Integrative Analysis of Promising Molecular Biomarkers and Pathways for Coronary Artery Disease Using WGCNA and Metade Methods
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

KO Kidney.Xlsx
Supplemental Table 18: Dietary Impact on the CGL KO Kidney Sulfhydrome DR/AL Accession Molecular Cysteine Spectral Protein Name Number Alternate ID Weight Residues Count Ratio P‐value Ig gamma‐2A chain C region, A allele P01863 (+1) Ighg 36 kDa 10 C 5.952 0.03767 Heterogeneous nuclear ribonucleoprotein M Q9D0E1 (+1) Hnrnpm 78 kDa 6 C 5.000 0.00595 Phospholipase D3 O35405 Pld3 54 kDa 8 C 4.167 0.04761 Ig kappa chain V‐V region L7 (Fragment) P01642 Gm10881 13 kDa 2 C 2.857 0.01232 UPF0160 protein MYG1, mitochondrial Q9JK81 Myg1 43 kDa 7 C 2.333 0.01613 Copper homeostasis protein cutC homolog Q9D8X1 Cutc 29 kDa 7 C 10.333 0.16419 Corticosteroid‐binding globulin Q06770 Serpina6 45 kDa 3 C 10.333 0.16419 28S ribosomal protein S22, mitochondrial Q9CXW2 Mrps22 41 kDa 2 C 7.333 0.3739 Isoform 3 of Agrin A2ASQ1‐3 Agrn 198 kDa 2 C 7.333 0.3739 3‐oxoacyl‐[acyl‐carrier‐protein] synthase, mitochondrial Q9D404 Oxsm 49 kDa 11 C 7.333 0.3739 Cordon‐bleu protein‐like 1 Q3UMF0 (+3)Cobll1 137 kDa 10 C 5.833 0.10658 ADP‐sugar pyrophosphatase Q9JKX6 Nudt5 24 kDa 5 C 4.167 0.15819 Complement C4‐B P01029 C4b 193 kDa 29 C 3.381 0.23959 Protein‐glutamine gamma‐glutamyltransferase 2 P21981 Tgm2 77 kDa 20 C 3.381 0.23959 Isochorismatase domain‐containing protein 1 Q91V64 Isoc1 32 kDa 5 C 3.333 0.10588 Serpin B8 O08800 Serpinb8 42 kDa 11 C 2.903 0.06902 Heterogeneous nuclear ribonucleoprotein A0 Q9CX86 Hnrnpa0 31 kDa 3 C 2.667 0.5461 Proteasome subunit beta type‐8 P28063 Psmb8 30 kDa 5 C 2.583 0.36848 Ig kappa chain V‐V region MOPC 149 P01636 12 kDa 2 C 2.583 0.36848 -

Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model
Downloaded from http://www.jimmunol.org/ by guest on September 25, 2021 T + is online at: average * The Journal of Immunology , 34 of which you can access for free at: 2016; 197:1477-1488; Prepublished online 1 July from submission to initial decision 4 weeks from acceptance to publication 2016; doi: 10.4049/jimmunol.1600589 http://www.jimmunol.org/content/197/4/1477 Molecular Profile of Tumor-Specific CD8 Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. Waugh, Sonia M. Leach, Brandon L. Moore, Tullia C. Bruno, Jonathan D. Buhrman and Jill E. Slansky J Immunol cites 95 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2016/07/01/jimmunol.160058 9.DCSupplemental This article http://www.jimmunol.org/content/197/4/1477.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 25, 2021. The Journal of Immunology Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. -

IMPDH2: a New Gene Associated with Dominant Juvenile-Onset Dystonia-Tremor Disorder
www.nature.com/ejhg BRIEF COMMUNICATION OPEN IMPDH2: a new gene associated with dominant juvenile-onset dystonia-tremor disorder 1,8 1,8 2 3 1,4 2 5 Anna Kuukasjärvi , Juan✉ C. Landoni , Jyrki Kaukonen , Mika Juhakoski , Mari Auranen , Tommi Torkkeli , Vidya Velagapudi and Anu Suomalainen 1,6,7 © The Author(s) 2021 The aetiology of dystonia disorders is complex, and next-generation sequencing has become a useful tool in elucidating the variable genetic background of these diseases. Here we report a deleterious heterozygous truncating variant in the inosine monophosphate dehydrogenasegene(IMPDH2) by whole-exome sequencing, co-segregating with a dominantly inherited dystonia-tremor disease in a large Finnish family. We show that the defect results in degradation of the gene product, causing IMPDH2 deficiency in patient cells. IMPDH2 is the first and rate-limiting enzyme in the de novo biosynthesis of guanine nucleotides, a dopamine synthetic pathway previously linked to childhood or adolescence-onset dystonia disorders. We report IMPDH2 as a new gene to the dystonia disease entity. The evidence underlines the important link between guanine metabolism, dopamine biosynthesis and dystonia. European Journal of Human Genetics; https://doi.org/10.1038/s41431-021-00939-1 INTRODUCTION The disease-onset was between 9 and 20 years of age. Table 1 Dystonias are rare movement disorders characterised by sustained or summarises the clinical presentations. intermittent muscle contractions causing abnormal, often repetitive, movements and/or postures. Dystonia can manifest as an isolated Case report symptom or combined with e.g. parkinsonism or myoclonus [1]. While Patient II-6 is a 46-year-old woman. -

Chuanxiong Rhizoma Compound on HIF-VEGF Pathway and Cerebral Ischemia-Reperfusion Injury’S Biological Network Based on Systematic Pharmacology
ORIGINAL RESEARCH published: 25 June 2021 doi: 10.3389/fphar.2021.601846 Exploring the Regulatory Mechanism of Hedysarum Multijugum Maxim.-Chuanxiong Rhizoma Compound on HIF-VEGF Pathway and Cerebral Ischemia-Reperfusion Injury’s Biological Network Based on Systematic Pharmacology Kailin Yang 1†, Liuting Zeng 1†, Anqi Ge 2†, Yi Chen 1†, Shanshan Wang 1†, Xiaofei Zhu 1,3† and Jinwen Ge 1,4* Edited by: 1 Takashi Sato, Key Laboratory of Hunan Province for Integrated Traditional Chinese and Western Medicine on Prevention and Treatment of 2 Tokyo University of Pharmacy and Life Cardio-Cerebral Diseases, Hunan University of Chinese Medicine, Changsha, China, Galactophore Department, The First 3 Sciences, Japan Hospital of Hunan University of Chinese Medicine, Changsha, China, School of Graduate, Central South University, Changsha, China, 4Shaoyang University, Shaoyang, China Reviewed by: Hui Zhao, Capital Medical University, China Background: Clinical research found that Hedysarum Multijugum Maxim.-Chuanxiong Maria Luisa Del Moral, fi University of Jaén, Spain Rhizoma Compound (HCC) has de nite curative effect on cerebral ischemic diseases, *Correspondence: such as ischemic stroke and cerebral ischemia-reperfusion injury (CIR). However, its Jinwen Ge mechanism for treating cerebral ischemia is still not fully explained. [email protected] †These authors share first authorship Methods: The traditional Chinese medicine related database were utilized to obtain the components of HCC. The Pharmmapper were used to predict HCC’s potential targets. Specialty section: The CIR genes were obtained from Genecards and OMIM and the protein-protein This article was submitted to interaction (PPI) data of HCC’s targets and IS genes were obtained from String Ethnopharmacology, a section of the journal database. -

Identification and Characterization of Genes That Control Fat
Claire D’Andre et al. Journal of Animal Science and Biotechnology 2013, 4:43 http://www.jasbsci.com/content/4/1/43 JOURNAL OF ANIMAL SCIENCE AND BIOTECHNOLOGY RESEARCH Open Access Identification and characterization of genes that control fat deposition in chickens Hirwa Claire D’Andre1,3*, Wallace Paul2, Xu Shen3, Xinzheng Jia3, Rong Zhang3, Liang Sun3 and Xiquan Zhang3 Abstract Background: Fat deposits in chickens contribute significantly to meat quality attributes such as juiciness, flavor, taste and other organoleptic properties. The quantity of fat deposited increases faster and earlier in the fast- growing chickens than in slow-growing chickens. In this study, Affymetrix Genechip® Chicken Genome Arrays 32773 transcripts were used to compare gene expression profiles in liver and hypothalamus tissues of fast-growing and slow-growing chicken at 8 wk of age. Real-time RT-PCR was used to validate the differential expression of genes selected from the microarray analysis. The mRNA expression of the genes was further examined in fat tissues. The association of single nucleotide polymorphisms of four lipid-related genes with fat traits was examined in a F2 resource population. Results: Four hundred genes in the liver tissues and 220 genes hypothalamus tissues, respectively, were identified to be differentially expressed in fast-growing chickens and slow-growing chickens. Expression levels of genes for lipid metabolism (SULT1B1, ACSBG2, PNPLA3, LPL, AOAH) carbohydrate metabolism (MGAT4B, XYLB, GBE1, PGM1, HKDC1)cholesttrol biosynthesis (FDPS, LSS, HMGCR, NSDHL, DHCR24, IDI1, ME1) HSD17B7 and other reaction or pro- cesses (CYP1A4, CYP1A1, AKR1B1, CYP4V2, DDO) were higher in the fast-growing White Recessive Rock chickens than in the slow-growing Xinghua chickens. -

MMP-25 Metalloprotease Regulates Innate Immune Response Through NF- Κb Signaling Clara Soria-Valles, Ana Gutiérrez-Fernández, Fernando G
MMP-25 Metalloprotease Regulates Innate Immune Response through NF- κB Signaling Clara Soria-Valles, Ana Gutiérrez-Fernández, Fernando G. Osorio, Dido Carrero, Adolfo A. Ferrando, Enrique Colado, This information is current as M. Soledad Fernández-García, Elena Bonzon-Kulichenko, of September 27, 2021. Jesús Vázquez, Antonio Fueyo and Carlos López-Otín J Immunol 2016; 197:296-302; Prepublished online 3 June 2016; doi: 10.4049/jimmunol.1600094 Downloaded from http://www.jimmunol.org/content/197/1/296 Supplementary http://www.jimmunol.org/content/suppl/2016/06/01/jimmunol.160009 Material 4.DCSupplemental http://www.jimmunol.org/ References This article cites 41 articles, 15 of which you can access for free at: http://www.jimmunol.org/content/197/1/296.full#ref-list-1 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision by guest on September 27, 2021 • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology MMP-25 Metalloprotease Regulates Innate Immune Response through NF-kB Signaling Clara Soria-Valles,* Ana Gutie´rrez-Ferna´ndez,* Fernando G. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -
Type of the Paper (Article
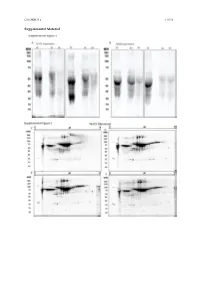
Cells 2020, 9, x 1 of 19 Supplemental Material Cells 2020, 9, x 2 of 19 Figure S1. Secretome enrichment: protocol optimization. 1D SDS-PAGE documentation of washing steps: Culture medium was substituted with FCS-free medium, which was changed every 2 h. The supernatants were then collected, and the proteins isolated and separated in 1D SDS-PAGE ((A) TK173 and (B) TK188). Proteins were stained with Flamingo fluorescent gel stain. Two-dimensional pattern of the proteins isolated from supernatant of TK173, (C) 2 h, (D) 4 h, (E) 6 h, and (F) 8 h after changing to FCS-free medium. (G) Cell secretome collected 24 h after elimination of the contaminating FCS-proteins with different washing steps. Proteins were stained with Flamingo fluorescent gel stain. Cells 2020, 9, x 3 of 19 Figure S2. 2-DE reference maps of secretomes; 150 μg proteins were loaded on an 11 cm IPG strip with a linear pH gradient PI 5–8 for IEF; 12% SDS-polyacrylamide gels were used for the second dimension. Proteins were stained with Flamingo fluorescent gel stain. Identified spots were assigned a number corresponding to that in their table. 2-DE maps from secretome of (A) TK173 control and (B) TGFβ1- treated ones. The 2-DE patterns revealed an alteration of secretome in stimulated TK173. Secretome patterns from TK173 treated with (C) ANG II and (D) PDGF. Cells 2020, 9, x 4 of 19 A Figure S3. Classification of the differentially expressed proteins upon ANG II, TGFβ1, or PDGF treatment in TK173. (A) Bar charts of the cellular component analyzed by STRAP biological function analysis in which the identified proteins from all treatments in both cell types are involved. -

Supplementary Methods
Supplementary methods Human lung tissues and tissue microarray (TMA) All human tissues were obtained from the Lung Cancer Specialized Program of Research Excellence (SPORE) Tissue Bank at the M.D. Anderson Cancer Center (Houston, TX). A collection of 26 lung adenocarcinomas and 24 non-tumoral paired tissues were snap-frozen and preserved in liquid nitrogen for total RNA extraction. For each tissue sample, the percentage of malignant tissue was calculated and the cellular composition of specimens was determined by histological examination (I.I.W.) following Hematoxylin-Eosin (H&E) staining. All malignant samples retained contained more than 50% tumor cells. Specimens resected from NSCLC stages I-IV patients who had no prior chemotherapy or radiotherapy were used for TMA analysis by immunohistochemistry. Patients who had smoked at least 100 cigarettes in their lifetime were defined as smokers. Samples were fixed in formalin, embedded in paraffin, stained with H&E, and reviewed by an experienced pathologist (I.I.W.). The 413 tissue specimens collected from 283 patients included 62 normal bronchial epithelia, 61 bronchial hyperplasias (Hyp), 15 squamous metaplasias (SqM), 9 squamous dysplasias (Dys), 26 carcinomas in situ (CIS), as well as 98 squamous cell carcinomas (SCC) and 141 adenocarcinomas. Normal bronchial epithelia, hyperplasia, squamous metaplasia, dysplasia, CIS, and SCC were considered to represent different steps in the development of SCCs. All tumors and lesions were classified according to the World Health Organization (WHO) 2004 criteria. The TMAs were prepared with a manual tissue arrayer (Advanced Tissue Arrayer ATA100, Chemicon International, Temecula, CA) using 1-mm-diameter cores in triplicate for tumors and 1.5 to 2-mm cores for normal epithelial and premalignant lesions. -

Development of Potent and Selective Inhibitors of Aldo−Keto Reductase
Article pubs.acs.org/jmc Development of Potent and Selective Inhibitors of Aldo−Keto Reductase 1C3 (Type 5 17β-Hydroxysteroid Dehydrogenase) Based on N-Phenyl-Aminobenzoates and Their Structure−Activity Relationships † § ‡ § † † † ‡ Adegoke O. Adeniji, , Barry M. Twenter, , Michael C. Byrns, Yi Jin, Mo Chen, Jeffrey D. Winkler,*, † and Trevor M. Penning*, † Department of Pharmacology and Center of Excellence in Environmental Toxicology, Perelman School of Medicine, University of Pennsylvania, 130C John Morgan Building, 3620 Hamilton Walk, Philadelphia, Pennsylvania 19104-6084, United States ‡ Department of Chemistry, University of Pennsylvania, 231 South 34th Street, Philadelphia, Pennsylvania 19104, United States *S Supporting Information ABSTRACT: Aldo−keto reductase 1C3 (AKR1C3; type 5 17β-hydroxy- steroid dehydrogenase) is overexpressed in castration resistant prostate cancer (CRPC) and is implicated in the intratumoral biosynthesis of testosterone and 5α-dihydrotestosterone. Selective AKR1C3 inhibitors are required because compounds should not inhibit the highly related AKR1C1 and AKR1C2 isoforms which are involved in the inactivation of 5α-dihydrotestosterone. NSAIDs, N-phenylanthranilates in particular, are potent but nonselective AKR1C3 inhibitors. Using flufenamic acid, 2-{[3-(trifluoromethyl)phenyl]amino}- benzoic acid, as lead compound, five classes of structural analogues were synthesized and evaluated for AKR1C3 inhibitory potency and selectivity. Structure−activity relationship (SAR) studies revealed that a meta-carboxylic acid group relative to the amine conferred pronounced AKR1C3 selectivity without loss of potency, while electron withdrawing groups on the phenylamino B-ring were optimal for AKR1C3 inhibition. Lead compounds did not inhibit COX-1 or COX-2 but blocked the AKR1C3 mediated production of testosterone in LNCaP-AKR1C3 cells. These compounds offer promising leads toward new therapeutics for CRPC. -

1 T CELL ACTIVATION TRIGGERS REVERSIBLE INOSINE-5'-MONOPHOSPHATE DEHYDROGENASE ASSEMBLY Krisna C. Duong-Ly 1, Yin-Ming Kuo 2
bioRxiv preprint doi: https://doi.org/10.1101/315929; this version posted May 7, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. T CELL ACTIVATION TRIGGERS REVERSIBLE INOSINE-5’-MONOPHOSPHATE DEHYDROGENASE ASSEMBLY Krisna C. Duong-Ly1, Yin-Ming Kuo2, Matthew C. Johnson3, Justin M. Kollman3, Jonathan Soboloff4, Glenn F. Rall5, Andrew J. Andrews2, and Jeffrey R. Peterson1 1Cancer Biology Program, Fox Chase Cancer Center, Philadelphia, PA 2Cancer Epigenetics Program, Fox Chase Cancer Center, Philadelphia, PA 3Department of Biochemistry, University of Washington, Seattle, WA 4Fels Institute for Cancer Research and Molecular Biology, Temple University School of Medicine, Philadelphia, PA 5Blood Cell Development and Function Program, Fox Chase Cancer Center, Philadelphia, PA Correspondence to Jeffrey R. Peterson: [email protected], Fox Chase Cancer Center, 333 Cottman Avenue, Philadelphia, PA 19111, ORCID: 0000-0002-0604-718X; Yin-Ming Kuo’s present address is Department of Radiology, University of Pennsylvania School of Medicine, Philadelphia, PA Short running title: IMPDH FILAMENT ASSEMBLY IN T CELLS Abbreviations used: IMP: inosine-5’-monophosphate IMPDH: inosine-5’-monophosphate dehydrogenase 1 bioRxiv preprint doi: https://doi.org/10.1101/315929; this version posted May 7, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. -

Transcriptome Analysis of Different Multidrug-Resistant Gastric Carcinoma Cells
in vivo 19: 583-590 (2005) Transcriptome Analysis of Different Multidrug-resistant Gastric Carcinoma Cells STEFFEN HEIM1 and HERMANN LAGE2 1Europroteome AG, Berlin-Hennigsdorf; 2Institute of Pathology, Charité Campus Mitte, Schumannstr. 20/21, D-10117 Berlin, Germany Abstract. Multidrug resistance (MDR) of human cancers is advanced cancer. These therapeutic protocols produce an the major cause of failure of chemotherapy. To better overall response rate of about 20% at best using a single- understand the molecular events associated with the agent, and up to 50% using combination chemotherapy. development of different types of MDR, two different multidrug- Thus, a large number of malignancies are incurable. resistant gastric carcinoma cell lines, the MDR1/P-glycoprotein- Generally, gastrointestinal cancers, including gastric expressing cell line EPG85-257RDB and the MDR1/P- carcinoma, are naturally resistant to many anticancer drugs. glycoprotein-negative cell variant EPG85-257RNOV, as well as Additionally, these tumors are able to develop acquired the corresponding drug-sensitive parental cell line EPG85-257P, drug resistance phenotypes, which include the multidrug were used for analyses of the mRNA expression profiles by resistance (MDR) phenomenon. cDNA array hybridization. Of more than 12,000 genes spotted The MDR phenotype is characterized by simultaneous on the arrays, 156 genes were detected as being significantly resistance of tumor cells to various antineoplastic agents regulated in the cell line EPG85-257RDB in comparison to the which are structurally and functionally unrelated. Besides non-resistant cell variant, and 61 genes were found to be the classical MDR phenotype, mediated by the enhanced differentially expressed in the cell line EPG85-257RNOV.