Induction of Therapeutic Tissue Tolerance Foxp3 Expression Is
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Ciliated Cell Transcriptome A
THE CILIATED CELL TRANSCRIPTOME A DISSERTATION SUBMITTED TO THE DEPARTMENT OF BIOLOGICAL SCIENCES AND THE COMMITTEE ON GRADUATE STUDIES OF STANFORD UNIVERSITY IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY Ramona Hoh March 2010 ! © 2010 by Ramona Amy Hoh. All Rights Reserved. Re-distributed by Stanford University under license with the author. This work is licensed under a Creative Commons Attribution- Noncommercial 3.0 United States License. http://creativecommons.org/licenses/by-nc/3.0/us/ This dissertation is online at: http://purl.stanford.edu/sk794dv5857 ii I certify that I have read this dissertation and that, in my opinion, it is fully adequate in scope and quality as a dissertation for the degree of Doctor of Philosophy. Timothy Stearns, Primary Adviser I certify that I have read this dissertation and that, in my opinion, it is fully adequate in scope and quality as a dissertation for the degree of Doctor of Philosophy. Mark Krasnow I certify that I have read this dissertation and that, in my opinion, it is fully adequate in scope and quality as a dissertation for the degree of Doctor of Philosophy. Maxence Nachury I certify that I have read this dissertation and that, in my opinion, it is fully adequate in scope and quality as a dissertation for the degree of Doctor of Philosophy. William Nelson Approved for the Stanford University Committee on Graduate Studies. Patricia J. Gumport, Vice Provost Graduate Education This signature page was generated electronically upon submission of this dissertation in electronic format. An original signed hard copy of the signature page is on file in University Archives. -

Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model
Downloaded from http://www.jimmunol.org/ by guest on September 25, 2021 T + is online at: average * The Journal of Immunology , 34 of which you can access for free at: 2016; 197:1477-1488; Prepublished online 1 July from submission to initial decision 4 weeks from acceptance to publication 2016; doi: 10.4049/jimmunol.1600589 http://www.jimmunol.org/content/197/4/1477 Molecular Profile of Tumor-Specific CD8 Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. Waugh, Sonia M. Leach, Brandon L. Moore, Tullia C. Bruno, Jonathan D. Buhrman and Jill E. Slansky J Immunol cites 95 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2016/07/01/jimmunol.160058 9.DCSupplemental This article http://www.jimmunol.org/content/197/4/1477.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 25, 2021. The Journal of Immunology Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. -

ASPH-Notch Axis Guided Exosomal Delivery of Prometastatic Secretome
Lin et al. Molecular Cancer (2019) 18:156 https://doi.org/10.1186/s12943-019-1077-0 RESEARCH Open Access ASPH-notch Axis guided Exosomal delivery of Prometastatic Secretome renders breast Cancer multi-organ metastasis Qiushi Lin1†, Xuesong Chen2†, Fanzheng Meng3†, Kosuke Ogawa4†, Min Li5, Ruipeng Song3, Shugeng Zhang3, Ziran Zhang3, Xianglu Kong3, Qinggang Xu1,6, Fuliang He1,7, Xuewei Bai8, Bei Sun8, Mien-Chie Hung9,10, Lianxin Liu3,11*, Jack Wands4* and Xiaoqun Dong12,1* Abstract Background: Aspartate β-hydroxylase (ASPH) is silent in normal adult tissues only to re-emerge during oncogenesis where its function is required for generation and maintenance of malignant phenotypes. Exosomes enable prooncogenic secretome delivering and trafficking for long-distance cell-to-cell communication. This study aims to explore molecular mechanisms underlying how ASPH network regulates designated exosomes to program development and progression of breast cancer. Methods: Stable cell lines overexpressing or knocking-out of ASPH were established using lentivirus transfection or CRISPR-CAS9 systems. Western blot, MTT, immunofluorescence, luciferase reporter, co-immunoprecipitation, 2D/3-D invasion, tube formation, mammosphere formation, immunohistochemistry and newly developed in vitro metastasis were applied. Results: Through physical interactions with Notch receptors, ligands (JAGs) and regulators (ADAM10/17), ASPH activates Notch cascade to provide raw materials (especially MMPs/ADAMs) for synthesis/release of pro-metastatic exosomes. Exosomes orchestrate EMT, 2-D/3-D invasion, stemness, angiogenesis, and premetastatic niche formation. Small molecule inhibitors (SMIs) of ASPH’s β-hydroxylase specifically/efficiently abrogated in vitro metastasis, which mimics basement membrane invasion at primary site, intravasation/extravasation (transendothelial migration), and colonization/outgrowth at distant sites. -

Transcriptomic Analysis of the Aquaporin (AQP) Gene Family
Pancreatology 19 (2019) 436e442 Contents lists available at ScienceDirect Pancreatology journal homepage: www.elsevier.com/locate/pan Transcriptomic analysis of the Aquaporin (AQP) gene family interactome identifies a molecular panel of four prognostic markers in patients with pancreatic ductal adenocarcinoma Dimitrios E. Magouliotis a, b, Vasiliki S. Tasiopoulou c, Konstantinos Dimas d, * Nikos Sakellaridis d, Konstantina A. Svokos e, Alexis A. Svokos f, Dimitris Zacharoulis b, a Division of Surgery and Interventional Science, Faculty of Medical Sciences, UCL, London, UK b Department of Surgery, University of Thessaly, Biopolis, Larissa, Greece c Faculty of Medicine, School of Health Sciences, University of Thessaly, Biopolis, Larissa, Greece d Department of Pharmacology, Faculty of Medicine, School of Health Sciences, University of Thessaly, Biopolis, Larissa, Greece e The Warren Alpert Medical School of Brown University, Providence, RI, USA f Riverside Regional Medical Center, Newport News, VA, USA article info abstract Article history: Background: This study aimed to assess the differential gene expression of aquaporin (AQP) gene family Received 14 October 2018 interactome in pancreatic ductal adenocarcinoma (PDAC) using data mining techniques to identify novel Received in revised form candidate genes intervening in the pathogenicity of PDAC. 29 January 2019 Method: Transcriptome data mining techniques were used in order to construct the interactome of the Accepted 9 February 2019 AQP gene family and to determine which genes members are differentially expressed in PDAC as Available online 11 February 2019 compared to controls. The same techniques were used in order to evaluate the potential prognostic role of the differentially expressed genes. Keywords: PDAC Results: Transcriptome microarray data of four GEO datasets were incorporated, including 142 primary Aquaporin tumor samples and 104 normal pancreatic tissue samples. -

Regulation of Transcription and Regulatory Networks for Muscle Growth * * * * A
Regulation Of Transcription And Regulatory Networks For Muscle Growth * * * * A. Reverter , N.J. Hudson , Q. Gu and B.P. Dalrymple Introduction The advent of microarray gene expression technology has provided animal scientists with an unprecedented ability to profile the transcriptional changes during skeletal muscle growth. With respect to meat quality, most of the effort has concentrated on the understanding of fat and energy metabolism (reviewed by Hausman et al . (2009)). Graugnard et al . (2009) explored the network among 31 genes associated with aspects of adipogenesis and energy metabolism in bovine skeletal muscle and in response to two distinct diets. Also, Freyssenet (2007) reviewed the roles that energy-sensing molecules and mitochondria have in the regulation of gene expression in muscle. However, other mechanisms such as cell cycle, glycolysis, extra-cellular matrix, ribosomal proteins and the immune system play a significant role in development, and this role can work in a tissue-specific manner. Hudson et al . (2009a) reported various functional modules underpinning the transcriptional regulation of bovine skeletal muscle. The authors integrated a total of six gene co-expression networks, each developed using the PCIT algorithm (Reverter and Chan (2008)), and proposed a Module-to-Regulator heuristic by which those transcription factors (TF) with the highest average absolute correlation co-expression with the genes present in each module are deemed to be the relevant regulators. However, this Module-to-Regulator approach failed to capture some well-known regulators of muscle fibre type composition, and the use of more sophisticated methods such as the differential wiring approach of Hudson et al . -

Iron Depletion Reduces Abce1 Transcripts While Inducing The
Preprints (www.preprints.org) | NOT PEER-REVIEWED | Posted: 22 October 2019 doi:10.20944/preprints201910.0252.v1 1 Research Article 2 Iron depletion Reduces Abce1 Transcripts While 3 Inducing the Mitophagy Factors Pink1 and Parkin 4 Jana Key 1,2, Nesli Ece Sen 1, Aleksandar Arsovic 1, Stella Krämer 1, Robert Hülse 1, Suzana 5 Gispert-Sanchez 1 and Georg Auburger 1,* 6 1 Experimental Neurology, Goethe University Medical School, 60590 Frankfurt am Main; 7 2 Faculty of Biosciences, Goethe-University Frankfurt am Main, Germany 8 * Correspondence: [email protected] 9 10 Abstract: Lifespan extension was recently achieved in Caenorhabditis elegans nematodes by 11 mitochondrial stress and mitophagy, triggered via iron depletion. Conversely in man, deficient 12 mitophagy due to Pink1/Parkin mutations triggers iron accumulation in patient brain and limits 13 survival. We now aimed to identify murine fibroblast factors, which adapt their mRNA expression 14 to acute iron manipulation, relate to mitochondrial dysfunction and may influence survival. After 15 iron depletion, expression of the plasma membrane receptor Tfrc with its activator Ireb2, the 16 mitochondrial membrane transporter Abcb10, the heme-release factor Pgrmc1, the heme- 17 degradation enzyme Hmox1, the heme-binding cholesterol metabolizer Cyp46a1, as well as the 18 mitophagy regulators Pink1 and Parkin showed a negative correlation to iron levels. After iron 19 overload, these factors did not change expression. Conversely, a positive correlation of mRNA levels 20 with both conditions of iron availability was observed for the endosomal factors Slc11a2 and Steap2, 21 as well as for the iron-sulfur-cluster (ISC)-containing factors Ppat, Bdh2 and Nthl1. -

Molecular Dissection of G-Protein Coupled Receptor Signaling and Oligomerization
MOLECULAR DISSECTION OF G-PROTEIN COUPLED RECEPTOR SIGNALING AND OLIGOMERIZATION BY MICHAEL RIZZO A Dissertation Submitted to the Graduate Faculty of WAKE FOREST UNIVERSITY GRADUATE SCHOOL OF ARTS AND SCIENCES in Partial Fulfillment of the Requirements for the Degree of DOCTOR OF PHILOSOPHY Biology December, 2019 Winston-Salem, North Carolina Approved By: Erik C. Johnson, Ph.D. Advisor Wayne E. Pratt, Ph.D. Chair Pat C. Lord, Ph.D. Gloria K. Muday, Ph.D. Ke Zhang, Ph.D. ACKNOWLEDGEMENTS I would first like to thank my advisor, Dr. Erik Johnson, for his support, expertise, and leadership during my time in his lab. Without him, the work herein would not be possible. I would also like to thank the members of my committee, Dr. Gloria Muday, Dr. Ke Zhang, Dr. Wayne Pratt, and Dr. Pat Lord, for their guidance and advice that helped improve the quality of the research presented here. I would also like to thank members of the Johnson lab, both past and present, for being valuable colleagues and friends. I would especially like to thank Dr. Jason Braco, Dr. Jon Fisher, Dr. Jake Saunders, and Becky Perry, all of whom spent a great deal of time offering me advice, proofreading grants and manuscripts, and overall supporting me through the ups and downs of the research process. Finally, I would like to thank my family, both for instilling in me a passion for knowledge and education, and for their continued support. In particular, I would like to thank my wife Emerald – I am forever indebted to you for your support throughout this process, and I will never forget the sacrifices you made to help me get to where I am today. -

Protein Identities in Evs Isolated from U87-MG GBM Cells As Determined by NG LC-MS/MS
Protein identities in EVs isolated from U87-MG GBM cells as determined by NG LC-MS/MS. No. Accession Description Σ Coverage Σ# Proteins Σ# Unique Peptides Σ# Peptides Σ# PSMs # AAs MW [kDa] calc. pI 1 A8MS94 Putative golgin subfamily A member 2-like protein 5 OS=Homo sapiens PE=5 SV=2 - [GG2L5_HUMAN] 100 1 1 7 88 110 12,03704523 5,681152344 2 P60660 Myosin light polypeptide 6 OS=Homo sapiens GN=MYL6 PE=1 SV=2 - [MYL6_HUMAN] 100 3 5 17 173 151 16,91913397 4,652832031 3 Q6ZYL4 General transcription factor IIH subunit 5 OS=Homo sapiens GN=GTF2H5 PE=1 SV=1 - [TF2H5_HUMAN] 98,59 1 1 4 13 71 8,048185945 4,652832031 4 P60709 Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 - [ACTB_HUMAN] 97,6 5 5 35 917 375 41,70973209 5,478027344 5 P13489 Ribonuclease inhibitor OS=Homo sapiens GN=RNH1 PE=1 SV=2 - [RINI_HUMAN] 96,75 1 12 37 173 461 49,94108966 4,817871094 6 P09382 Galectin-1 OS=Homo sapiens GN=LGALS1 PE=1 SV=2 - [LEG1_HUMAN] 96,3 1 7 14 283 135 14,70620005 5,503417969 7 P60174 Triosephosphate isomerase OS=Homo sapiens GN=TPI1 PE=1 SV=3 - [TPIS_HUMAN] 95,1 3 16 25 375 286 30,77169764 5,922363281 8 P04406 Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 - [G3P_HUMAN] 94,63 2 13 31 509 335 36,03039959 8,455566406 9 Q15185 Prostaglandin E synthase 3 OS=Homo sapiens GN=PTGES3 PE=1 SV=1 - [TEBP_HUMAN] 93,13 1 5 12 74 160 18,68541938 4,538574219 10 P09417 Dihydropteridine reductase OS=Homo sapiens GN=QDPR PE=1 SV=2 - [DHPR_HUMAN] 93,03 1 1 17 69 244 25,77302971 7,371582031 11 P01911 HLA class II histocompatibility antigen, -

4-6 Weeks Old Female C57BL/6 Mice Obtained from Jackson Labs Were Used for Cell Isolation
Methods Mice: 4-6 weeks old female C57BL/6 mice obtained from Jackson labs were used for cell isolation. Female Foxp3-IRES-GFP reporter mice (1), backcrossed to B6/C57 background for 10 generations, were used for the isolation of naïve CD4 and naïve CD8 cells for the RNAseq experiments. The mice were housed in pathogen-free animal facility in the La Jolla Institute for Allergy and Immunology and were used according to protocols approved by the Institutional Animal Care and use Committee. Preparation of cells: Subsets of thymocytes were isolated by cell sorting as previously described (2), after cell surface staining using CD4 (GK1.5), CD8 (53-6.7), CD3ε (145- 2C11), CD24 (M1/69) (all from Biolegend). DP cells: CD4+CD8 int/hi; CD4 SP cells: CD4CD3 hi, CD24 int/lo; CD8 SP cells: CD8 int/hi CD4 CD3 hi, CD24 int/lo (Fig S2). Peripheral subsets were isolated after pooling spleen and lymph nodes. T cells were enriched by negative isolation using Dynabeads (Dynabeads untouched mouse T cells, 11413D, Invitrogen). After surface staining for CD4 (GK1.5), CD8 (53-6.7), CD62L (MEL-14), CD25 (PC61) and CD44 (IM7), naïve CD4+CD62L hiCD25-CD44lo and naïve CD8+CD62L hiCD25-CD44lo were obtained by sorting (BD FACS Aria). Additionally, for the RNAseq experiments, CD4 and CD8 naïve cells were isolated by sorting T cells from the Foxp3- IRES-GFP mice: CD4+CD62LhiCD25–CD44lo GFP(FOXP3)– and CD8+CD62LhiCD25– CD44lo GFP(FOXP3)– (antibodies were from Biolegend). In some cases, naïve CD4 cells were cultured in vitro under Th1 or Th2 polarizing conditions (3, 4). -
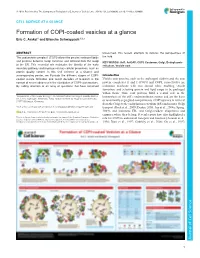
Formation of COPI-Coated Vesicles at a Glance Eric C
© 2018. Published by The Company of Biologists Ltd | Journal of Cell Science (2018) 131, jcs209890. doi:10.1242/jcs.209890 CELL SCIENCE AT A GLANCE Formation of COPI-coated vesicles at a glance Eric C. Arakel1 and Blanche Schwappach1,2,* ABSTRACT unresolved, this review attempts to refocus the perspectives of The coat protein complex I (COPI) allows the precise sorting of lipids the field. and proteins between Golgi cisternae and retrieval from the Golgi KEY WORDS: Arf1, ArfGAP, COPI, Coatomer, Golgi, Endoplasmic to the ER. This essential role maintains the identity of the early reticulum, Vesicle coat secretory pathway and impinges on key cellular processes, such as protein quality control. In this Cell Science at a Glance and accompanying poster, we illustrate the different stages of COPI- Introduction coated vesicle formation and revisit decades of research in the Vesicle coat proteins, such as the archetypal clathrin and the coat context of recent advances in the elucidation of COPI coat structure. protein complexes II and I (COPII and COPI, respectively) are By calling attention to an array of questions that have remained molecular machines with two central roles: enabling vesicle formation, and selecting protein and lipid cargo to be packaged within them. Thus, coat proteins fulfil a central role in the 1Department of Molecular Biology, Universitätsmedizin Göttingen, Humboldtallee homeostasis of the cell’s endomembrane system and are the basis 23, 37073 Göttingen, Germany. 2Max-Planck Institute for Biophysical Chemistry, 37077 Göttingen, Germany. of functionally segregated compartments. COPI operates in retrieval from the Golgi to the endoplasmic reticulum (ER) and in intra-Golgi *Author for correspondence ([email protected]) transport (Beck et al., 2009; Duden, 2003; Lee et al., 2004a; Spang, E.C.A., 0000-0001-7716-7149; B.S., 0000-0003-0225-6432 2009), and maintains ER- and Golgi-resident chaperones and enzymes where they belong. -

A2B Adenosine Receptors and T Cell Activation 493
Journal of Cell Science 112, 491-502 (1999) 491 Printed in Great Britain © The Company of Biologists Limited 1999 JCS0069 Expression of A2B adenosine receptors in human lymphocytes: their role in T cell activation Maribel Mirabet1, Carolina Herrera1, Oscar J. Cordero2, Josefa Mallol1, Carmen Lluis1 and Rafael Franco1,* 1Department of Biochemistry and Molecular Biology, Faculty of Chemistry, University of Barcelona, Barcelona, Catalonia, Spain 2Department of Biochemistry and Molecular Biology, Faculty of Biology, University of Santiago de Compostela, Spain *Author for correspondence (e-mail: [email protected]; homepage: www.bq.ub.es/recep/franco.html) Accepted 9 December 1998; published on WWW 25 January 1999 SUMMARY Extracellular adenosine has a key role in the development A2BRs but not of A2A receptors in these human cells. The and function of the cells of the immune system. Many of percentage of A2BR-expressing cells was similar in the the adenosine actions seem to be mediated by specific CD4+ or CD8+ T cell subpopulations. Interestingly surface receptors positively coupled to adenylate cyclase: activation signals delivered by either phytohemagglutinin A2A and A2B. Despite the fact that A2A receptors (A2ARs) or anti-T cell receptor/CD3 complex antibodies led to a can be easily studied due to the availability of the specific significant increase in both the percentage of cells agonist CGS21680, a pharmacological and physiological expressing the receptor and the intensity of the labeling. characterization of adenosine A2B receptors (A2BRs) in These receptors are functional since interleukin-2 lymphocytes has not been possible due to the lack of production in these cells is reduced by NECA but not by R- suitable reagents. -

Kinetochore Kinesin CENP-E Is a Processive Bi-Directional Tracker of Dynamic Microtubule Tips
ARTICLES Kinetochore kinesin CENP-E is a processive bi-directional tracker of dynamic microtubule tips Nikita Gudimchuk1,5, Benjamin Vitre2,5, Yumi Kim2,6, Anatoly Kiyatkin1, Don W. Cleveland2, Fazly I. Ataullakhanov3,4 and Ekaterina L. Grishchuk1,7 During vertebrate mitosis, the centromere-associated kinesin CENP-E (centromere protein E) transports misaligned chromosomes to the plus ends of spindle microtubules. Subsequently, the kinetochores that form at the centromeres establish stable associations with microtubule ends, which assemble and disassemble dynamically. Here we provide evidence that after chromosomes have congressed and bi-oriented, the CENP-E motor continues to play an active role at kinetochores, enhancing their links with dynamic microtubule ends. Using a combination of single-molecule approaches and laser trapping in vitro, we demonstrate that once reaching microtubule ends, CENP-E converts from a lateral transporter into a microtubule tip-tracker that maintains association with both assembling and disassembling microtubule tips. Computational modelling of this behaviour supports our proposal that CENP-E tip-tracks bi-directionally through a tethered motor mechanism, which relies on both the motor and tail domains of CENP-E. Our results provide a molecular framework for the contribution of CENP-E to the stability of attachments between kinetochores and dynamic microtubule ends. Accurate chromosome segregation depends on interactions between proportion of lagging chromosomes in anaphase in mouse liver microtubules and the kinetochore, a protein structure localized at cells and embryonic fibroblasts11,17. Fourth, after CENP-E-mediated each centromere1. Initially, kinetochores often attach to the walls congression, CENP-E-dependent localization of protein phosphatase of microtubules with the chromosomes then moving towards a 1 (PP1) to kinetochores is still required for stable microtubule spindle pole in a dynein-dependent manner2,3.