Anti-GPR77 / C5L2 Antibody (ARG56169)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Deciphering the Functions of Ets2, Pten and P53 in Stromal Fibroblasts in Multiple
Deciphering the Functions of Ets2, Pten and p53 in Stromal Fibroblasts in Multiple Breast Cancer Models DISSERTATION Presented in Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy in the Graduate School of The Ohio State University By Julie Wallace Graduate Program in Molecular, Cellular and Developmental Biology The Ohio State University 2013 Dissertation Committee: Michael C. Ostrowski, PhD, Advisor Gustavo Leone, PhD Denis Guttridge, PhD Dawn Chandler, PhD Copyright by Julie Wallace 2013 Abstract Breast cancer is the second most common cancer in American women, and is also the second leading cause of cancer death in women. It is estimated that nearly a quarter of a million new cases of invasive breast cancer will be diagnosed in women in the United States this year, and approximately 40,000 of these women will die from breast cancer. Although death rates have been on the decline for the past decade, there is still much we need to learn about this disease to improve prevention, detection and treatment strategies. The majority of early studies have focused on the malignant tumor cells themselves, and much has been learned concerning mutations, amplifications and other genetic and epigenetic alterations of these cells. However more recent work has acknowledged the strong influence of tumor stroma on the initiation, progression and recurrence of cancer. Under normal conditions this stroma has been shown to have protective effects against tumorigenesis, however the transformation of tumor cells manipulates this surrounding environment to actually promote malignancy. Fibroblasts in particular make up a significant portion of this stroma, and have been shown to impact various aspects of tumor cell biology. -

Targeting and Reprograming Cancer-Associated Fibroblasts and the Tumor Microenvironment in Pancreatic Cancer
cancers Review Targeting and Reprograming Cancer-Associated Fibroblasts and the Tumor Microenvironment in Pancreatic Cancer Yoshiaki Sunami * , Viktoria Böker and Jörg Kleeff Department of Visceral, Vascular and Endocrine Surgery, Martin-Luther-University Halle-Wittenberg, University Medical Center Halle, 06120 Halle, Germany; [email protected] (V.B.); [email protected] (J.K.) * Correspondence: [email protected]; Tel.: +49-345-557-2794 Simple Summary: The tumor microenvironment plays a major role in the progression and drug resistance of pancreatic cancer. Cancer-associated fibroblasts are the major stromal cells and source of extracellular matrix proteins forming the dense stromal tumor microenvironment. Targeting cancer-associated fibroblasts has been deemed a promising therapeutic strategy. However, deplet- ing cancer-associated fibroblasts may also have tumor-promoting effects due to their functional heterogeneity. It is therefore important to target selectively the tumor-promoting subtype of cancer- associated fibroblasts. Furthermore, deactivating fibroblasts, or reprograming tumor-promoting cancer-associated fibroblasts to tumor-restraining cancer-associated fibroblasts are considered as therapy for pancreatic cancer. Abstract: Pancreatic cancer is the fourth leading cause of cancer deaths in the United States both in female and male, and is projected to become the second deadliest cancer by 2030. The overall five-year survival rate remains at around 10%. Pancreatic cancer exhibits a remarkable resistance to established therapeutic options such as chemotherapy and radiotherapy, due to dense stromal Citation: Sunami, Y.; Böker, V.; tumor microenvironment. Cancer-associated fibroblasts are the major stromal cell type and source of Kleeff, J. Targeting and Reprograming extracellular matrix proteins shaping a physical and metabolic barrier thereby reducing therapeutic Cancer-Associated Fibroblasts and efficacy. -

Edinburgh Research Explorer
Edinburgh Research Explorer International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list Citation for published version: Davenport, AP, Alexander, SPH, Sharman, JL, Pawson, AJ, Benson, HE, Monaghan, AE, Liew, WC, Mpamhanga, CP, Bonner, TI, Neubig, RR, Pin, JP, Spedding, M & Harmar, AJ 2013, 'International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list: recommendations for new pairings with cognate ligands', Pharmacological reviews, vol. 65, no. 3, pp. 967-86. https://doi.org/10.1124/pr.112.007179 Digital Object Identifier (DOI): 10.1124/pr.112.007179 Link: Link to publication record in Edinburgh Research Explorer Document Version: Publisher's PDF, also known as Version of record Published In: Pharmacological reviews Publisher Rights Statement: U.S. Government work not protected by U.S. copyright General rights Copyright for the publications made accessible via the Edinburgh Research Explorer is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The University of Edinburgh has made every reasonable effort to ensure that Edinburgh Research Explorer content complies with UK legislation. If you believe that the public display of this file breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 02. Oct. 2021 1521-0081/65/3/967–986$25.00 http://dx.doi.org/10.1124/pr.112.007179 PHARMACOLOGICAL REVIEWS Pharmacol Rev 65:967–986, July 2013 U.S. -

Current Understanding of Epigenetics Mechanism As a Novel Target In
Keyvani‑Ghamsari et al. Clin Epigenet (2021) 13:120 https://doi.org/10.1186/s13148‑021‑01107‑4 REVIEW Open Access Current understanding of epigenetics mechanism as a novel target in reducing cancer stem cells resistance Saeedeh Keyvani‑Ghamsari1, Khatereh Khorsandi2* , Azhar Rasul3 and Muhammad Khatir Zaman4 Abstract At present, after extensive studies in the feld of cancer, cancer stem cells (CSCs) have been proposed as a major fac‑ tor in tumor initiation, progression, metastasis, and recurrence. CSCs are a subpopulation of bulk tumors, with stem cell‑like properties and tumorigenic capabilities, having the abilities of self‑renewal and diferentiation, thereby being able to generate heterogeneous lineages of cancer cells and lead to resistance toward anti‑tumor treatments. Highly resistant to conventional chemo‑ and radiotherapy, CSCs have heterogeneity and can migrate to diferent organs and metastasize. Recent studies have demonstrated that the population of CSCs and the progression of cancer are increased by the deregulation of diferent epigenetic pathways having efects on gene expression patterns and key pathways connected with cell proliferation and survival. Further, epigenetic modifcations (DNA methylation, histone modifcations, and RNA methylations) have been revealed to be key drivers in the formation and maintenance of CSCs. Hence, identifying CSCs and targeting epigenetic pathways therein can ofer new insights into the treatment of cancer. In the present review, recent studies are addressed in terms of the characteristics of CSCs, the resistance thereof, and the factors infuencing the development thereof, with an emphasis on diferent types of epigenetic changes in genes and main signaling pathways involved therein. Finally, targeted therapy for CSCs by epigenetic drugs is referred to, which is a new approach in overcoming resistance and recurrence of cancer. -

G Protein-Coupled Receptors
S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2015/16: G protein-coupled receptors. British Journal of Pharmacology (2015) 172, 5744–5869 THE CONCISE GUIDE TO PHARMACOLOGY 2015/16: G protein-coupled receptors Stephen PH Alexander1, Anthony P Davenport2, Eamonn Kelly3, Neil Marrion3, John A Peters4, Helen E Benson5, Elena Faccenda5, Adam J Pawson5, Joanna L Sharman5, Christopher Southan5, Jamie A Davies5 and CGTP Collaborators 1School of Biomedical Sciences, University of Nottingham Medical School, Nottingham, NG7 2UH, UK, 2Clinical Pharmacology Unit, University of Cambridge, Cambridge, CB2 0QQ, UK, 3School of Physiology and Pharmacology, University of Bristol, Bristol, BS8 1TD, UK, 4Neuroscience Division, Medical Education Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, DD1 9SY, UK, 5Centre for Integrative Physiology, University of Edinburgh, Edinburgh, EH8 9XD, UK Abstract The Concise Guide to PHARMACOLOGY 2015/16 provides concise overviews of the key properties of over 1750 human drug targets with their pharmacology, plus links to an open access knowledgebase of drug targets and their ligands (www.guidetopharmacology.org), which provides more detailed views of target and ligand properties. The full contents can be found at http://onlinelibrary.wiley.com/doi/ 10.1111/bph.13348/full. G protein-coupled receptors are one of the eight major pharmacological targets into which the Guide is divided, with the others being: ligand-gated ion channels, voltage-gated ion channels, other ion channels, nuclear hormone receptors, catalytic receptors, enzymes and transporters. These are presented with nomenclature guidance and summary information on the best available pharmacological tools, alongside key references and suggestions for further reading. -

G Protein‐Coupled Receptors
S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2019/20: G protein-coupled receptors. British Journal of Pharmacology (2019) 176, S21–S141 THE CONCISE GUIDE TO PHARMACOLOGY 2019/20: G protein-coupled receptors Stephen PH Alexander1 , Arthur Christopoulos2 , Anthony P Davenport3 , Eamonn Kelly4, Alistair Mathie5 , John A Peters6 , Emma L Veale5 ,JaneFArmstrong7 , Elena Faccenda7 ,SimonDHarding7 ,AdamJPawson7 , Joanna L Sharman7 , Christopher Southan7 , Jamie A Davies7 and CGTP Collaborators 1School of Life Sciences, University of Nottingham Medical School, Nottingham, NG7 2UH, UK 2Monash Institute of Pharmaceutical Sciences and Department of Pharmacology, Monash University, Parkville, Victoria 3052, Australia 3Clinical Pharmacology Unit, University of Cambridge, Cambridge, CB2 0QQ, UK 4School of Physiology, Pharmacology and Neuroscience, University of Bristol, Bristol, BS8 1TD, UK 5Medway School of Pharmacy, The Universities of Greenwich and Kent at Medway, Anson Building, Central Avenue, Chatham Maritime, Chatham, Kent, ME4 4TB, UK 6Neuroscience Division, Medical Education Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, DD1 9SY, UK 7Centre for Discovery Brain Sciences, University of Edinburgh, Edinburgh, EH8 9XD, UK Abstract The Concise Guide to PHARMACOLOGY 2019/20 is the fourth in this series of biennial publications. The Concise Guide provides concise overviews of the key properties of nearly 1800 human drug targets with an emphasis on selective pharmacology (where available), plus links to the open access knowledgebase source of drug targets and their ligands (www.guidetopharmacology.org), which provides more detailed views of target and ligand properties. Although the Concise Guide represents approximately 400 pages, the material presented is substantially reduced compared to information and links presented on the website. -

Early During Myelomagenesis Alterations in DNA Methylation That
Myeloma Is Characterized by Stage-Specific Alterations in DNA Methylation That Occur Early during Myelomagenesis This information is current as Christoph J. Heuck, Jayesh Mehta, Tushar Bhagat, Krishna of September 23, 2021. Gundabolu, Yiting Yu, Shahper Khan, Grigoris Chrysofakis, Carolina Schinke, Joseph Tariman, Eric Vickrey, Natalie Pulliam, Sangeeta Nischal, Li Zhou, Sanchari Bhattacharyya, Richard Meagher, Caroline Hu, Shahina Maqbool, Masako Suzuki, Samir Parekh, Frederic Reu, Ulrich Steidl, John Greally, Amit Verma and Seema B. Downloaded from Singhal J Immunol 2013; 190:2966-2975; Prepublished online 13 February 2013; doi: 10.4049/jimmunol.1202493 http://www.jimmunol.org/content/190/6/2966 http://www.jimmunol.org/ Supplementary http://www.jimmunol.org/content/suppl/2013/02/13/jimmunol.120249 Material 3.DC1 References This article cites 38 articles, 15 of which you can access for free at: http://www.jimmunol.org/content/190/6/2966.full#ref-list-1 by guest on September 23, 2021 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. -

Br J Pharmacol
Supplementary Information Supplementary Table 1. List of polypeptide cell surface receptor and their cognate ligand genes. Supplementary Table 2. List of coding SNPs with high FST ( > 0.5) in human GPCR and their cognate ligand genes. Supplementary Table 3. List of coding SNPs with high FST ( > 0.5) in human nonGPCR receptor and ligand genes. Supplementary Table 4. List of genotyped SNPs from 44246961 to 44542055 on chromosome 17. The HapMap II dataset was analyzed using HaploView. The 101 SNPs included in LD plots of Supplementary Fig. 3 (A-C) are highlighted by a grey background. The 37 SNPs used in the haplotype analysis of Fig. 2C are indicated by red letters. SNPs that are linked with rs2291725 are indicated by bold red letters. Supplementary Table 5. Allele frequency of rs2291725 in the HGDP-CEPH populations. Frequencies of GIP103T and GIP103C alleles in each of the 52 populations from the seven geographical regions and the number of chromosomes analyzed for each population. 1 Supplementary Table 6. EC50 values for GIP receptor activation at four different time points after incubation with pooled human serum or pooled complement-preserved human serum (N=4). Supplementary Table 7. EC50 values for GIP receptor activation at three different time points after incubation with a recombinant DPP IV enzyme (N=4). Supplementary Fig. 1. Cumulative distribution function (CDF) plots for the FST of coding SNPs in human GPCRs and their cognate ligand genes (blue) and all other human genes (magenta). The FST was computed between three HapMap II populations (CEU, YRI, and ASN), and coding SNPs have been split into synonymous and nonsynonymous groups. -

Adenylyl Cyclase 2 Selectively Regulates IL-6 Expression in Human Bronchial Smooth Muscle Cells Amy Sue Bogard University of Tennessee Health Science Center
University of Tennessee Health Science Center UTHSC Digital Commons Theses and Dissertations (ETD) College of Graduate Health Sciences 12-2013 Adenylyl Cyclase 2 Selectively Regulates IL-6 Expression in Human Bronchial Smooth Muscle Cells Amy Sue Bogard University of Tennessee Health Science Center Follow this and additional works at: https://dc.uthsc.edu/dissertations Part of the Medical Cell Biology Commons, and the Medical Molecular Biology Commons Recommended Citation Bogard, Amy Sue , "Adenylyl Cyclase 2 Selectively Regulates IL-6 Expression in Human Bronchial Smooth Muscle Cells" (2013). Theses and Dissertations (ETD). Paper 330. http://dx.doi.org/10.21007/etd.cghs.2013.0029. This Dissertation is brought to you for free and open access by the College of Graduate Health Sciences at UTHSC Digital Commons. It has been accepted for inclusion in Theses and Dissertations (ETD) by an authorized administrator of UTHSC Digital Commons. For more information, please contact [email protected]. Adenylyl Cyclase 2 Selectively Regulates IL-6 Expression in Human Bronchial Smooth Muscle Cells Document Type Dissertation Degree Name Doctor of Philosophy (PhD) Program Biomedical Sciences Track Molecular Therapeutics and Cell Signaling Research Advisor Rennolds Ostrom, Ph.D. Committee Elizabeth Fitzpatrick, Ph.D. Edwards Park, Ph.D. Steven Tavalin, Ph.D. Christopher Waters, Ph.D. DOI 10.21007/etd.cghs.2013.0029 Comments Six month embargo expired June 2014 This dissertation is available at UTHSC Digital Commons: https://dc.uthsc.edu/dissertations/330 Adenylyl Cyclase 2 Selectively Regulates IL-6 Expression in Human Bronchial Smooth Muscle Cells A Dissertation Presented for The Graduate Studies Council The University of Tennessee Health Science Center In Partial Fulfillment Of the Requirements for the Degree Doctor of Philosophy From The University of Tennessee By Amy Sue Bogard December 2013 Copyright © 2013 by Amy Sue Bogard. -

MRGPRX4 Is a Novel Bile Acid Receptor in Cholestatic Itch Huasheng Yu1,2,3, Tianjun Zhao1,2,3, Simin Liu1, Qinxue Wu4, Omar
bioRxiv preprint doi: https://doi.org/10.1101/633446; this version posted May 9, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 MRGPRX4 is a novel bile acid receptor in cholestatic itch 2 Huasheng Yu1,2,3, Tianjun Zhao1,2,3, Simin Liu1, Qinxue Wu4, Omar Johnson4, Zhaofa 3 Wu1,2, Zihao Zhuang1, Yaocheng Shi5, Renxi He1,2, Yong Yang6, Jianjun Sun7, 4 Xiaoqun Wang8, Haifeng Xu9, Zheng Zeng10, Xiaoguang Lei3,5, Wenqin Luo4*, Yulong 5 Li1,2,3* 6 7 1State Key Laboratory of Membrane Biology, Peking University School of Life 8 Sciences, Beijing 100871, China 9 2PKU-IDG/McGovern Institute for Brain Research, Beijing 100871, China 10 3Peking-Tsinghua Center for Life Sciences, Beijing 100871, China 11 4Department of Neuroscience, Perelman School of Medicine, University of 12 Pennsylvania, Philadelphia, PA 19104, USA 13 5Department of Chemical Biology, College of Chemistry and Molecular Engineering, 14 Peking University, Beijing 100871, China 15 6Department of Dermatology, Peking University First Hospital, Beijing Key Laboratory 16 of Molecular Diagnosis on Dermatoses, Beijing 100034, China 17 7Department of Neurosurgery, Peking University Third Hospital, Peking University, 18 Beijing, 100191, China 19 8State Key Laboratory of Brain and Cognitive Science, CAS Center for Excellence in 20 Brain Science and Intelligence Technology (Shanghai), Institute of Biophysics, 21 Chinese Academy of Sciences, Beijing, 100101, China 22 9Department of Liver Surgery, Peking Union Medical College Hospital, Chinese bioRxiv preprint doi: https://doi.org/10.1101/633446; this version posted May 9, 2019. -

Receptor Signaling Through Osteoclast-Associated Monocyte
Downloaded from http://www.jimmunol.org/ by guest on September 29, 2021 is online at: average * The Journal of Immunology The Journal of Immunology , 20 of which you can access for free at: 2015; 194:3169-3179; Prepublished online 27 from submission to initial decision 4 weeks from acceptance to publication February 2015; doi: 10.4049/jimmunol.1402800 http://www.jimmunol.org/content/194/7/3169 Collagen Induces Maturation of Human Monocyte-Derived Dendritic Cells by Signaling through Osteoclast-Associated Receptor Heidi S. Schultz, Louise M. Nitze, Louise H. Zeuthen, Pernille Keller, Albrecht Gruhler, Jesper Pass, Jianhe Chen, Li Guo, Andrew J. Fleetwood, John A. Hamilton, Martin W. Berchtold and Svetlana Panina J Immunol cites 43 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Author Choice option Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Freely available online through http://www.jimmunol.org/content/suppl/2015/02/27/jimmunol.140280 0.DCSupplemental This article http://www.jimmunol.org/content/194/7/3169.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Author Choice Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2015 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. -
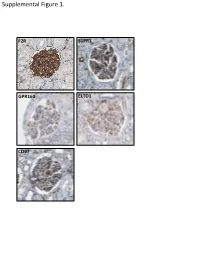
Supplemental Data
Supplemental Figure 1. F2R S1PR1 GPR160 ELTD1 CD97 Supplemental Figure 2. A B brain heart lung liver kidney spleen testis skm Expression in mouse ssues glom rok Gprc5a Gapdh Human Protein Atlas RNAseq database Supplemental Figure 3. A Gprc5a Pdgfrb Merged CL CL Gprc5a CD31 Merged CL CL B C 1.4 * 1.2 mc 1 matrix 0.8 0.6 0.4 matrix 0.2 0 pod end 1 2 mes3 D Vector Gprc5a E 40 kD - - Gprc5a - acn Supplemental Figure 4. Control 12 month-old KO 12 month-old A B C D E F 1 2 0.8 * 1.5 score 0.6 m µ 1 0.4 0.2 Slits/ 0.5 Mesangial 0 0 Ctrl KO Ctrl KO Supplemental Figure 5. A B 1.2 Vector Gprc5a 1 * 0.8 0.6 * 0.4 * Normalized Density 0.2 0 pEGFR/tEGR pSmad/tSmad TGF-β1 C 1.8 siCON 1.6 siGprc5a * 1.4 * * 1.2 1 0.8 0.6 NormalizedDensity 0.4 0.2 0 pEGFR/tEGR pSmad/tSmad TGF-β1 Supplemental table 1. List of glomerulus-expressed GPCRs as detected by qPCR. Data shown as mean ± standard deviation (Glom=glomerulus, Rok=rest of kidney). GPCR Glom Rok Glom/Rok LPAR6 41892,11 ± 38478,89 1040,06 ± 1370,12 39,28 ELTD1 30275,64 ± 14085,26 33,23 ± 46,99 910,13 GPR116 24020,06 ± 7789,84 55,1 ± 47,75 434,96 PTH1R 15402,81 ± 17644,32 7521,01 ± 3264,57 1,05 CALCRL 14096,09 ± 3854,84 199,06 ± 222,52 69,81 HPRT1 13342,04 ± 10677,69 1824,77 ± 1767,23 6,31 S1PR5 9474,7 ± 10124,3 110,84 ± 29,54 84,48 LPHN2 8645,89 ± 914,74 256,04 ± 293,87 32,77 FZD1 8176,2 ± 4947,45 1321,97 ± 1311,91 5,18 CXCR4 7097,31 ± 4388,91 535,98 ± 640,28 12,24 GPR160 6446,59 ± 1550,07 4816,3 ± 5918,99 0,34 NPY1R 6177,74 ± 6282,76 209,64 ± 296,48 28,47 PTGER4 5323,66 ± 3789,51 179,13 ± 210 28,72 RXFP1