda1 1 University of California San Francisco, Department of Dermatology, Mt. Zion Cancer Research -

The Role of Casein Kinase 1 in Meiosis
e & Ch m rom ro o d s n o y m S Zhao and Liang, J Down Syndr Chr Abnorm 2016, 2:1 e n A Journal of Down Syndrome & w b o n DOI: 10.4172/2472-1115.1000106 D o f r m o l ISSN: 2472-1115a a l i n t r i e u s o J Chromosome Abnormalities Review Article Open Access The Role of Casein Kinase 1 in Meiosis Yue-Fang Zhao and Cheng-Guang Liang* The Key Laboratory of National Education Ministry for Mammalian Reproductive Biology and Biotechnology, The Research Center for Laboratory Animal Science, College of Life Science, Inner Mongolia University, Hohhot, Inner Mongolia, People’s Republic of China Abstract The casein kinase 1 (CK1) belongs to the serine/threonine protein kinases. CK1 members, which commonly exist in all eukaryotes, are involved in the regulation of many cellular processes linked to cell cycle progression, spindle- dynamics, and chromosome segregation. Additionally, CK1 regulate key signaling pathways such as Wnt (Wingless/ Int-1), Hh (Hedgehog), and Hippo, known to be critically involved in tumor progression. Considering the importance of CK1 for accurate cell division and regulation of tumor suppressor functions, it is not surprising scientific effort has enormously increased. In mammals, CK1 regulate the transition from interphase to metaphase in mitosis. In budding yeast and fission yeast, CK1 phosphorylate Rec8 subunits of cohesin complex and regulate chromosome segregation in meiosis. During meiosis, two rounds of chromosome segregation after a single round of DNA replication produce haploid gametes from diploid precursors. Any mistake in chromosome segregation may result in aneuploidy, which in meiosis are one of the main causes of infertility, abortion and many genetic diseases in humans.