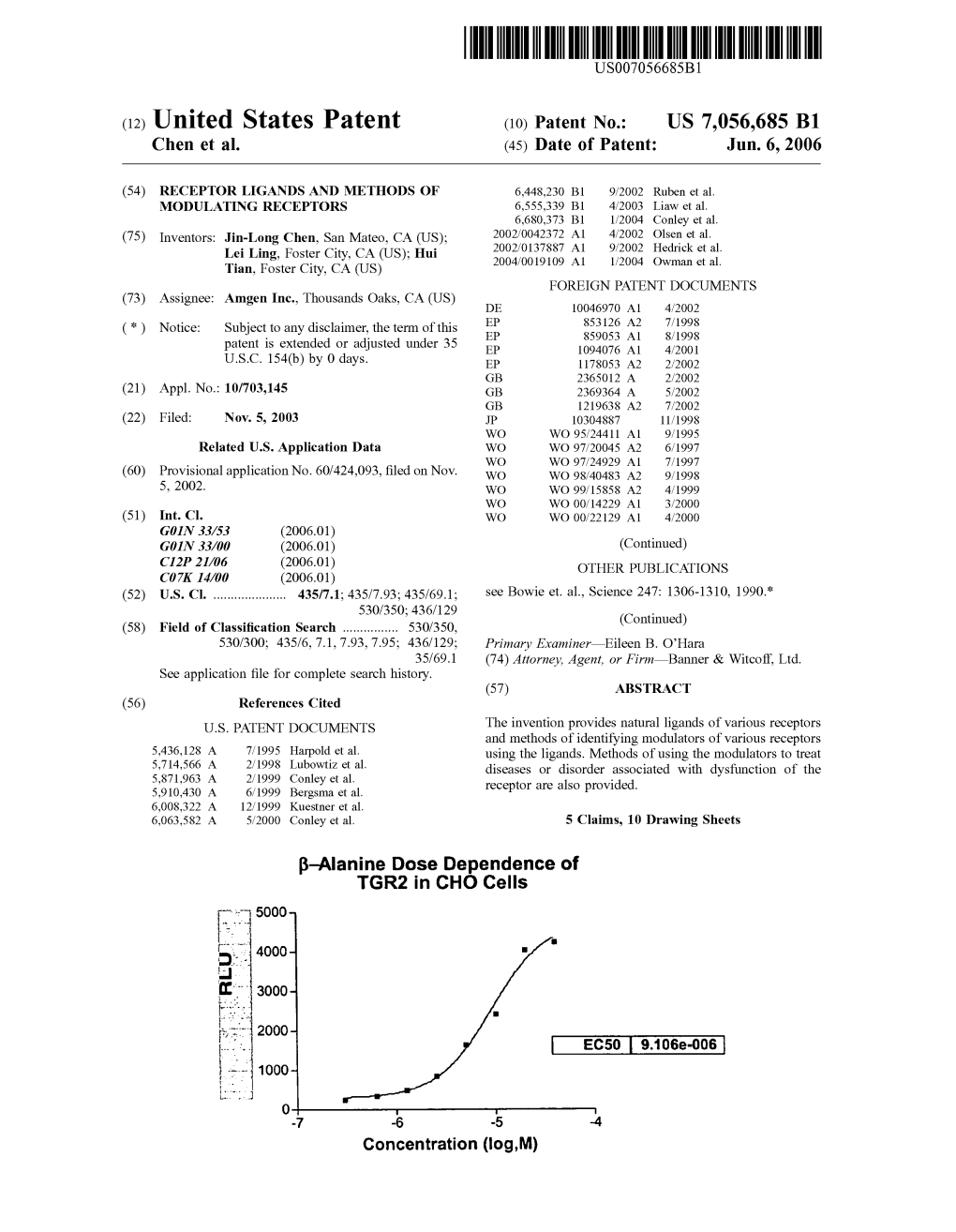
United States Patent B-Alanine Dose Dependence Of
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Deciphering the Functions of Ets2, Pten and P53 in Stromal Fibroblasts in Multiple
Deciphering the Functions of Ets2, Pten and p53 in Stromal Fibroblasts in Multiple Breast Cancer Models DISSERTATION Presented in Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy in the Graduate School of The Ohio State University By Julie Wallace Graduate Program in Molecular, Cellular and Developmental Biology The Ohio State University 2013 Dissertation Committee: Michael C. Ostrowski, PhD, Advisor Gustavo Leone, PhD Denis Guttridge, PhD Dawn Chandler, PhD Copyright by Julie Wallace 2013 Abstract Breast cancer is the second most common cancer in American women, and is also the second leading cause of cancer death in women. It is estimated that nearly a quarter of a million new cases of invasive breast cancer will be diagnosed in women in the United States this year, and approximately 40,000 of these women will die from breast cancer. Although death rates have been on the decline for the past decade, there is still much we need to learn about this disease to improve prevention, detection and treatment strategies. The majority of early studies have focused on the malignant tumor cells themselves, and much has been learned concerning mutations, amplifications and other genetic and epigenetic alterations of these cells. However more recent work has acknowledged the strong influence of tumor stroma on the initiation, progression and recurrence of cancer. Under normal conditions this stroma has been shown to have protective effects against tumorigenesis, however the transformation of tumor cells manipulates this surrounding environment to actually promote malignancy. Fibroblasts in particular make up a significant portion of this stroma, and have been shown to impact various aspects of tumor cell biology. -

Targeting and Reprograming Cancer-Associated Fibroblasts and the Tumor Microenvironment in Pancreatic Cancer
cancers Review Targeting and Reprograming Cancer-Associated Fibroblasts and the Tumor Microenvironment in Pancreatic Cancer Yoshiaki Sunami * , Viktoria Böker and Jörg Kleeff Department of Visceral, Vascular and Endocrine Surgery, Martin-Luther-University Halle-Wittenberg, University Medical Center Halle, 06120 Halle, Germany; [email protected] (V.B.); [email protected] (J.K.) * Correspondence: [email protected]; Tel.: +49-345-557-2794 Simple Summary: The tumor microenvironment plays a major role in the progression and drug resistance of pancreatic cancer. Cancer-associated fibroblasts are the major stromal cells and source of extracellular matrix proteins forming the dense stromal tumor microenvironment. Targeting cancer-associated fibroblasts has been deemed a promising therapeutic strategy. However, deplet- ing cancer-associated fibroblasts may also have tumor-promoting effects due to their functional heterogeneity. It is therefore important to target selectively the tumor-promoting subtype of cancer- associated fibroblasts. Furthermore, deactivating fibroblasts, or reprograming tumor-promoting cancer-associated fibroblasts to tumor-restraining cancer-associated fibroblasts are considered as therapy for pancreatic cancer. Abstract: Pancreatic cancer is the fourth leading cause of cancer deaths in the United States both in female and male, and is projected to become the second deadliest cancer by 2030. The overall five-year survival rate remains at around 10%. Pancreatic cancer exhibits a remarkable resistance to established therapeutic options such as chemotherapy and radiotherapy, due to dense stromal Citation: Sunami, Y.; Böker, V.; tumor microenvironment. Cancer-associated fibroblasts are the major stromal cell type and source of Kleeff, J. Targeting and Reprograming extracellular matrix proteins shaping a physical and metabolic barrier thereby reducing therapeutic Cancer-Associated Fibroblasts and efficacy. -

Genome-Wide Prediction of Small Molecule Binding to Remote
bioRxiv preprint doi: https://doi.org/10.1101/2020.08.04.236729; this version posted August 5, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Genome-wide Prediction of Small Molecule Binding 2 to Remote Orphan Proteins Using Distilled Sequence 3 Alignment Embedding 1 2 3 4 4 Tian Cai , Hansaim Lim , Kyra Alyssa Abbu , Yue Qiu , 5,6 1,2,3,4,7,* 5 Ruth Nussinov , and Lei Xie 1 6 Ph.D. Program in Computer Science, The Graduate Center, The City University of New York, New York, 10016, USA 2 7 Ph.D. Program in Biochemistry, The Graduate Center, The City University of New York, New York, 10016, USA 3 8 Department of Computer Science, Hunter College, The City University of New York, New York, 10065, USA 4 9 Ph.D. Program in Biology, The Graduate Center, The City University of New York, New York, 10016, USA 5 10 Computational Structural Biology Section, Basic Science Program, Frederick National Laboratory for Cancer Research, 11 Frederick, MD 21702, USA 6 12 Department of Human Molecular Genetics and Biochemistry, Sackler School of Medicine, Tel Aviv University, Tel 13 Aviv, Israel 7 14 Helen and Robert Appel Alzheimer’s Disease Research Institute, Feil Family Brain & Mind Research Institute, Weill 15 Cornell Medicine, Cornell University, New York, 10021, USA * 16 [email protected] 17 July 27, 2020 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.08.04.236729; this version posted August 5, 2020. -

Edinburgh Research Explorer
Edinburgh Research Explorer International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list Citation for published version: Davenport, AP, Alexander, SPH, Sharman, JL, Pawson, AJ, Benson, HE, Monaghan, AE, Liew, WC, Mpamhanga, CP, Bonner, TI, Neubig, RR, Pin, JP, Spedding, M & Harmar, AJ 2013, 'International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list: recommendations for new pairings with cognate ligands', Pharmacological reviews, vol. 65, no. 3, pp. 967-86. https://doi.org/10.1124/pr.112.007179 Digital Object Identifier (DOI): 10.1124/pr.112.007179 Link: Link to publication record in Edinburgh Research Explorer Document Version: Publisher's PDF, also known as Version of record Published In: Pharmacological reviews Publisher Rights Statement: U.S. Government work not protected by U.S. copyright General rights Copyright for the publications made accessible via the Edinburgh Research Explorer is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The University of Edinburgh has made every reasonable effort to ensure that Edinburgh Research Explorer content complies with UK legislation. If you believe that the public display of this file breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 02. Oct. 2021 1521-0081/65/3/967–986$25.00 http://dx.doi.org/10.1124/pr.112.007179 PHARMACOLOGICAL REVIEWS Pharmacol Rev 65:967–986, July 2013 U.S. -

Current Understanding of Epigenetics Mechanism As a Novel Target In
Keyvani‑Ghamsari et al. Clin Epigenet (2021) 13:120 https://doi.org/10.1186/s13148‑021‑01107‑4 REVIEW Open Access Current understanding of epigenetics mechanism as a novel target in reducing cancer stem cells resistance Saeedeh Keyvani‑Ghamsari1, Khatereh Khorsandi2* , Azhar Rasul3 and Muhammad Khatir Zaman4 Abstract At present, after extensive studies in the feld of cancer, cancer stem cells (CSCs) have been proposed as a major fac‑ tor in tumor initiation, progression, metastasis, and recurrence. CSCs are a subpopulation of bulk tumors, with stem cell‑like properties and tumorigenic capabilities, having the abilities of self‑renewal and diferentiation, thereby being able to generate heterogeneous lineages of cancer cells and lead to resistance toward anti‑tumor treatments. Highly resistant to conventional chemo‑ and radiotherapy, CSCs have heterogeneity and can migrate to diferent organs and metastasize. Recent studies have demonstrated that the population of CSCs and the progression of cancer are increased by the deregulation of diferent epigenetic pathways having efects on gene expression patterns and key pathways connected with cell proliferation and survival. Further, epigenetic modifcations (DNA methylation, histone modifcations, and RNA methylations) have been revealed to be key drivers in the formation and maintenance of CSCs. Hence, identifying CSCs and targeting epigenetic pathways therein can ofer new insights into the treatment of cancer. In the present review, recent studies are addressed in terms of the characteristics of CSCs, the resistance thereof, and the factors infuencing the development thereof, with an emphasis on diferent types of epigenetic changes in genes and main signaling pathways involved therein. Finally, targeted therapy for CSCs by epigenetic drugs is referred to, which is a new approach in overcoming resistance and recurrence of cancer. -

Rabbit Anti-FFAR3/GPR42/FITC Conjugated Antibody-SL16076R
SunLong Biotech Co.,LTD Tel: 0086-571- 56623320 Fax:0086-571- 56623318 E-mail:[email protected] www.sunlongbiotech.com Rabbit Anti-FFAR3/GPR42/FITC Conjugated antibody SL16076R-FITC Product Name: Anti-FFAR3/GPR42/FITC Chinese Name: FITC标记的G protein-coupled receptor42抗体 FFA3R; Ffar3; FFAR3_HUMAN; Free fatty acid receptor 3; G protein coupled Alias: receptor 41; G-protein coupled receptor 41; gpcr41; GPR41; gpr42. Organism Species: Rabbit Clonality: Polyclonal React Species: Human, ICC=1:50-200IF=1:50-200 Applications: not yet tested in other applications. optimal dilutions/concentrations should be determined by the end user. Molecular weight: 39kDa Form: Lyophilized or Liquid Concentration: 1mg/ml immunogen: KLH conjugated synthetic peptide derived from human FFAR3 Lsotype: IgG Purification: affinitywww.sunlongbiotech.com purified by Protein A Storage Buffer: 0.01M TBS(pH7.4) with 1% BSA, 0.03% Proclin300 and 50% Glycerol. Store at -20 °C for one year. Avoid repeated freeze/thaw cycles. The lyophilized antibody is stable at room temperature for at least one month and for greater than a year Storage: when kept at -20°C. When reconstituted in sterile pH 7.4 0.01M PBS or diluent of antibody the antibody is stable for at least two weeks at 2-4 °C. background: G protein-coupled receptors (GPRs), also known as seven transmembrane receptors, heptahelical receptors or 7TM receptors, comprise a superfamily of proteins that play a Product Detail: role in many different stimulus-response pathways. GPRs translate extracellular signals into intracellular signals (a process called G-protein activation) and they respond to a variety of signaling molecules, such as hormones and neurotransmitters. -

G Protein-Coupled Receptors
S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2015/16: G protein-coupled receptors. British Journal of Pharmacology (2015) 172, 5744–5869 THE CONCISE GUIDE TO PHARMACOLOGY 2015/16: G protein-coupled receptors Stephen PH Alexander1, Anthony P Davenport2, Eamonn Kelly3, Neil Marrion3, John A Peters4, Helen E Benson5, Elena Faccenda5, Adam J Pawson5, Joanna L Sharman5, Christopher Southan5, Jamie A Davies5 and CGTP Collaborators 1School of Biomedical Sciences, University of Nottingham Medical School, Nottingham, NG7 2UH, UK, 2Clinical Pharmacology Unit, University of Cambridge, Cambridge, CB2 0QQ, UK, 3School of Physiology and Pharmacology, University of Bristol, Bristol, BS8 1TD, UK, 4Neuroscience Division, Medical Education Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, DD1 9SY, UK, 5Centre for Integrative Physiology, University of Edinburgh, Edinburgh, EH8 9XD, UK Abstract The Concise Guide to PHARMACOLOGY 2015/16 provides concise overviews of the key properties of over 1750 human drug targets with their pharmacology, plus links to an open access knowledgebase of drug targets and their ligands (www.guidetopharmacology.org), which provides more detailed views of target and ligand properties. The full contents can be found at http://onlinelibrary.wiley.com/doi/ 10.1111/bph.13348/full. G protein-coupled receptors are one of the eight major pharmacological targets into which the Guide is divided, with the others being: ligand-gated ion channels, voltage-gated ion channels, other ion channels, nuclear hormone receptors, catalytic receptors, enzymes and transporters. These are presented with nomenclature guidance and summary information on the best available pharmacological tools, alongside key references and suggestions for further reading. -

G Protein‐Coupled Receptors
S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2019/20: G protein-coupled receptors. British Journal of Pharmacology (2019) 176, S21–S141 THE CONCISE GUIDE TO PHARMACOLOGY 2019/20: G protein-coupled receptors Stephen PH Alexander1 , Arthur Christopoulos2 , Anthony P Davenport3 , Eamonn Kelly4, Alistair Mathie5 , John A Peters6 , Emma L Veale5 ,JaneFArmstrong7 , Elena Faccenda7 ,SimonDHarding7 ,AdamJPawson7 , Joanna L Sharman7 , Christopher Southan7 , Jamie A Davies7 and CGTP Collaborators 1School of Life Sciences, University of Nottingham Medical School, Nottingham, NG7 2UH, UK 2Monash Institute of Pharmaceutical Sciences and Department of Pharmacology, Monash University, Parkville, Victoria 3052, Australia 3Clinical Pharmacology Unit, University of Cambridge, Cambridge, CB2 0QQ, UK 4School of Physiology, Pharmacology and Neuroscience, University of Bristol, Bristol, BS8 1TD, UK 5Medway School of Pharmacy, The Universities of Greenwich and Kent at Medway, Anson Building, Central Avenue, Chatham Maritime, Chatham, Kent, ME4 4TB, UK 6Neuroscience Division, Medical Education Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, DD1 9SY, UK 7Centre for Discovery Brain Sciences, University of Edinburgh, Edinburgh, EH8 9XD, UK Abstract The Concise Guide to PHARMACOLOGY 2019/20 is the fourth in this series of biennial publications. The Concise Guide provides concise overviews of the key properties of nearly 1800 human drug targets with an emphasis on selective pharmacology (where available), plus links to the open access knowledgebase source of drug targets and their ligands (www.guidetopharmacology.org), which provides more detailed views of target and ligand properties. Although the Concise Guide represents approximately 400 pages, the material presented is substantially reduced compared to information and links presented on the website. -

Early During Myelomagenesis Alterations in DNA Methylation That
Myeloma Is Characterized by Stage-Specific Alterations in DNA Methylation That Occur Early during Myelomagenesis This information is current as Christoph J. Heuck, Jayesh Mehta, Tushar Bhagat, Krishna of September 23, 2021. Gundabolu, Yiting Yu, Shahper Khan, Grigoris Chrysofakis, Carolina Schinke, Joseph Tariman, Eric Vickrey, Natalie Pulliam, Sangeeta Nischal, Li Zhou, Sanchari Bhattacharyya, Richard Meagher, Caroline Hu, Shahina Maqbool, Masako Suzuki, Samir Parekh, Frederic Reu, Ulrich Steidl, John Greally, Amit Verma and Seema B. Downloaded from Singhal J Immunol 2013; 190:2966-2975; Prepublished online 13 February 2013; doi: 10.4049/jimmunol.1202493 http://www.jimmunol.org/content/190/6/2966 http://www.jimmunol.org/ Supplementary http://www.jimmunol.org/content/suppl/2013/02/13/jimmunol.120249 Material 3.DC1 References This article cites 38 articles, 15 of which you can access for free at: http://www.jimmunol.org/content/190/6/2966.full#ref-list-1 by guest on September 23, 2021 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. -

The Pharmacology and Function of Receptors for Short-Chain Fatty Acids
1521-0111/89/3/388–398$25.00 http://dx.doi.org/10.1124/mol.115.102301 MOLECULAR PHARMACOLOGY Mol Pharmacol 89:388–398, March 2016 Copyright ª 2016 by The American Society for Pharmacology and Experimental Therapeutics MINIREVIEW The Pharmacology and Function of Receptors for Short-Chain Fatty Acids Daniele Bolognini, Andrew B. Tobin, Graeme Milligan, and Catherine E. Moss Molecular Pharmacology Group, Institute of Molecular, Cell and Systems Biology, University of Glasgow, Glasgow, Scotland, Downloaded from United Kingdom (D.B., G.M.); and Medical Research Council Toxicology Unit, University of Leicester, Leicester, United Kingdom (A.B.T., C.E.M.) Received November 2, 2015; accepted December 29, 2015 ABSTRACT molpharm.aspetjournals.org Despite some blockbuster G protein–coupled receptor (GPCR) and butyrate, ligands that originate largely as fermentation by- drugs, only a small fraction (∼15%) of the more than 390 products of anaerobic bacteria in the gut. However, the presence nonodorant GPCRs have been successfully targeted by the of FFA2 and FFA3 on pancreatic b-cells, FFA3 on neurons, and pharmaceutical industry. One way that this issue might be FFA2 on leukocytes and adipocytes means that the biologic role of addressed is via translation of recent deorphanization programs these receptors likely extends beyond the widely accepted role of that have opened the prospect of extending the reach of new regulating peptide hormone release from enteroendocrine cells in medicine design to novel receptor types with potential therapeutic the gut. Here, we review the physiologic roles of FFA2 and FFA3, value. Prominent among these receptors are those that respond to the recent development and use of receptor-selective pharmaco- short-chain free fatty acids of carbon chain length 2–6. -

Br J Pharmacol
Supplementary Information Supplementary Table 1. List of polypeptide cell surface receptor and their cognate ligand genes. Supplementary Table 2. List of coding SNPs with high FST ( > 0.5) in human GPCR and their cognate ligand genes. Supplementary Table 3. List of coding SNPs with high FST ( > 0.5) in human nonGPCR receptor and ligand genes. Supplementary Table 4. List of genotyped SNPs from 44246961 to 44542055 on chromosome 17. The HapMap II dataset was analyzed using HaploView. The 101 SNPs included in LD plots of Supplementary Fig. 3 (A-C) are highlighted by a grey background. The 37 SNPs used in the haplotype analysis of Fig. 2C are indicated by red letters. SNPs that are linked with rs2291725 are indicated by bold red letters. Supplementary Table 5. Allele frequency of rs2291725 in the HGDP-CEPH populations. Frequencies of GIP103T and GIP103C alleles in each of the 52 populations from the seven geographical regions and the number of chromosomes analyzed for each population. 1 Supplementary Table 6. EC50 values for GIP receptor activation at four different time points after incubation with pooled human serum or pooled complement-preserved human serum (N=4). Supplementary Table 7. EC50 values for GIP receptor activation at three different time points after incubation with a recombinant DPP IV enzyme (N=4). Supplementary Fig. 1. Cumulative distribution function (CDF) plots for the FST of coding SNPs in human GPCRs and their cognate ligand genes (blue) and all other human genes (magenta). The FST was computed between three HapMap II populations (CEU, YRI, and ASN), and coding SNPs have been split into synonymous and nonsynonymous groups. -

Adenylyl Cyclase 2 Selectively Regulates IL-6 Expression in Human Bronchial Smooth Muscle Cells Amy Sue Bogard University of Tennessee Health Science Center
University of Tennessee Health Science Center UTHSC Digital Commons Theses and Dissertations (ETD) College of Graduate Health Sciences 12-2013 Adenylyl Cyclase 2 Selectively Regulates IL-6 Expression in Human Bronchial Smooth Muscle Cells Amy Sue Bogard University of Tennessee Health Science Center Follow this and additional works at: https://dc.uthsc.edu/dissertations Part of the Medical Cell Biology Commons, and the Medical Molecular Biology Commons Recommended Citation Bogard, Amy Sue , "Adenylyl Cyclase 2 Selectively Regulates IL-6 Expression in Human Bronchial Smooth Muscle Cells" (2013). Theses and Dissertations (ETD). Paper 330. http://dx.doi.org/10.21007/etd.cghs.2013.0029. This Dissertation is brought to you for free and open access by the College of Graduate Health Sciences at UTHSC Digital Commons. It has been accepted for inclusion in Theses and Dissertations (ETD) by an authorized administrator of UTHSC Digital Commons. For more information, please contact [email protected]. Adenylyl Cyclase 2 Selectively Regulates IL-6 Expression in Human Bronchial Smooth Muscle Cells Document Type Dissertation Degree Name Doctor of Philosophy (PhD) Program Biomedical Sciences Track Molecular Therapeutics and Cell Signaling Research Advisor Rennolds Ostrom, Ph.D. Committee Elizabeth Fitzpatrick, Ph.D. Edwards Park, Ph.D. Steven Tavalin, Ph.D. Christopher Waters, Ph.D. DOI 10.21007/etd.cghs.2013.0029 Comments Six month embargo expired June 2014 This dissertation is available at UTHSC Digital Commons: https://dc.uthsc.edu/dissertations/330 Adenylyl Cyclase 2 Selectively Regulates IL-6 Expression in Human Bronchial Smooth Muscle Cells A Dissertation Presented for The Graduate Studies Council The University of Tennessee Health Science Center In Partial Fulfillment Of the Requirements for the Degree Doctor of Philosophy From The University of Tennessee By Amy Sue Bogard December 2013 Copyright © 2013 by Amy Sue Bogard.