FASTKD3 Rabbit Polyclonal Antibody – TA349969 | Origene
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -
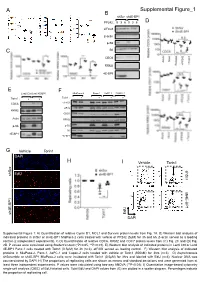
H Supplemental Figure 1 I D E B a C
A Supplemental Figure_1 B shScr sh4E-BP1 PP242 : 0 3 6 0 3 6 D eIF4G1 β-actin p-S6 C S6 CDC6 RRM2 4E-BP1 E F Lenti Ctrl Lenti 4E-BP1 MiaPaca-2 Panc-1 AsPC-1 Capan-2 Torin1 : Torin1 : + + + + + + CDC6 eIF4G1 eIF3a RRM2 CDC6 Actin RRM2 p-S6 p-S6 S6 4E-BP1 4E-BP1 G Vehicle Torin1 DAPI H I Vehicle Torin1 62.20% 45.30% shScr sh4E-BP1 EdU shScr 59.58% 49.47% EdU sh4E-BP1 DAPI Supplemental Figure 1: A) Quantification of relative Cyclin D1, MCL1 and Survivin protein levels from Fig. 1A. B) Western blot analysis of indicated proteins in shScr or sh4E-BP1 MiaPaca-2 cells treated with vehicle or PP242 (5µM) for 3h and 6h. β-actin served as a loading control (2 independent experiments). C-D) Quantification of relative CDC6, RRM2 and CDC7 protein levels from (C) Fig. 2C and (D) Fig. 2D. P values were calculated using Student’s t-test (*P<0.05, **P<0.01). E) Western blot analysis of indicated proteins in Lenti Ctrl or Lenti 4E-BP1 Panc-1 cells treated with Torin1 (0.5µM) for 3h (n=3). eIF4GI served as loading control. F) Western blot analysis of indicated proteins in MiaPaca-2, Panc-1, AsPC-1 and Capan-2 cells treated with vehicle or Torin1 (500nM) for 3hrs (n=3). G) Asynchronous shScramble or sh4E-BP1 MiaPaca-2 cells were incubated with Torin1 (0.5μM) for 3hrs and labeled with EdU (n=3). Nuclear DNA was counterstained by DAPI. H) The proportions of replicating cells are shown as means and standard deviations and were generated from at least three independent experiments. -

Molecular Risk Factors of Pulmonary Arterial Hypertension Hamza M
Florida International University FIU Digital Commons FIU Electronic Theses and Dissertations University Graduate School 9-22-2017 Molecular Risk Factors of Pulmonary Arterial Hypertension Hamza M. Assaggaf Florida International University, [email protected] DOI: 10.25148/etd.FIDC004013 Follow this and additional works at: https://digitalcommons.fiu.edu/etd Part of the Environmental Public Health Commons Recommended Citation Assaggaf, Hamza M., "Molecular Risk Factors of Pulmonary Arterial Hypertension" (2017). FIU Electronic Theses and Dissertations. 3554. https://digitalcommons.fiu.edu/etd/3554 This work is brought to you for free and open access by the University Graduate School at FIU Digital Commons. It has been accepted for inclusion in FIU Electronic Theses and Dissertations by an authorized administrator of FIU Digital Commons. For more information, please contact [email protected]. FLORIDA INTERNATIONAL UNIVERSITY Miami, Florida MOLECULAR RISK FACTORS OF PULMONARY ARTERIAL HYPERTENSION A dissertation submitted in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY in PUBLIC HEALTH by Hamza Assaggaf 2017 To: Dean Tomás R. Guilarte Robert Stempel College of Public Health and Social Work This dissertation, written by Hamza Assaggaf, entitled Molecular Risk Factors of Pulmonary Arterial Hypertension, having been approved in respect to style and intellectual content, is referred to you for judgment. We have read the dissertation and recommend that it be approved. __________________________________________ Alok Deoraj __________________________________________ Juan Luizzi __________________________________________ Deodutta Roy __________________________________________ ChangwonYoo, Co-Major Professor __________________________________________ Quentin Felty, Co-Major Professor Date of Defense: September 22, 2017 The dissertation of Hamza Assaggaf is approved. __________________________________________ Dean Tomás R. Guilarte Robert Stempel College of Public Health and Social Work __________________________________________ Andres G. -

Induction of Therapeutic Tissue Tolerance Foxp3 Expression Is
Downloaded from http://www.jimmunol.org/ by guest on October 2, 2021 is online at: average * The Journal of Immunology , 13 of which you can access for free at: 2012; 189:3947-3956; Prepublished online 17 from submission to initial decision 4 weeks from acceptance to publication September 2012; doi: 10.4049/jimmunol.1200449 http://www.jimmunol.org/content/189/8/3947 Foxp3 Expression Is Required for the Induction of Therapeutic Tissue Tolerance Frederico S. Regateiro, Ye Chen, Adrian R. Kendal, Robert Hilbrands, Elizabeth Adams, Stephen P. Cobbold, Jianbo Ma, Kristian G. Andersen, Alexander G. Betz, Mindy Zhang, Shruti Madhiwalla, Bruce Roberts, Herman Waldmann, Kathleen F. Nolan and Duncan Howie J Immunol cites 35 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription http://www.jimmunol.org/content/suppl/2012/09/17/jimmunol.120044 9.DC1 This article http://www.jimmunol.org/content/189/8/3947.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2012 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of October 2, 2021. -

ATA29119-FASTKD3 Rabbit Polyclonal Antibody
ATAGENIX LABORATORIES Catalog Number:ATA29119 FASTKD3 rabbit Polyclonal Antibody Product overview product name FASTKD3 rabbit Polyclonal Antibody catalog No. ATA29119 Category Primary antibodies Host Rabbit Species specificity Human Tested applications WB,IHC-p,ELISA Clonality Polyclonal Conjugation Unconjugated Immunogen The antiserum was produced against synthesized peptide derived from human FAKD3. AA range:121-170 Product performance Form Liquid Storage Use a manual defrost freezer and avoid repeated freeze thaw cycles. Store at 4 °C for frequent use. Store at -20 to -80 °C for twelve months from the date of receipt. Purity The antibody was affinity-purified from rabbit antiserum by affinity-chromatography using epitope-specific immunogen. Dilution range Western Blot: 1/500 - 1/2000. Immunohistochemistry: 1/100 - 1/300. ELISA: 1/20000. Not yet tested in other applications. Product background FAST kinase domains 3(FASTKD3) Homo sapiens This gene encodes a member of a small family of Fas-activated serine /threonine kinase domain (FASTKD) containing proteins that share an amino terminal mitochondrial targeting domain and multiple carboxy terminal FAST domains as well as a putative RNA-binding RAP domain. The members of this family are ubiquitously expressed and are generally most abundant in mitochondria-enriched tissues such as heart, skeletal muscle and brown-adipose tissue. Some members of this protein family may play a role in apoptosis. The protein encoded by this gene interacts with components of the mitochondrial respiratory and translation networks. A pseudogene of this gene is also present on chromosome 5. Alternative splicing results in multiple transcript variants. [provided by RefSeq, Jan 2013], Web:www.atagenix.com E-mail: [email protected] Tel: 027-87433958. -
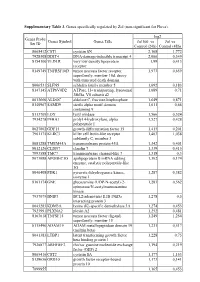
Supplementary Table 3. Genes Specifically Regulated by Zol (Non-Significant for Fluva)
Supplementary Table 3. Genes specifically regulated by Zol (non-significant for Fluva). log2 Genes Probe Genes Symbol Genes Title Zol100 vs Zol vs Set ID Control (24h) Control (48h) 8065412 CST1 cystatin SN 2,168 1,772 7928308 DDIT4 DNA-damage-inducible transcript 4 2,066 0,349 8154100 VLDLR very low density lipoprotein 1,99 0,413 receptor 8149749 TNFRSF10D tumor necrosis factor receptor 1,973 0,659 superfamily, member 10d, decoy with truncated death domain 8006531 SLFN5 schlafen family member 5 1,692 0,183 8147145 ATP6V0D2 ATPase, H+ transporting, lysosomal 1,689 0,71 38kDa, V0 subunit d2 8013660 ALDOC aldolase C, fructose-bisphosphate 1,649 0,871 8140967 SAMD9 sterile alpha motif domain 1,611 0,66 containing 9 8113709 LOX lysyl oxidase 1,566 0,524 7934278 P4HA1 prolyl 4-hydroxylase, alpha 1,527 0,428 polypeptide I 8027002 GDF15 growth differentiation factor 15 1,415 0,201 7961175 KLRC3 killer cell lectin-like receptor 1,403 1,038 subfamily C, member 3 8081288 TMEM45A transmembrane protein 45A 1,342 0,401 8012126 CLDN7 claudin 7 1,339 0,415 7993588 TMC7 transmembrane channel-like 7 1,318 0,3 8073088 APOBEC3G apolipoprotein B mRNA editing 1,302 0,174 enzyme, catalytic polypeptide-like 3G 8046408 PDK1 pyruvate dehydrogenase kinase, 1,287 0,382 isozyme 1 8161174 GNE glucosamine (UDP-N-acetyl)-2- 1,283 0,562 epimerase/N-acetylmannosamine kinase 7937079 BNIP3 BCL2/adenovirus E1B 19kDa 1,278 0,5 interacting protein 3 8043283 KDM3A lysine (K)-specific demethylase 3A 1,274 0,453 7923991 PLXNA2 plexin A2 1,252 0,481 8163618 TNFSF15 tumor necrosis -

Identification of Novel Regulatory Genes in Acetaminophen
IDENTIFICATION OF NOVEL REGULATORY GENES IN ACETAMINOPHEN INDUCED HEPATOCYTE TOXICITY BY A GENOME-WIDE CRISPR/CAS9 SCREEN A THESIS IN Cell Biology and Biophysics and Bioinformatics Presented to the Faculty of the University of Missouri-Kansas City in partial fulfillment of the requirements for the degree DOCTOR OF PHILOSOPHY By KATHERINE ANNE SHORTT B.S, Indiana University, Bloomington, 2011 M.S, University of Missouri, Kansas City, 2014 Kansas City, Missouri 2018 © 2018 Katherine Shortt All Rights Reserved IDENTIFICATION OF NOVEL REGULATORY GENES IN ACETAMINOPHEN INDUCED HEPATOCYTE TOXICITY BY A GENOME-WIDE CRISPR/CAS9 SCREEN Katherine Anne Shortt, Candidate for the Doctor of Philosophy degree, University of Missouri-Kansas City, 2018 ABSTRACT Acetaminophen (APAP) is a commonly used analgesic responsible for over 56,000 overdose-related emergency room visits annually. A long asymptomatic period and limited treatment options result in a high rate of liver failure, generally resulting in either organ transplant or mortality. The underlying molecular mechanisms of injury are not well understood and effective therapy is limited. Identification of previously unknown genetic risk factors would provide new mechanistic insights and new therapeutic targets for APAP induced hepatocyte toxicity or liver injury. This study used a genome-wide CRISPR/Cas9 screen to evaluate genes that are protective against or cause susceptibility to APAP-induced liver injury. HuH7 human hepatocellular carcinoma cells containing CRISPR/Cas9 gene knockouts were treated with 15mM APAP for 30 minutes to 4 days. A gene expression profile was developed based on the 1) top screening hits, 2) overlap with gene expression data of APAP overdosed human patients, and 3) biological interpretation including assessment of known and suspected iii APAP-associated genes and their therapeutic potential, predicted affected biological pathways, and functionally validated candidate genes. -
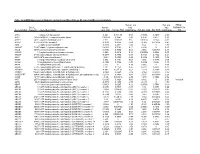
Mrna Expression in Human Leiomyoma and Eker Rats As Measured by Microarray Analysis
Table 3S: mRNA Expression in Human Leiomyoma and Eker Rats as Measured by Microarray Analysis Human_avg Rat_avg_ PENG_ Entrez. Human_ log2_ log2_ RAPAMYCIN Gene.Symbol Gene.ID Gene Description avg_tstat Human_FDR foldChange Rat_avg_tstat Rat_FDR foldChange _DN A1BG 1 alpha-1-B glycoprotein 4.982 9.52E-05 0.68 -0.8346 0.4639 -0.38 A1CF 29974 APOBEC1 complementation factor -0.08024 0.9541 -0.02 0.9141 0.421 0.10 A2BP1 54715 ataxin 2-binding protein 1 2.811 0.01093 0.65 0.07114 0.954 -0.01 A2LD1 87769 AIG2-like domain 1 -0.3033 0.8056 -0.09 -3.365 0.005704 -0.42 A2M 2 alpha-2-macroglobulin -0.8113 0.4691 -0.03 6.02 0 1.75 A4GALT 53947 alpha 1,4-galactosyltransferase 0.4383 0.7128 0.11 6.304 0 2.30 AACS 65985 acetoacetyl-CoA synthetase 0.3595 0.7664 0.03 3.534 0.00388 0.38 AADAC 13 arylacetamide deacetylase (esterase) 0.569 0.6216 0.16 0.005588 0.9968 0.00 AADAT 51166 aminoadipate aminotransferase -0.9577 0.3876 -0.11 0.8123 0.4752 0.24 AAK1 22848 AP2 associated kinase 1 -1.261 0.2505 -0.25 0.8232 0.4689 0.12 AAMP 14 angio-associated, migratory cell protein 0.873 0.4351 0.07 1.656 0.1476 0.06 AANAT 15 arylalkylamine N-acetyltransferase -0.3998 0.7394 -0.08 0.8486 0.456 0.18 AARS 16 alanyl-tRNA synthetase 5.517 0 0.34 8.616 0 0.69 AARS2 57505 alanyl-tRNA synthetase 2, mitochondrial (putative) 1.701 0.1158 0.35 0.5011 0.6622 0.07 AARSD1 80755 alanyl-tRNA synthetase domain containing 1 4.403 9.52E-05 0.52 1.279 0.2609 0.13 AASDH 132949 aminoadipate-semialdehyde dehydrogenase -0.8921 0.4247 -0.12 -2.564 0.02993 -0.32 AASDHPPT 60496 aminoadipate-semialdehyde -

MOL #82305 TITLE PAGE Title: Induced CYP3A4 Expression In
Downloaded from molpharm.aspetjournals.org at ASPET Journals on September 28, 2021 1 This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. The final version may differ from this version. This article has not been copyedited and formatted. -

Nanostring®: Product Data Sheet | Ncounter® GX Human Kinase
PRODUCT DATA SHEET nCounter® GX Human Kinase Kit nCounter® GX Human Kinase Kit Product Highlights Highly Curated • Our expert bio-informaticists use a very rigorous process in selecting the most meaningful set of genes Efficient • Multiplexed assay profiles 522 human kinase genes in a single reaction Cost-effective • Gold-standard data at a fraction of the cost Quick Turnaround Time • Complete kit with all consumables ready to ship next day nCounter® GX Human Kinase Kit The nCounter GX Human Kinase Kit is a comprehensive list of 522 human The nCounter Human Kinase Kit represents 99% of the KinBase content for genes known to be differentially expressed in the kinome. Human. With the nCounter GX Human Kinase Kit, scientists can leverage a pre-designed The final nCounter GX Human Kinase Kit consists of 522 protein kinase- panel to accelerate their research and quickly generate expression data for a related genes and 14 internal reference genes. For the gene list and additional large panel of protein kinase-related genes. information about this gene set, visit the nCounter Pre-built Panels product page at: www.nanostring.com. The gene list was compiled using the KinBase database at www.kinase.com. Home > Products > nCounter Gene Expression CodeSets > Pre-built Panels The database is based on the publication, The Protein Kinase Complement of the Human Genome, by G Manning, DB Whyte, R Martinez, T Hunter, S Sudarsanam (2002). Science 298:1912-1934. Molecules That Count® Translational Research Gene Expression miRNA Expression Copy Number Variation 1 PRODUCT DATA SHEET nCounter® GX Human Kinase Kit nCounter® Analysis System Overview The nCounter Analysis System from NanoString offers a cost-effective way to easily profile hundreds of gene transcripts simultaneously with high sensitivity and precision. -
Genome-Wide Association Study Reveals Two New Risk Loci for Bipolar Disorder
ARTICLE Received 14 Aug 2013 | Accepted 29 Jan 2014 | Published 11 Mar 2014 DOI: 10.1038/ncomms4339 Genome-wide association study reveals two new risk loci for bipolar disorder Thomas W. Mu¨hleisen1,2,3,*, Markus Leber4,5,*, Thomas G. Schulze6,*, Jana Strohmaier7, Franziska Degenhardt1,2, Jens Treutlein7, Manuel Mattheisen8,9, Andreas J. Forstner1,2, Johannes Schumacher1,2,Rene´ Breuer7, Sandra Meier7,10, Stefan Herms1,2,11,Per Hoffmann1,2,3,11, Andre´ Lacour5, Stephanie H. Witt7, Andreas Reif12, Bertram Mu¨ller-Myhsok13,14,15, Susanne Lucae13, Wolfgang Maier16, Markus Schwarz17, Helmut Vedder17, Jutta Kammerer-Ciernioch17, Andrea Pfennig18, Michael Bauer18, Martin Hautzinger19, Susanne Moebus20, Lutz Priebe1,2, Piotr M. Czerski21, Joanna Hauser21, Jolanta Lissowska22, Neonila Szeszenia-Dabrowska23, Paul Brennan24, James D. McKay25, Adam Wright26,27, Philip B. Mitchell26,27, Janice M. Fullerton28,29, Peter R. Schofield28,29, Grant W. Montgomery30, Sarah E. Medland30, Scott D. Gordon30, Nicholas G. Martin30, Valery Krasnow31, Alexander Chuchalin32, Gulja Babadjanova32, Galina Pantelejeva33, Lilia I. Abramova33, Alexander S. Tiganov33, Alexey Polonikov34, Elza Khusnutdinova35, Martin Alda36,37, Paul Grof37,38,39, Guy A. Rouleau40, Gustavo Turecki41, Catherine Laprise42, Fabio Rivas43, Fermin Mayoral43, Manolis Kogevinas44, Maria Grigoroiu- Serbanescu45, Peter Propping1, Tim Becker5,4, Marcella Rietschel7,*, Markus M. No¨then1,2,* & Sven Cichon1,2,3,11,* Bipolar disorder (BD) is a common and highly heritable mental illness and genome-wide asso- ciation studies (GWAS) have robustly identified the first common genetic variants involved in disease aetiology. The data also provide strong evidence for the presence of multiple additional risk loci, each contributing a relatively small effect to BD susceptibility. -

Thesis Reference
Thesis Regulation of mitochondrial RNA expression by FASTK proteins BOEHM, Erik Abstract This work focused on the regulation of Mitochondrial RNA by the FASTK proteins. CRISPR KOs were used to investigate the effects of the loss of FASTKD1, FASTKD3, and FASTKD4, while work with FASTK and FASTKD2 was done with siRNAs and MEF KOs. All FASTK proteins were found to affect mitochondrial RNA, but the specificity and effect on RNA was varied. In particular loss of FASTKD1 or FASTKD3 had opposite effects to FASTKD4 on the loss of the ND3 RNA. FASTKD4 loss showed a striking phenotype in which there was a loss of mature ND5 and CYB RNA and a large increase in the ND5-CYB precursor. Structural modeling and mutagenesis studies suggested a similarity of a domain of the FASTK proteins to an endonuclease. In contrast the N terminal region appears to resemble PPR proteins and determines if the protein is incorporated into mitochondrial RNA granules or not. Reference BOEHM, Erik. Regulation of mitochondrial RNA expression by FASTK proteins. Thèse de doctorat : Univ. Genève, 2016, no. Sc. 4929 URN : urn:nbn:ch:unige-877228 DOI : 10.13097/archive-ouverte/unige:87722 Available at: http://archive-ouverte.unige.ch/unige:87722 Disclaimer: layout of this document may differ from the published version. 1 / 1 UNIVERSITÉ DE GENÈVE Département de biologie cellulaire FACULTÉ DES SCIENCES Professeur Jean-Claude Martinou Regulation of mitochondrial RNA expression by FASTK proteins THÈSE présentée à la Faculté des sciences de l'Université de Genève pour obtenir le grade de Docteur ès sciences, mention biologie Erik BOEHM de Milpitas, États-Unis Thèse n° 4929 Genève (imprimeur) 2016 1 I.