Transcriptome Analysis of Different Multidrug-Resistant Gastric Carcinoma Cells
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Molecular and Physiological Basis for Hair Loss in Near Naked Hairless and Oak Ridge Rhino-Like Mouse Models: Tracking the Role of the Hairless Gene
University of Tennessee, Knoxville TRACE: Tennessee Research and Creative Exchange Doctoral Dissertations Graduate School 5-2006 Molecular and Physiological Basis for Hair Loss in Near Naked Hairless and Oak Ridge Rhino-like Mouse Models: Tracking the Role of the Hairless Gene Yutao Liu University of Tennessee - Knoxville Follow this and additional works at: https://trace.tennessee.edu/utk_graddiss Part of the Life Sciences Commons Recommended Citation Liu, Yutao, "Molecular and Physiological Basis for Hair Loss in Near Naked Hairless and Oak Ridge Rhino- like Mouse Models: Tracking the Role of the Hairless Gene. " PhD diss., University of Tennessee, 2006. https://trace.tennessee.edu/utk_graddiss/1824 This Dissertation is brought to you for free and open access by the Graduate School at TRACE: Tennessee Research and Creative Exchange. It has been accepted for inclusion in Doctoral Dissertations by an authorized administrator of TRACE: Tennessee Research and Creative Exchange. For more information, please contact [email protected]. To the Graduate Council: I am submitting herewith a dissertation written by Yutao Liu entitled "Molecular and Physiological Basis for Hair Loss in Near Naked Hairless and Oak Ridge Rhino-like Mouse Models: Tracking the Role of the Hairless Gene." I have examined the final electronic copy of this dissertation for form and content and recommend that it be accepted in partial fulfillment of the requirements for the degree of Doctor of Philosophy, with a major in Life Sciences. Brynn H. Voy, Major Professor We have read this dissertation and recommend its acceptance: Naima Moustaid-Moussa, Yisong Wang, Rogert Hettich Accepted for the Council: Carolyn R. -

BTG2: a Rising Star of Tumor Suppressors (Review)
INTERNATIONAL JOURNAL OF ONCOLOGY 46: 459-464, 2015 BTG2: A rising star of tumor suppressors (Review) BIjING MAO1, ZHIMIN ZHANG1,2 and GE WANG1 1Cancer Center, Institute of Surgical Research, Daping Hospital, Third Military Medical University, Chongqing 400042; 2Department of Oncology, Wuhan General Hospital of Guangzhou Command, People's Liberation Army, Wuhan, Hubei 430070, P.R. China Received September 22, 2014; Accepted November 3, 2014 DOI: 10.3892/ijo.2014.2765 Abstract. B-cell translocation gene 2 (BTG2), the first 1. Discovery of BTG2 in TOB/BTG gene family gene identified in the BTG/TOB gene family, is involved in many biological activities in cancer cells acting as a tumor The TOB/BTG genes belong to the anti-proliferative gene suppressor. The BTG2 expression is downregulated in many family that includes six different genes in vertebrates: TOB1, human cancers. It is an instantaneous early response gene and TOB2, BTG1 BTG2/TIS21/PC3, BTG3 and BTG4 (Fig. 1). plays important roles in cell differentiation, proliferation, DNA The conserved domain of BTG N-terminal contains two damage repair, and apoptosis in cancer cells. Moreover, BTG2 regions, named box A and box B, which show a high level of is regulated by many factors involving different signal path- homology to the other domains (1-5). Box A has a major effect ways. However, the regulatory mechanism of BTG2 is largely on cell proliferation, while box B plays a role in combination unknown. Recently, the relationship between microRNAs and with many target molecules. Compared with other family BTG2 has attracted much attention. MicroRNA-21 (miR-21) members, BTG1 and BTG2 have an additional region named has been found to regulate BTG2 gene during carcinogenesis. -

Chuanxiong Rhizoma Compound on HIF-VEGF Pathway and Cerebral Ischemia-Reperfusion Injury’S Biological Network Based on Systematic Pharmacology
ORIGINAL RESEARCH published: 25 June 2021 doi: 10.3389/fphar.2021.601846 Exploring the Regulatory Mechanism of Hedysarum Multijugum Maxim.-Chuanxiong Rhizoma Compound on HIF-VEGF Pathway and Cerebral Ischemia-Reperfusion Injury’s Biological Network Based on Systematic Pharmacology Kailin Yang 1†, Liuting Zeng 1†, Anqi Ge 2†, Yi Chen 1†, Shanshan Wang 1†, Xiaofei Zhu 1,3† and Jinwen Ge 1,4* Edited by: 1 Takashi Sato, Key Laboratory of Hunan Province for Integrated Traditional Chinese and Western Medicine on Prevention and Treatment of 2 Tokyo University of Pharmacy and Life Cardio-Cerebral Diseases, Hunan University of Chinese Medicine, Changsha, China, Galactophore Department, The First 3 Sciences, Japan Hospital of Hunan University of Chinese Medicine, Changsha, China, School of Graduate, Central South University, Changsha, China, 4Shaoyang University, Shaoyang, China Reviewed by: Hui Zhao, Capital Medical University, China Background: Clinical research found that Hedysarum Multijugum Maxim.-Chuanxiong Maria Luisa Del Moral, fi University of Jaén, Spain Rhizoma Compound (HCC) has de nite curative effect on cerebral ischemic diseases, *Correspondence: such as ischemic stroke and cerebral ischemia-reperfusion injury (CIR). However, its Jinwen Ge mechanism for treating cerebral ischemia is still not fully explained. [email protected] †These authors share first authorship Methods: The traditional Chinese medicine related database were utilized to obtain the components of HCC. The Pharmmapper were used to predict HCC’s potential targets. Specialty section: The CIR genes were obtained from Genecards and OMIM and the protein-protein This article was submitted to interaction (PPI) data of HCC’s targets and IS genes were obtained from String Ethnopharmacology, a section of the journal database. -

Pinin (PNN) Rabbit Polyclonal Antibody – TA337687 | Origene
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for TA337687 Pinin (PNN) Rabbit Polyclonal Antibody Product data: Product Type: Primary Antibodies Applications: WB Recommended Dilution: WB Reactivity: Human Host: Rabbit Isotype: IgG Clonality: Polyclonal Immunogen: The immunogen for anti-PNN antibody: synthetic peptide directed towards the N terminal of human PNN. Synthetic peptide located within the following region: MAVAVRTLQEQLEKAKESLKNVDENIRKLTGRDPNDVRPIQARLLALSGP Formulation: Liquid. Purified antibody supplied in 1x PBS buffer with 0.09% (w/v) sodium azide and 2% sucrose. Note that this product is shipped as lyophilized powder to China customers. Purification: Affinity Purified Conjugation: Unconjugated Storage: Store at -20°C as received. Stability: Stable for 12 months from date of receipt. Predicted Protein Size: 81 kDa Gene Name: pinin, desmosome associated protein Database Link: NP_002678 Entrez Gene 5411 Human Q9H307 Background: PNN is the transcriptional activator binding to the E-box 1 core sequence of the E-cadherin promoter gene; the core-binding sequence is 5'CAGGTG-3'. PNN is capable of reversing CTBP1-mediated transcription repression. Component of a splicing-dependent mul Synonyms: DRS; DRSP; memA; SDK3 Note: Immunogen Sequence Homology: Dog: 100%; Pig: 100%; Rat: 100%; Human: 100%; Mouse: 100%; Bovine: 100%; Rabbit: 100%; Zebrafish: 100%; Guinea pig: 100% This product -

Original Article Long Noncoding RNA TOB1-AS1, an Epigenetically Silenced Gene, Functioned As a Novel Tumor Suppressor by Sponging Mir-27B in Cervical Cancer
Am J Cancer Res 2018;8(8):1483-1498 www.ajcr.us /ISSN:2156-6976/ajcr0079106 Original Article Long noncoding RNA TOB1-AS1, an epigenetically silenced gene, functioned as a novel tumor suppressor by sponging miR-27b in cervical cancer Jihang Yao1, Zhenghong Li1, Ziwei Yang2, Hui Xue1, Hua Chang1, Xue Zhang1, Tianren Li1, Kejun Guo1 Departments of 1Gynecology, 2Clinical Laboratory, The First Hospital of China Medical University, Shenyang 110001, Liaoning, China Received April 21, 2018; Accepted July 9, 2018; Epub August 1, 2018; Published August 15, 2018 Abstract: Cervical cancer is one of the most common cancers in females, accounting for a majority of cancer- related deaths in worldwide. Long non-coding RNAs (lncRNAs) have been identified as critical regulators in many tumor-related biological processes. Thus, investigation into the function and mechanism of lncRNAs in the develop- ment of cervical cancer is very necessary. In this study, we found that the expression of TOB1-AS1 was significantly decreased in cervical cancer tissues compared with the adjacent normal tissues. The methylation status of TOB1- AS1-related CpG island was analyzed using methylation specific PCR and bisulfite sequencing analysis, revealing that the aberrant hypermethylation of TOB1-AS1-related CpG island was frequently observed in primary tumors and cervical cancer cells. The expression of TOB1-AS1 in cervical cancer cells could be reversed by demethylation agent treatment. Functionally, overexpression of TOB1-AS1 significantly inhibited cell proliferation, cell cycle progression, invasion and induced apoptosis, while knockdown of TOB1-AS1 exhibited the opposite effect. Furthermore, it was determined that TOB1-AS1 was able to bind and degrade the expression of miR-27b. -

Identification and Characterization of Genes That Control Fat
Claire D’Andre et al. Journal of Animal Science and Biotechnology 2013, 4:43 http://www.jasbsci.com/content/4/1/43 JOURNAL OF ANIMAL SCIENCE AND BIOTECHNOLOGY RESEARCH Open Access Identification and characterization of genes that control fat deposition in chickens Hirwa Claire D’Andre1,3*, Wallace Paul2, Xu Shen3, Xinzheng Jia3, Rong Zhang3, Liang Sun3 and Xiquan Zhang3 Abstract Background: Fat deposits in chickens contribute significantly to meat quality attributes such as juiciness, flavor, taste and other organoleptic properties. The quantity of fat deposited increases faster and earlier in the fast- growing chickens than in slow-growing chickens. In this study, Affymetrix Genechip® Chicken Genome Arrays 32773 transcripts were used to compare gene expression profiles in liver and hypothalamus tissues of fast-growing and slow-growing chicken at 8 wk of age. Real-time RT-PCR was used to validate the differential expression of genes selected from the microarray analysis. The mRNA expression of the genes was further examined in fat tissues. The association of single nucleotide polymorphisms of four lipid-related genes with fat traits was examined in a F2 resource population. Results: Four hundred genes in the liver tissues and 220 genes hypothalamus tissues, respectively, were identified to be differentially expressed in fast-growing chickens and slow-growing chickens. Expression levels of genes for lipid metabolism (SULT1B1, ACSBG2, PNPLA3, LPL, AOAH) carbohydrate metabolism (MGAT4B, XYLB, GBE1, PGM1, HKDC1)cholesttrol biosynthesis (FDPS, LSS, HMGCR, NSDHL, DHCR24, IDI1, ME1) HSD17B7 and other reaction or pro- cesses (CYP1A4, CYP1A1, AKR1B1, CYP4V2, DDO) were higher in the fast-growing White Recessive Rock chickens than in the slow-growing Xinghua chickens. -

MMP-25 Metalloprotease Regulates Innate Immune Response Through NF- Κb Signaling Clara Soria-Valles, Ana Gutiérrez-Fernández, Fernando G
MMP-25 Metalloprotease Regulates Innate Immune Response through NF- κB Signaling Clara Soria-Valles, Ana Gutiérrez-Fernández, Fernando G. Osorio, Dido Carrero, Adolfo A. Ferrando, Enrique Colado, This information is current as M. Soledad Fernández-García, Elena Bonzon-Kulichenko, of September 27, 2021. Jesús Vázquez, Antonio Fueyo and Carlos López-Otín J Immunol 2016; 197:296-302; Prepublished online 3 June 2016; doi: 10.4049/jimmunol.1600094 Downloaded from http://www.jimmunol.org/content/197/1/296 Supplementary http://www.jimmunol.org/content/suppl/2016/06/01/jimmunol.160009 Material 4.DCSupplemental http://www.jimmunol.org/ References This article cites 41 articles, 15 of which you can access for free at: http://www.jimmunol.org/content/197/1/296.full#ref-list-1 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision by guest on September 27, 2021 • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology MMP-25 Metalloprotease Regulates Innate Immune Response through NF-kB Signaling Clara Soria-Valles,* Ana Gutie´rrez-Ferna´ndez,* Fernando G. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Exon Junction Complexes Can Have Distinct Functional Flavours To
www.nature.com/scientificreports OPEN Exon Junction Complexes can have distinct functional favours to regulate specifc splicing events Received: 9 November 2017 Zhen Wang1, Lionel Ballut2, Isabelle Barbosa1 & Hervé Le Hir1 Accepted: 11 June 2018 The exon junction complex (EJC) deposited on spliced mRNAs, plays a central role in the post- Published: xx xx xxxx transcriptional gene regulation and specifc gene expression. The EJC core complex is associated with multiple peripheral factors involved in various post-splicing events. Here, using recombinant complex reconstitution and transcriptome-wide analysis, we showed that the EJC peripheral protein complexes ASAP and PSAP form distinct complexes with the EJC core and can confer to EJCs distinct alternative splicing regulatory activities. This study provides the frst evidence that diferent EJCs can have distinct functions, illuminating EJC-dependent gene regulation. Te Exon Junction Complex (EJC) plays a central role in post-transcriptional gene expression control. EJCs tag mRNA exon junctions following intron removal by spliceosomes and accompany spliced mRNAs from the nucleus to the cytoplasm where they are displaced by the translating ribosomes1,2. Te EJC is organized around a core complex made of the proteins eIF4A3, MAGOH, Y14 and MLN51, and this EJC core serves as platforms for multiple peripheral factors during diferent post-transcriptional steps3,4. Dismantled during translation, EJCs mark a very precise period in mRNA life between nuclear splicing and cytoplasmic translation. In this window, EJCs contribute to alternative splicing5–7, intra-cellular RNA localization8, translation efciency9–11 and mRNA stability control by nonsense-mediated mRNA decay (NMD)12–14. At a physiological level, developmental defects and human pathological disorders due to altered expression of EJC proteins show that the EJC dosage is critical for specifc cell fate determinations, such as specifcation of embryonic body axis in drosophila, or Neural Stem Cells division in the mouse8,15,16. -
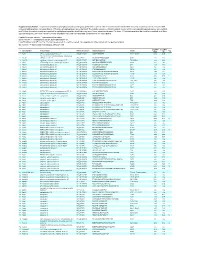
Supplementary Table 1
Supplementary Table 1. Large-scale quantitative phosphoproteomic profiling was performed on paired vehicle- and hormone-treated mTAL-enriched suspensions (n=3). A total of 654 unique phosphopeptides corresponding to 374 unique phosphoproteins were identified. The peptide sequence, phosphorylation site(s), and the corresponding protein name, gene symbol, and RefSeq Accession number are reported for each phosphopeptide identified in any one of three experimental pairs. For those 414 phosphopeptides that could be quantified in all three experimental pairs, the mean Hormone:Vehicle abundance ratio and corresponding standard error are also reported. Peptide Sequence column: * = phosphorylated residue Site(s) column: ^ = ambiguously assigned phosphorylation site Log2(H/V) Mean and SE columns: H = hormone-treated, V = vehicle-treated, n/a = peptide not observable in all 3 experimental pairs Sig. column: * = significantly changed Log 2(H/V), p<0.05 Log (H/V) Log (H/V) # Gene Symbol Protein Name Refseq Accession Peptide Sequence Site(s) 2 2 Sig. Mean SE 1 Aak1 AP2-associated protein kinase 1 NP_001166921 VGSLT*PPSS*PK T622^, S626^ 0.24 0.95 PREDICTED: ATP-binding cassette, sub-family A 2 Abca12 (ABC1), member 12 XP_237242 GLVQVLS*FFSQVQQQR S251^ 1.24 2.13 3 Abcc10 multidrug resistance-associated protein 7 NP_001101671 LMT*ELLS*GIRVLK T464, S468 -2.68 2.48 4 Abcf1 ATP-binding cassette sub-family F member 1 NP_001103353 QLSVPAS*DEEDEVPVPVPR S109 n/a n/a 5 Ablim1 actin-binding LIM protein 1 NP_001037859 PGSSIPGS*PGHTIYAK S51 -3.55 1.81 6 Ablim1 actin-binding -

Dissertation
Regulation of gene silencing: From microRNA biogenesis to post-translational modifications of TNRC6 complexes DISSERTATION zur Erlangung des DOKTORGRADES DER NATURWISSENSCHAFTEN (Dr. rer. nat.) der Fakultät Biologie und Vorklinische Medizin der Universität Regensburg vorgelegt von Johannes Danner aus Eggenfelden im Jahr 2017 Das Promotionsgesuch wurde eingereicht am: 12.09.2017 Die Arbeit wurde angeleitet von: Prof. Dr. Gunter Meister Johannes Danner Summary ‘From microRNA biogenesis to post-translational modifications of TNRC6 complexes’ summarizes the two main projects, beginning with the influence of specific RNA binding proteins on miRNA biogenesis processes. The fate of the mature miRNA is determined by the incorporation into Argonaute proteins followed by a complex formation with TNRC6 proteins as core molecules of gene silencing complexes. miRNAs are transcribed as stem-loop structured primary transcripts (pri-miRNA) by Pol II. The further nuclear processing is carried out by the microprocessor complex containing the RNase III enzyme Drosha, which cleaves the pri-miRNA to precursor-miRNA (pre-miRNA). After Exportin-5 mediated transport of the pre-miRNA to the cytoplasm, the RNase III enzyme Dicer cleaves off the terminal loop resulting in a 21-24 nt long double-stranded RNA. One of the strands is incorporated in the RNA-induced silencing complex (RISC), where it directly interacts with a member of the Argonaute protein family. The miRNA guides the mature RISC complex to partially complementary target sites on mRNAs leading to gene silencing. During this process TNRC6 proteins interact with Argonaute and recruit additional factors to mediate translational repression and target mRNA destabilization through deadenylation and decapping leading to mRNA decay. -

A Deficiency in the Region Homologous to Human 17Q21.33
Copyright Ó 2006 by the Genetics Society of America DOI: 10.1534/genetics.105.054833 A Deficiency in the Region Homologous to Human 17q21.33–q23.2 Causes Heart Defects in Mice Y. Eugene Yu,*,†,1,2 Masae Morishima,‡ Annie Pao,* Ding-Yan Wang,§ Xiao-Yan Wen,§ Antonio Baldini*,‡,** and Allan Bradley††,1 *Department of Molecular and Human Genetics, ‡Department of Pediatrics (Cardiology), **Center for Cardiovascular Development, Baylor College of Medicine, Houston, Texas 77030, †Department of Cancer Genetics and Genetics Program, Roswell Park Cancer Institute, Buffalo, New York 14263, §Division of Cellular and Molecular Biology, Toronto General Research Institute, University of Toronto, Toronto, Ontario M5G 2C1, Canada and ††Wellcome Trust Sanger Institute, Wellcome Trust Genome Campus, Hinxton, Cambridge CB10 1SA, United Kingdom Manuscript received December 17, 2005 Accepted for publication February 14, 2006 ABSTRACT Several constitutional chromosomal rearrangements occur on human chromosome 17. Patients who carry constitutional deletions of 17q21.3–q24 exhibit distinct phenotypic features. Within the deletion interval, there is a genomic segment that is bounded by the myeloperoxidase and homeobox B1 genes. This genomic segment is syntenically conserved on mouse chromosome 11 and is bounded by the mouse homologs of the same genes (Mpo and HoxB1). To attain functional information about this syntenic segment in mice, we have generated a 6.9-Mb deletion [Df(11)18], the reciprocal duplication [Dp(11)18] between Mpo and Chad (the chondroadherin gene), and a 1.8-Mb deletion between Chad and HoxB1. Phenotypic analyses of the mutant mouse lines showed that the Dp(11)18/Dp(11)18 genotype was responsible for embryonic or adolescent lethality, whereas the Df(11)18/1 genotype was responsible for heart defects.