Polymerase Ribozyme with Promoter Recognition
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Molecular Mechanisms Involved Involved in the Interaction Effects of HCV and Ethanol on Liver Cirrhosis
Virginia Commonwealth University VCU Scholars Compass Theses and Dissertations Graduate School 2010 Molecular Mechanisms Involved Involved in the Interaction Effects of HCV and Ethanol on Liver Cirrhosis Ryan Fassnacht Virginia Commonwealth University Follow this and additional works at: https://scholarscompass.vcu.edu/etd Part of the Physiology Commons © The Author Downloaded from https://scholarscompass.vcu.edu/etd/2246 This Thesis is brought to you for free and open access by the Graduate School at VCU Scholars Compass. It has been accepted for inclusion in Theses and Dissertations by an authorized administrator of VCU Scholars Compass. For more information, please contact [email protected]. Ryan C. Fassnacht 2010 All Rights Reserved Molecular Mechanisms Involved in the Interaction Effects of HCV and Ethanol on Liver Cirrhosis A thesis submitted in partial fulfillment of the requirements for the degree of Master of Science at Virginia Commonwealth University. by Ryan Christopher Fassnacht, B.S. Hampden Sydney University, 2005 M.S. Virginia Commonwealth University, 2010 Director: Valeria Mas, Ph.D., Associate Professor of Surgery and Pathology Division of Transplant Department of Surgery Virginia Commonwealth University Richmond, Virginia July 9, 2010 Acknowledgement The Author wishes to thank his family and close friends for their support. He would also like to thank the members of the molecular transplant team for their help and advice. This project would not have been possible with out the help of Dr. Valeria Mas and her endearing -

Yeast Genome Gazetteer P35-65
gazetteer Metabolism 35 tRNA modification mitochondrial transport amino-acid metabolism other tRNA-transcription activities vesicular transport (Golgi network, etc.) nitrogen and sulphur metabolism mRNA synthesis peroxisomal transport nucleotide metabolism mRNA processing (splicing) vacuolar transport phosphate metabolism mRNA processing (5’-end, 3’-end processing extracellular transport carbohydrate metabolism and mRNA degradation) cellular import lipid, fatty-acid and sterol metabolism other mRNA-transcription activities other intracellular-transport activities biosynthesis of vitamins, cofactors and RNA transport prosthetic groups other transcription activities Cellular organization and biogenesis 54 ionic homeostasis organization and biogenesis of cell wall and Protein synthesis 48 plasma membrane Energy 40 ribosomal proteins organization and biogenesis of glycolysis translation (initiation,elongation and cytoskeleton gluconeogenesis termination) organization and biogenesis of endoplasmic pentose-phosphate pathway translational control reticulum and Golgi tricarboxylic-acid pathway tRNA synthetases organization and biogenesis of chromosome respiration other protein-synthesis activities structure fermentation mitochondrial organization and biogenesis metabolism of energy reserves (glycogen Protein destination 49 peroxisomal organization and biogenesis and trehalose) protein folding and stabilization endosomal organization and biogenesis other energy-generation activities protein targeting, sorting and translocation vacuolar and lysosomal -

(12) Patent Application Publication (10) Pub. No.: US 2010/0317005 A1 Hardin Et Al
US 20100317005A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2010/0317005 A1 Hardin et al. (43) Pub. Date: Dec. 16, 2010 (54) MODIFIED NUCLEOTIDES AND METHODS (22) Filed: Mar. 15, 2010 FOR MAKING AND USE SAME Related U.S. Application Data (63) Continuation of application No. 11/007,794, filed on Dec. 8, 2004, now abandoned, which is a continuation (75) Inventors: Susan H. Hardin, College Station, in-part of application No. 09/901,782, filed on Jul. 9, TX (US); Hongyi Wang, Pearland, 2001. TX (US); Brent A. Mulder, (60) Provisional application No. 60/527,909, filed on Dec. Sugarland, TX (US); Nathan K. 8, 2003, provisional application No. 60/216,594, filed Agnew, Richmond, TX (US); on Jul. 7, 2000. Tommie L. Lincecum, JR., Publication Classification Houston, TX (US) (51) Int. Cl. CI2O I/68 (2006.01) Correspondence Address: (52) U.S. Cl. ............................................................ 435/6 LIFE TECHNOLOGES CORPORATION (57) ABSTRACT CFO INTELLEVATE Labeled nucleotide triphosphates are disclosed having a label P.O. BOX S2OSO bonded to the gamma phosphate of the nucleotide triphos MINNEAPOLIS, MN 55402 (US) phate. Methods for using the gamma phosphate labeled nucleotide are also disclosed where the gamma phosphate labeled nucleotide are used to attach the labeled gamma phos (73) Assignees: LIFE TECHNOLOGIES phate in a catalyzed (enzyme or man-made catalyst) reaction to a target biomolecule or to exchange a phosphate on a target CORPORATION, Carlsbad, CA biomolecule with a labeled gamme phosphate. Preferred tar (US); VISIGEN get biomolecules are DNAs, RNAs, DNA/RNAs, PNA, BIOTECHNOLOGIES, INC. polypeptide (e.g., proteins enzymes, protein, assemblages, etc.), Sugars and polysaccharides or mixed biomolecules hav ing two or more of DNAs, RNAs, DNA/RNAs, polypeptide, (21) Appl. -

Characterization of Human UMP/CMP Kinase and Its Phosphorylation of D- and 1 L-Form Deoxycytidine Analogue Monophosphates
[CANCER RESEARCH 62, 1624–1631, March 15, 2002] Characterization of Human UMP/CMP Kinase and Its Phosphorylation of D- and 1 L-Form Deoxycytidine Analogue Monophosphates Jieh-Yuan Liou, Ginger E. Dutschman, Wing Lam, Zaoli Jiang, and Yung-Chi Cheng2 Department of Pharmacology, Yale University School of Medicine, New Haven, Connecticut 06520 ABSTRACT with leukemia, lymphoma, or solid tumors (11). Deoxycytidine ana- logues, such as -D-2Ј,3Ј-dideoxycytidine and L-(Ϫ)-SddC (Lamivu- Pyrimidine nucleoside monophosphate kinase [UMP/CMP kinase dine), have been shown to have anti-HIV and antihuman hepatitis B (UMP/CMPK); EC 2.7.4.14] plays a crucial role in the formation of UDP, virus activities (12–17). L-(Ϫ)-SddC was the first nucleoside analogue CDP, and dCDP, which are required for cellular nucleic acid synthesis. Several cytidine and deoxycytidine analogues are important anticancer with an L configuration to show therapeutic activity and, thus, defined  Ј Ј and antiviral drugs. These drugs require stepwise phosphorylation to their a new category for the design of nucleoside analogues. -L-2 ,3 - triphosphate forms to exert their therapeutic effects. The role of UMP/ dideoxy-5-fluoro-3Ј-thia-cytidine and -L-2Ј,3Ј-dideoxy-2Ј,3Ј-dide- CMPK for the phosphorylation of nucleoside analogues has been indi- hydro-5-fluorocytidine have been shown to be potent antihuman hep- cated. Thus, we cloned the human UMP/CMPK gene, expressed it in atitis B virus agents in vitro and in animal studies (18–22). In studies Escherichia coli, and purified it to homogeneity. Its kinetic properties of other -L-(Ϫ)-2Ј,3Ј-dideoxycytidine analogues, it was observed were determined. -

Structure-Function Studies of Enzymes from Ribose Metabolism
Comprehensive Summaries of Uppsala Dissertations from the Faculty of Science and Technology 939 Structure-Function Studies of Enzymes from Ribose Metabolism BY C. EVALENA ANDERSSON ACTA UNIVERSITATIS UPSALIENSIS UPPSALA 2004 !"" #$"" % & % % ' ( ) * + &( , +( !""( - . - % + / % 0 ( , ( 1#1( ( ( 2-3 1. 45 ." 2 * & & * % * &( , % . * % % ( ) % / ( 0 6 / % ,)' & % % & ( )* % 6 % 6 * ( 0 6 * * % ( - % & 7 % & % & && ( ' && ,)' % /( 2 8 * ,)' & ,'.'' ( ) * % / % * 6 & & / 6 ( 0 . . . ( - * & * % %% & ( 9 * 6 / %% % ( -: % & * . & . , /( , & % * /( ) % / % & % ( ! 6 . . & / 6 % " # $ % # %& '()# %$# # *+',-. # ; ( + , !"" 2--3 ".!#!< 2-3 1. 45 ." $ $$$ .#111 = $>> (6(> ? @ $ $$$ .#111A List of Papers This thesis is based on the following papers, which are referred to in the text by their Roman numerals: I Andersson, C. E. & Mowbray, S. L. (2002). Activation of ribokinase by monovalent cations. J. Mol. Biol. 315, 409-19 II Zhang, R., Andersson, C. E., Savchenko, -

The Microbiota-Produced N-Formyl Peptide Fmlf Promotes Obesity-Induced Glucose
Page 1 of 230 Diabetes Title: The microbiota-produced N-formyl peptide fMLF promotes obesity-induced glucose intolerance Joshua Wollam1, Matthew Riopel1, Yong-Jiang Xu1,2, Andrew M. F. Johnson1, Jachelle M. Ofrecio1, Wei Ying1, Dalila El Ouarrat1, Luisa S. Chan3, Andrew W. Han3, Nadir A. Mahmood3, Caitlin N. Ryan3, Yun Sok Lee1, Jeramie D. Watrous1,2, Mahendra D. Chordia4, Dongfeng Pan4, Mohit Jain1,2, Jerrold M. Olefsky1 * Affiliations: 1 Division of Endocrinology & Metabolism, Department of Medicine, University of California, San Diego, La Jolla, California, USA. 2 Department of Pharmacology, University of California, San Diego, La Jolla, California, USA. 3 Second Genome, Inc., South San Francisco, California, USA. 4 Department of Radiology and Medical Imaging, University of Virginia, Charlottesville, VA, USA. * Correspondence to: 858-534-2230, [email protected] Word Count: 4749 Figures: 6 Supplemental Figures: 11 Supplemental Tables: 5 1 Diabetes Publish Ahead of Print, published online April 22, 2019 Diabetes Page 2 of 230 ABSTRACT The composition of the gastrointestinal (GI) microbiota and associated metabolites changes dramatically with diet and the development of obesity. Although many correlations have been described, specific mechanistic links between these changes and glucose homeostasis remain to be defined. Here we show that blood and intestinal levels of the microbiota-produced N-formyl peptide, formyl-methionyl-leucyl-phenylalanine (fMLF), are elevated in high fat diet (HFD)- induced obese mice. Genetic or pharmacological inhibition of the N-formyl peptide receptor Fpr1 leads to increased insulin levels and improved glucose tolerance, dependent upon glucagon- like peptide-1 (GLP-1). Obese Fpr1-knockout (Fpr1-KO) mice also display an altered microbiome, exemplifying the dynamic relationship between host metabolism and microbiota. -

Product Sheet Info
Master Clone List for NR-19279 ® Vibrio cholerae Gateway Clone Set, Recombinant in Escherichia coli, Plates 1-46 Catalog No. NR-19279 Table 1: Vibrio cholerae Gateway® Clones, Plate 1 (NR-19679) Clone ID Well ORF Locus ID Symbol Product Accession Position Length Number 174071 A02 367 VC2271 ribD riboflavin-specific deaminase NP_231902.1 174346 A03 336 VC1877 lpxK tetraacyldisaccharide 4`-kinase NP_231511.1 174354 A04 342 VC0953 holA DNA polymerase III, delta subunit NP_230600.1 174115 A05 388 VC2085 sucC succinyl-CoA synthase, beta subunit NP_231717.1 174310 A06 506 VC2400 murC UDP-N-acetylmuramate--alanine ligase NP_232030.1 174523 A07 132 VC0644 rbfA ribosome-binding factor A NP_230293.2 174632 A08 322 VC0681 ribF riboflavin kinase-FMN adenylyltransferase NP_230330.1 174930 A09 433 VC0720 phoR histidine protein kinase PhoR NP_230369.1 174953 A10 206 VC1178 conserved hypothetical protein NP_230823.1 174976 A11 213 VC2358 hypothetical protein NP_231988.1 174898 A12 369 VC0154 trmA tRNA (uracil-5-)-methyltransferase NP_229811.1 174059 B01 73 VC2098 hypothetical protein NP_231730.1 174075 B02 82 VC0561 rpsP ribosomal protein S16 NP_230212.1 174087 B03 378 VC1843 cydB-1 cytochrome d ubiquinol oxidase, subunit II NP_231477.1 174099 B04 383 VC1798 eha eha protein NP_231433.1 174294 B05 494 VC0763 GTP-binding protein NP_230412.1 174311 B06 314 VC2183 prsA ribose-phosphate pyrophosphokinase NP_231814.1 174603 B07 108 VC0675 thyA thymidylate synthase NP_230324.1 174474 B08 466 VC1297 asnS asparaginyl-tRNA synthetase NP_230942.2 174933 B09 198 -

200703 Supplemental Magnani Et Al Clean Version
Supplemental Data Sleeping Beauty-engineered CAR T cells achieve anti-leukemic activity without severe toxicities Chiara F. Magnani, Giuseppe Gaipa, Federico Lussana, Daniela Belotti, Giuseppe Gritti, Sara Napolitano, Giada Matera, Benedetta Cabiati, Chiara Buracchi, Gianmaria Borleri, Grazia Fazio, Silvia Zaninelli, Sarah Tettamanti, Stefania Cesana, Valentina Colombo, Michele Quaroni, Giovanni Cazzaniga, Attilio Rovelli, Ettore Biagi, Stefania Galimberti, Andrea Calabria, Fabrizio Benedicenti, Eugenio Montini, Silvia Ferrari, Martino Introna, Adriana Balduzzi, Maria Grazia Valsecchi, Giuseppe Dastoli, Alessandro Rambaldi, Andrea Biondi Table of Contents: Supplemental Methods ...................................................................................................... 2 Supplemental Figure 1. .................................................................................................... 13 Supplemental Figure 2. .................................................................................................... 14 Supplemental Figure 3. .................................................................................................... 15 Supplemental Figure 4. .................................................................................................... 16 Supplemental Figure 5. .................................................................................................... 17 Supplemental Figure 6. .................................................................................................... 18 Supplemental -

Supplementary Information
Supplementary information (a) (b) Figure S1. Resistant (a) and sensitive (b) gene scores plotted against subsystems involved in cell regulation. The small circles represent the individual hits and the large circles represent the mean of each subsystem. Each individual score signifies the mean of 12 trials – three biological and four technical. The p-value was calculated as a two-tailed t-test and significance was determined using the Benjamini-Hochberg procedure; false discovery rate was selected to be 0.1. Plots constructed using Pathway Tools, Omics Dashboard. Figure S2. Connectivity map displaying the predicted functional associations between the silver-resistant gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S3. Connectivity map displaying the predicted functional associations between the silver-sensitive gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S4. Metabolic overview of the pathways in Escherichia coli. The pathways involved in silver-resistance are coloured according to respective normalized score. Each individual score represents the mean of 12 trials – three biological and four technical. Amino acid – upward pointing triangle, carbohydrate – square, proteins – diamond, purines – vertical ellipse, cofactor – downward pointing triangle, tRNA – tee, and other – circle. -

Enzymatic and Structural Characterization of the Naegleria Fowleri Glucokinase 1 2 Jillian E. Milanes1, Jimmy Suryadi1, Jan
bioRxiv preprint doi: https://doi.org/10.1101/469072; this version posted November 14, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Enzymatic and structural characterization of the Naegleria fowleri glucokinase 2 3 Jillian E. Milanes1, Jimmy Suryadi1, Jan Abendroth2, Wesley C. Van Voorhis3, Kayleigh F. 4 Barrett3, David M. Dranow2, Isabelle Q. Phan4, Stephen L. Patrick1, Soren D. Rozema5, 5 Muhammad M. Khalifa5, Jennifer E. Golden5, and James C. Morris1# 6 7 8 1Eukaryotic Pathogens Innovation Center 9 Department of Genetics and Biochemistry 10 Clemson University 11 Clemson, SC 29634 12 13 2CrystalCore 14 Beryllium Discovery 15 Bainbridge Island, WA 98110 16 17 3Center for Emerging and Re-emerging Infectious Diseases 18 Division of Allergy and Infectious Diseases, Department of Medicine 19 University of Washington 20 Seattle, WA 98109 21 22 4Seattle Structural Genomics Center for Infectious Disease 23 Center for Global Infection Disease Research 24 Seattle Children’s Research Institute 25 Seattle, WA 98109 26 27 5School of Pharmacy 28 Pharmaceutical Sciences Division 29 University of Wisconsin-Madison 30 Madison, WI 53705 31 32 33 Running title: Crystal structure of the glucokinase from the human pathogen N. fowleri 34 35 #Address correspondence to: James C. Morris, PhD, 249 LSB, 190 Collings Street, 36 Clemson SC 29634. FAX: (864) 656-0393; Email: [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/469072; this version posted November 14, 2018. -
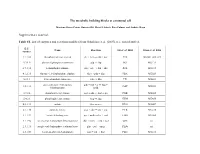
The Metabolic Building Blocks of a Minimal Cell Supplementary
The metabolic building blocks of a minimal cell Mariana Reyes-Prieto, Rosario Gil, Mercè Llabrés, Pere Palmer and Andrés Moya Supplementary material. Table S1. List of enzymes and reactions modified from Gabaldon et. al. (2007). n.i.: non identified. E.C. Name Reaction Gil et. al. 2004 Glass et. al. 2006 number 2.7.1.69 phosphotransferase system glc + pep → g6p + pyr PTS MG041, 069, 429 5.3.1.9 glucose-6-phosphate isomerase g6p ↔ f6p PGI MG111 2.7.1.11 6-phosphofructokinase f6p + atp → fbp + adp PFK MG215 4.1.2.13 fructose-1,6-bisphosphate aldolase fbp ↔ gdp + dhp FBA MG023 5.3.1.1 triose-phosphate isomerase gdp ↔ dhp TPI MG431 glyceraldehyde-3-phosphate gdp + nad + p ↔ bpg + 1.2.1.12 GAP MG301 dehydrogenase nadh 2.7.2.3 phosphoglycerate kinase bpg + adp ↔ 3pg + atp PGK MG300 5.4.2.1 phosphoglycerate mutase 3pg ↔ 2pg GPM MG430 4.2.1.11 enolase 2pg ↔ pep ENO MG407 2.7.1.40 pyruvate kinase pep + adp → pyr + atp PYK MG216 1.1.1.27 lactate dehydrogenase pyr + nadh ↔ lac + nad LDH MG460 1.1.1.94 sn-glycerol-3-phosphate dehydrogenase dhp + nadh → g3p + nad GPS n.i. 2.3.1.15 sn-glycerol-3-phosphate acyltransferase g3p + pal → mag PLSb n.i. 2.3.1.51 1-acyl-sn-glycerol-3-phosphate mag + pal → dag PLSc MG212 acyltransferase 2.7.7.41 phosphatidate cytidyltransferase dag + ctp → cdp-dag + pp CDS MG437 cdp-dag + ser → pser + 2.7.8.8 phosphatidylserine synthase PSS n.i. cmp 4.1.1.65 phosphatidylserine decarboxylase pser → peta PSD n.i. -

(RBKS) from Chinese Banna Mini-Pig I
African Journal of Biotechnology Vol. 11(1), pp. 46-53, 3 January, 2012 Available online at http://www.academicjournals.org/AJB DOI: 10.5897/AJB11.2885 ISSN 1684–5315 © 2012 Academic Journals Full Length Research Paper Isolation, sequence identification and tissue expression profile of a novel ribokinase gene ( RBKS ) from Chinese Banna mini-pig inbred line (BMI) Jinlong Huo 1,2,3 , Pei Wang 2,3 , Yongwang Miao 3, Hailong Huo 1,4 , Lixian Liu 4, Yangzhi Zeng 2,3 and Heng Xiao 1* 1Faculty of Life Science, Yunnan University, Kunming 650091, Yunnan, China. 2Key Laboratory of Banna Mini-pig Inbred Line of Yunnan Province, Kunming 650201, Yunnan, China. 3Faculty of Animal Science and Technology, Yunnan Agricultural University, Kunming 650201, Yunnan, China. 4Department of Husbandry and Veterinary, Yunnan Vocational and Technical College of Agriculture, Kunming 650031, China. Accepted 23 November, 2011 The complete expressed sequence tag (CDS) sequence of Banna mini-pig inbred line (BMI) ribokinase gene ( RBKS ) was amplified using the reverse transcription-polymerase chain reaction (RT-PCR) based on the conserved sequence information of the cattle or other mammals and known highly homologous swine ESTs. This novel gene was then deposited into NCBI database and assigned to accession number JF944892. Sequence analysis revealed that the BMI RBKS encodes a protein of 323 amino acids that has high homology with the ribokinase proteins of seven species: cattle (99%), horse (99%), orangutan (99%), human (89%), monkey (89%), rat (88%) and mouse (80%). The phylogenetic tree analysis revealed that the BMI RBKS gene has a closer genetic relationship with the RBKS genes of bovine and horse than with those of orangutan, human, monkey, rat and mouse.