AKR1C1 As a Biomarker for Differentiating the Biological Effects of Combustible from Non-Combustible Tobacco Products
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Chuanxiong Rhizoma Compound on HIF-VEGF Pathway and Cerebral Ischemia-Reperfusion Injury’S Biological Network Based on Systematic Pharmacology
ORIGINAL RESEARCH published: 25 June 2021 doi: 10.3389/fphar.2021.601846 Exploring the Regulatory Mechanism of Hedysarum Multijugum Maxim.-Chuanxiong Rhizoma Compound on HIF-VEGF Pathway and Cerebral Ischemia-Reperfusion Injury’s Biological Network Based on Systematic Pharmacology Kailin Yang 1†, Liuting Zeng 1†, Anqi Ge 2†, Yi Chen 1†, Shanshan Wang 1†, Xiaofei Zhu 1,3† and Jinwen Ge 1,4* Edited by: 1 Takashi Sato, Key Laboratory of Hunan Province for Integrated Traditional Chinese and Western Medicine on Prevention and Treatment of 2 Tokyo University of Pharmacy and Life Cardio-Cerebral Diseases, Hunan University of Chinese Medicine, Changsha, China, Galactophore Department, The First 3 Sciences, Japan Hospital of Hunan University of Chinese Medicine, Changsha, China, School of Graduate, Central South University, Changsha, China, 4Shaoyang University, Shaoyang, China Reviewed by: Hui Zhao, Capital Medical University, China Background: Clinical research found that Hedysarum Multijugum Maxim.-Chuanxiong Maria Luisa Del Moral, fi University of Jaén, Spain Rhizoma Compound (HCC) has de nite curative effect on cerebral ischemic diseases, *Correspondence: such as ischemic stroke and cerebral ischemia-reperfusion injury (CIR). However, its Jinwen Ge mechanism for treating cerebral ischemia is still not fully explained. [email protected] †These authors share first authorship Methods: The traditional Chinese medicine related database were utilized to obtain the components of HCC. The Pharmmapper were used to predict HCC’s potential targets. Specialty section: The CIR genes were obtained from Genecards and OMIM and the protein-protein This article was submitted to interaction (PPI) data of HCC’s targets and IS genes were obtained from String Ethnopharmacology, a section of the journal database. -

Taylor Et Al REVISED MS28926-1.Pdf
1 The effect of micronutrient supplementation on growth and hepatic 2 metabolism in diploid and triploid Atlantic salmon (Salmo salar) parr 3 fed a low marine ingredient diet 4 5 John F. Taylor a*, Luisa M. Vera a, Christian De Santis a, Erik-Jan Lock b, Marit Espe b, 6 Kaja H. Skjærven b, Daniel Leeming c, Jorge del Pozo d, Jose Mota-Velasco e, Herve 7 Migaud a, Kristin Hamre b, Douglas R. Tocher a 8 9 a Institute of Aquaculture, University of Stirling, Stirling, FK9 4LA, UK 10 b Institute of Marine Research, PO box 1870 Nordnes, 5817 Bergen, Norway 11 c BioMar Ltd., Grangemouth, FK3 8UL, UK 12 d The Royal (Dick) School of Veterinary Studies, Edinburgh, EH25 9RG, UK 13 e Hendrix Genetics, Landcatch Natural Selection Ltd., Lochgilphead, PA31 8PE, UK 14 15 Running Title: Dietary micronutrient supplementation in Atlantic salmon 16 17 ms. has 31 pg.s, 4 figures, 9 tables, 4 suppl. files 18 19 Corresponding Author: 20 Dr John F. Taylor 21 Institute of Aquaculture, University of Stirling, Stirling, FK9 4LA, UK 22 Tel: +44-01786 467929 ; Fax: +44-01768 472133 23 [email protected] 24 Accepted refereed manuscript of: Taylor JF, Vera LM, De Santis C, Lock E, Espe M, Skjaerven KH, Leeming D, Del Pozo J, Mota-Velasco J, Migaud H, Hamre K & Tocher DR (2019) The effect of micronutrient supplementation on growth and hepatic metabolism in diploid and triploid Atlantic salmon (Salmo salar) parr fed a low marine ingredient diet. Comparative Biochemistry and Physiology. Part B, Biochemistry and Molecular Biology, 227, pp. -

The Identification of Human Aldo-Keto Reductase AKR7A2 As a Novel
Li et al. Cellular & Molecular Biology Letters (2016) 21:25 Cellular & Molecular DOI 10.1186/s11658-016-0026-9 Biology Letters SHORTCOMMUNICATION Open Access The identification of human aldo-keto reductase AKR7A2 as a novel cytoglobin- binding partner Xin Li, Shanshan Zou, Zhen Li, Gaotai Cai, Bohong Chen, Ping Wang and Wenqi Dong* * Correspondence: [email protected] Abstract Department of Biopharmaceutics, School of Laboratory Medicine and Cytoglobin (CYGB), a member of the globin family, is thought to protect cells from Biotechnology, Southern Medical reactive oxygen and nitrogen species and deal with hypoxic conditions and University, 1838 North Guangzhou oxidative stress. However, its molecular mechanisms of action are not clearly Avenue, Guangzhou 510515, China understood. Through immunoprecipitation combined with a two-dimensional electrophoresis–mass spectrometry assay, we identified a CYGB interactor: aldo-keto reductase family 7 member A2 (AKR7A2). The interaction was further confirmed using yeast two-hybrid and co-immunoprecipitation assays. Our results show that AKR7A2 physically interacts with CYGB. Keywords: CYGB, AKR7A2, Protein-protein interactions, Yeast two-hybrid assay, Co-immunoprecipitation, 2-DE, Oxidative stress Introduction Cytoglobin (CYGB), which is a member of the globin family, was discovered more than a decade ago in a proteomic screen of fibrotic liver [1]. It was originally named STAP (stellate activating protein). Human CYGB is a 190-amino acid, 21-kDa protein [2], encoded by a single copy gene mapped at the 17q25.3 chromosomal segment [3]. It has a compact helical conformation, giving it the ability to bind to heme, which allows reversible binding of gaseous, diatomic molecules, including oxygen (O2), nitric oxide (NO) and carbon monoxide (CO), just like hemoglobin (Hb), myoglobin (Mb) and neuroglobin (Ngb) [4]. -

Gene-Expression Signature Regulated by the KEAP1-NRF2-CUL3 Axis Is Associated with a Poor Prognosis in Head and Neck Squamous Cell Cancer Akhileshwar Namani1†, Md
Namani et al. BMC Cancer (2018) 18:46 DOI 10.1186/s12885-017-3907-z RESEARCH ARTICLE Open Access Gene-expression signature regulated by the KEAP1-NRF2-CUL3 axis is associated with a poor prognosis in head and neck squamous cell cancer Akhileshwar Namani1†, Md. Matiur Rahaman2†, Ming Chen2* and Xiuwen Tang1* Abstract Background: NRF2 is the key regulator of oxidative stress in normal cells and aberrant expression of the NRF2 pathway due to genetic alterations in the KEAP1 (Kelch-like ECH-associated protein 1)-NRF2 (nuclear factor erythroid 2 like 2)-CUL3 (cullin 3) axis leads to tumorigenesis and drug resistance in many cancers including head and neck squamous cell cancer (HNSCC). The main goal of this study was to identify specific genes regulated by the KEAP1-NRF2-CUL3 axis in HNSCC patients, to assess the prognostic value of this gene signature in different cohorts, and to reveal potential biomarkers. Methods: RNA-Seq V2 level 3 data from 279 tumor samples along with 37 adjacent normal samples from patients enrolled in the The Cancer Genome Atlas (TCGA)-HNSCC study were used to identify upregulated genes using two methods (altered KEAP1-NRF2-CUL3 versus normal, and altered KEAP1-NRF2-CUL3 versus wild-type). We then used a new approach to identify the combined gene signature by integrating both datasets and subsequently tested this signature in 4 independent HNSCC datasets to assess its prognostic value. In addition, functional annotation using the DAVID v6.8 database and protein-protein interaction (PPI) analysis using the STRING v10 databasewereperformedonthesignature. Results: A signature composed of a subset of 17 genes regulated by the KEAP1-NRF2-CUL3 axis was identified by overlapping both the upregulated genes of altered versus normal (251 genes) and altered versus wild-type (25 genes) datasets. -

Identification and Characterization of Genes That Control Fat
Claire D’Andre et al. Journal of Animal Science and Biotechnology 2013, 4:43 http://www.jasbsci.com/content/4/1/43 JOURNAL OF ANIMAL SCIENCE AND BIOTECHNOLOGY RESEARCH Open Access Identification and characterization of genes that control fat deposition in chickens Hirwa Claire D’Andre1,3*, Wallace Paul2, Xu Shen3, Xinzheng Jia3, Rong Zhang3, Liang Sun3 and Xiquan Zhang3 Abstract Background: Fat deposits in chickens contribute significantly to meat quality attributes such as juiciness, flavor, taste and other organoleptic properties. The quantity of fat deposited increases faster and earlier in the fast- growing chickens than in slow-growing chickens. In this study, Affymetrix Genechip® Chicken Genome Arrays 32773 transcripts were used to compare gene expression profiles in liver and hypothalamus tissues of fast-growing and slow-growing chicken at 8 wk of age. Real-time RT-PCR was used to validate the differential expression of genes selected from the microarray analysis. The mRNA expression of the genes was further examined in fat tissues. The association of single nucleotide polymorphisms of four lipid-related genes with fat traits was examined in a F2 resource population. Results: Four hundred genes in the liver tissues and 220 genes hypothalamus tissues, respectively, were identified to be differentially expressed in fast-growing chickens and slow-growing chickens. Expression levels of genes for lipid metabolism (SULT1B1, ACSBG2, PNPLA3, LPL, AOAH) carbohydrate metabolism (MGAT4B, XYLB, GBE1, PGM1, HKDC1)cholesttrol biosynthesis (FDPS, LSS, HMGCR, NSDHL, DHCR24, IDI1, ME1) HSD17B7 and other reaction or pro- cesses (CYP1A4, CYP1A1, AKR1B1, CYP4V2, DDO) were higher in the fast-growing White Recessive Rock chickens than in the slow-growing Xinghua chickens. -

MMP-25 Metalloprotease Regulates Innate Immune Response Through NF- Κb Signaling Clara Soria-Valles, Ana Gutiérrez-Fernández, Fernando G
MMP-25 Metalloprotease Regulates Innate Immune Response through NF- κB Signaling Clara Soria-Valles, Ana Gutiérrez-Fernández, Fernando G. Osorio, Dido Carrero, Adolfo A. Ferrando, Enrique Colado, This information is current as M. Soledad Fernández-García, Elena Bonzon-Kulichenko, of September 27, 2021. Jesús Vázquez, Antonio Fueyo and Carlos López-Otín J Immunol 2016; 197:296-302; Prepublished online 3 June 2016; doi: 10.4049/jimmunol.1600094 Downloaded from http://www.jimmunol.org/content/197/1/296 Supplementary http://www.jimmunol.org/content/suppl/2016/06/01/jimmunol.160009 Material 4.DCSupplemental http://www.jimmunol.org/ References This article cites 41 articles, 15 of which you can access for free at: http://www.jimmunol.org/content/197/1/296.full#ref-list-1 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision by guest on September 27, 2021 • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology MMP-25 Metalloprotease Regulates Innate Immune Response through NF-kB Signaling Clara Soria-Valles,* Ana Gutie´rrez-Ferna´ndez,* Fernando G. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Functional Role of Pentose Phosphate Pathway and Glutamine in Cancer Cell
Functional Role of Pentose Phosphate Pathway and Glutamine in Cancer Cell Ibrahim Halil Polat ADVERTIMENT. La consulta d’aquesta tesi queda condicionada a l’acceptació de les següents condicions d'ús: La difusió d’aquesta tesi per mitjà del servei TDX (www.tdx.cat) i a través del Dipòsit Digital de la UB (diposit.ub.edu) ha estat autoritzada pels titulars dels drets de propietat intel·lectual únicament per a usos privats emmarcats en activitats d’investigació i docència. No s’autoritza la seva reproducció amb finalitats de lucre ni la seva difusió i posada a disposició des d’un lloc aliè al servei TDX ni al Dipòsit Digital de la UB. No s’autoritza la presentació del seu contingut en una finestra o marc aliè a TDX o al Dipòsit Digital de la UB (framing). Aquesta reserva de drets afecta tant al resum de presentació de la tesi com als seus continguts. En la utilització o cita de parts de la tesi és obligat indicar el nom de la persona autora. ADVERTENCIA. La consulta de esta tesis queda condicionada a la aceptación de las siguientes condiciones de uso: La difusión de esta tesis por medio del servicio TDR (www.tdx.cat) y a través del Repositorio Digital de la UB (diposit.ub.edu) ha sido autorizada por los titulares de los derechos de propiedad intelectual únicamente para usos privados enmarcados en actividades de investigación y docencia. No se autoriza su reproducción con finalidades de lucro ni su difusión y puesta a disposición desde un sitio ajeno al servicio TDR o al Repositorio Digital de la UB. -

Development of Potent and Selective Inhibitors of Aldo−Keto Reductase
Article pubs.acs.org/jmc Development of Potent and Selective Inhibitors of Aldo−Keto Reductase 1C3 (Type 5 17β-Hydroxysteroid Dehydrogenase) Based on N-Phenyl-Aminobenzoates and Their Structure−Activity Relationships † § ‡ § † † † ‡ Adegoke O. Adeniji, , Barry M. Twenter, , Michael C. Byrns, Yi Jin, Mo Chen, Jeffrey D. Winkler,*, † and Trevor M. Penning*, † Department of Pharmacology and Center of Excellence in Environmental Toxicology, Perelman School of Medicine, University of Pennsylvania, 130C John Morgan Building, 3620 Hamilton Walk, Philadelphia, Pennsylvania 19104-6084, United States ‡ Department of Chemistry, University of Pennsylvania, 231 South 34th Street, Philadelphia, Pennsylvania 19104, United States *S Supporting Information ABSTRACT: Aldo−keto reductase 1C3 (AKR1C3; type 5 17β-hydroxy- steroid dehydrogenase) is overexpressed in castration resistant prostate cancer (CRPC) and is implicated in the intratumoral biosynthesis of testosterone and 5α-dihydrotestosterone. Selective AKR1C3 inhibitors are required because compounds should not inhibit the highly related AKR1C1 and AKR1C2 isoforms which are involved in the inactivation of 5α-dihydrotestosterone. NSAIDs, N-phenylanthranilates in particular, are potent but nonselective AKR1C3 inhibitors. Using flufenamic acid, 2-{[3-(trifluoromethyl)phenyl]amino}- benzoic acid, as lead compound, five classes of structural analogues were synthesized and evaluated for AKR1C3 inhibitory potency and selectivity. Structure−activity relationship (SAR) studies revealed that a meta-carboxylic acid group relative to the amine conferred pronounced AKR1C3 selectivity without loss of potency, while electron withdrawing groups on the phenylamino B-ring were optimal for AKR1C3 inhibition. Lead compounds did not inhibit COX-1 or COX-2 but blocked the AKR1C3 mediated production of testosterone in LNCaP-AKR1C3 cells. These compounds offer promising leads toward new therapeutics for CRPC. -

Bioinformatic and Modelling Approaches for a System-Level Understanding of Oxidative Stress Toxicity Elias Zgheib
Bioinformatic and modelling approaches for a system-level understanding of oxidative stress toxicity Elias Zgheib To cite this version: Elias Zgheib. Bioinformatic and modelling approaches for a system-level understanding of oxidative stress toxicity. Quantitative Methods [q-bio.QM]. Université de Technologie de Compiègne, 2018. English. NNT : 2018COMP2464. tel-02088169 HAL Id: tel-02088169 https://tel.archives-ouvertes.fr/tel-02088169 Submitted on 2 Apr 2019 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Par Elias ZGHEIB Bioinformatic and modelling approaches for a system- level understanding of oxidative stress toxicity Thèse présentée pour l’obtention du grade de Docteur de l’UTC Soutenue le 18 décembre 2018 Spécialité : Bio-ingénierie et Mathématiques Appliquées : Unité de Recherche Biomécanique et Bio-ingénierie (UMR-7338) D2464 BIOINFORMATIC AND MODELLING APPROACHES FOR A SYSTEM-LEVEL UNDERSTANDING OF OXIDATIVE STRESS TOXICITY A THESIS SUBMITTED TO THE UNIVERSITE DE TECHNOLOGIE DE COMPIEGNE SORBONNE UNIVERSITES LABORATOIRE DE BIO-MECANIQUE ET BIOINGENIERIE UMR CNRS 7338 – BMBI 18TH OF DECEMBER 2018 For the degree of Doctor Spécialité : Bio-ingénierie et Mathématiques Appliquées Elias ZGHEIB SUPERVISED BY Prof. Frédéric Y. BOIS JURY MEMBERS Mme. Karine AUDOUZE Rapporteur Mr. -

Transcriptome Analysis of Different Multidrug-Resistant Gastric Carcinoma Cells
in vivo 19: 583-590 (2005) Transcriptome Analysis of Different Multidrug-resistant Gastric Carcinoma Cells STEFFEN HEIM1 and HERMANN LAGE2 1Europroteome AG, Berlin-Hennigsdorf; 2Institute of Pathology, Charité Campus Mitte, Schumannstr. 20/21, D-10117 Berlin, Germany Abstract. Multidrug resistance (MDR) of human cancers is advanced cancer. These therapeutic protocols produce an the major cause of failure of chemotherapy. To better overall response rate of about 20% at best using a single- understand the molecular events associated with the agent, and up to 50% using combination chemotherapy. development of different types of MDR, two different multidrug- Thus, a large number of malignancies are incurable. resistant gastric carcinoma cell lines, the MDR1/P-glycoprotein- Generally, gastrointestinal cancers, including gastric expressing cell line EPG85-257RDB and the MDR1/P- carcinoma, are naturally resistant to many anticancer drugs. glycoprotein-negative cell variant EPG85-257RNOV, as well as Additionally, these tumors are able to develop acquired the corresponding drug-sensitive parental cell line EPG85-257P, drug resistance phenotypes, which include the multidrug were used for analyses of the mRNA expression profiles by resistance (MDR) phenomenon. cDNA array hybridization. Of more than 12,000 genes spotted The MDR phenotype is characterized by simultaneous on the arrays, 156 genes were detected as being significantly resistance of tumor cells to various antineoplastic agents regulated in the cell line EPG85-257RDB in comparison to the which are structurally and functionally unrelated. Besides non-resistant cell variant, and 61 genes were found to be the classical MDR phenotype, mediated by the enhanced differentially expressed in the cell line EPG85-257RNOV. -
Data Supplement
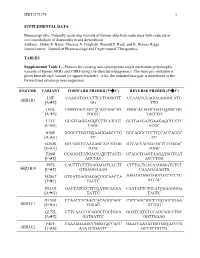
JPET#173179 1 SUPPLEMENTAL DATA Manuscript title: Naturally occurring variants of human aldo-keto reductases with reduced in vitro metabolism of daunorubicin and doxorubicin Authors: Onkar S. Bains, Thomas A. Grigliatti, Ronald E. Reid, and K. Wayne Riggs Journal name: Journal of Pharmacology and Experimental Therapeutics TABLES Supplemental Table 1—Primers for creating non-synonymous single nucleotide polymorphic variants of human AKRs and CBR4 using site-directed mutagenesis. The base pair mutation is given beneath each variant [in square brackets]. Also, the mutated base pair is underlined in the forward and reverse primer sequences. ENZYME VARIANT FORWARD PRIMER (5'3') REVERSE PRIMER (5'3') I15F CAAGATGCCCTTCCTGGGGTT CCAACCCCAGGAAGGGCATC AKR1B1 [AT] GG TTG H42L CGGGTACCGCCTCATCGACTG GGGCACAGTCGATGAGGCGG [AT] TGCCC TACCCG L73V GCGTGAGGAGGTCTTCATCGT GCTGACGATGAAGACCTCCTC [CG] CAGC ACGC K90E GGGCCTGGTGGAAGGAGCCTG GGCAGGCTCCTTCCACCAGGC [AG] CC CC G204S GCCAGTCCAAAAGCATCGTGG GTCACCACGATGCTTTTGGAC [GA] TGAC TGGC T288I CCAGGATATGACCATCTTACTC GTAGCTGAGTAAGATGGTCAT [CT] AGCTAC ATCCTGG P87S CACTTTCTTTGAGAGATCCCTT CTTTCCTCACAAGGGATCTCT AKR1B10 [CT] GTGAGGAAAG CAAAGAAAGTG M286T GTGATGAGGAGACGGCAACCA GAGTATGGTTGCCGTCTCCTC [TC] TACTC ATCAC N313D GACTATCCCTTCGATGCAGAA CAATATTCTGCATCGAAGGGA [AG] TATTG TAGTC R170H CCAACTTCAACCACAGGCAGC CTCCAGCTGCCTGTGGTTGAA AKR1C1 [GA] TGGAG GTTGG Q172L CTTCAACCGCAGGCTGCTGGA GGATCATCTCCAGCAGCCTGC [AT] GATGATCC GGTTGAAG F46Y CAATAGAAGCCGGGTACCACC GAATCAATATGGTGGTACCCG AKR1C2 [TA] ATATTTGATTC GCTTCTATTG JPET#173179