L-Histidine Monohydrochloride Monohydrate, from Non-Animal Source Cell Culture Tested, Meets EP Testing Specifications Product
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Amino Acid Recognition by Aminoacyl-Trna Synthetases
www.nature.com/scientificreports OPEN The structural basis of the genetic code: amino acid recognition by aminoacyl‑tRNA synthetases Florian Kaiser1,2,4*, Sarah Krautwurst3,4, Sebastian Salentin1, V. Joachim Haupt1,2, Christoph Leberecht3, Sebastian Bittrich3, Dirk Labudde3 & Michael Schroeder1 Storage and directed transfer of information is the key requirement for the development of life. Yet any information stored on our genes is useless without its correct interpretation. The genetic code defnes the rule set to decode this information. Aminoacyl-tRNA synthetases are at the heart of this process. We extensively characterize how these enzymes distinguish all natural amino acids based on the computational analysis of crystallographic structure data. The results of this meta-analysis show that the correct read-out of genetic information is a delicate interplay between the composition of the binding site, non-covalent interactions, error correction mechanisms, and steric efects. One of the most profound open questions in biology is how the genetic code was established. While proteins are encoded by nucleic acid blueprints, decoding this information in turn requires proteins. Te emergence of this self-referencing system poses a chicken-or-egg dilemma and its origin is still heavily debated 1,2. Aminoacyl-tRNA synthetases (aaRSs) implement the correct assignment of amino acids to their codons and are thus inherently connected to the emergence of genetic coding. Tese enzymes link tRNA molecules with their amino acid cargo and are consequently vital for protein biosynthesis. Beside the correct recognition of tRNA features3, highly specifc non-covalent interactions in the binding sites of aaRSs are required to correctly detect the designated amino acid4–7 and to prevent errors in biosynthesis5,8. -

Development of a Phage Display Library for Discovery of Antigenic Brucella Peptides Jeffrey Williams Iowa State University
Iowa State University Capstones, Theses and Graduate Theses and Dissertations Dissertations 2018 Development of a phage display library for discovery of antigenic Brucella peptides Jeffrey Williams Iowa State University Follow this and additional works at: https://lib.dr.iastate.edu/etd Part of the Microbiology Commons Recommended Citation Williams, Jeffrey, "Development of a phage display library for discovery of antigenic Brucella peptides" (2018). Graduate Theses and Dissertations. 16896. https://lib.dr.iastate.edu/etd/16896 This Thesis is brought to you for free and open access by the Iowa State University Capstones, Theses and Dissertations at Iowa State University Digital Repository. It has been accepted for inclusion in Graduate Theses and Dissertations by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. Development of a phage display library for discovery of antigenic Brucella peptides by Jeffrey Williams A thesis submitted to the graduate faculty in partial fulfillment of the requirements for the degree of MASTER OF SCIENCE Major: Microbiology Program of Study Committee: Bryan H. Bellaire, Major Professor Steven Olsen Steven Carlson The student author, whose presentation of the scholarship herein was approved by the program of study committee, is solely responsible for the content of this thesis. The Graduate College will ensure this thesis is globally accessible and will not permit alterations after a degree is conferred. Iowa State University -

Convenient Preparation of Poly(L-Histidine) by the Direct Polymerization of L-Histidine Or Nim Benzyl-L-Histidine with Diphenylphosphoryl Azide
Polymer Journal, Vol. 13, No. 12, pp 1151-1154 (1981) SHORT COMMUNICATION Convenient Preparation of Poly(L-histidine) by the Direct Polymerization of L-Histidine or Nim_Benzyl-L-histidine with Diphenylphosphoryl Azide Takumi NARUSE, Bun-ichiro NAKAJIMA, Akihiro TSUTSUMI, and Norio NISHI Department of Polymer Science, Faculty of Science, Hokkaido University, Nishi 8-chome, Kita 10-jo, Kita-ku, Sapporo 060, Japan. (Received August 14, 1981) KEY WORDS Polymerization I Diphenylphosphoryl Azide (DPPA) I L- Histidine I N'm-Benzyl-L-histidine I Poly(L-histidine) I Poly(N'm-benzyl-L histidine) I IR Spectrum I 13C NMR Spectrum I Intrinsic Viscosity I Poly(L-histidine) is a very interesting poly(a histidine) by the polymerization of L-histidine with amino acid) as a synthetic functional polymer or as iodine-phosphonic acid esters was unsuccessful. The a model for the functional biopolymer such as simplification of the synthetic route for poly(L enzymes. It is known, however, that poly(L histidine) is still necessary. histidine) can not be prepared by the Fuchs Diphenylphosphoryl azide (DPPA) has been used Farthing method 1 which is very popular for prepar as a convenient reagent for racemization-free pep ing poly(a-amino acid)s. A synthetic route with tide synthesis, since it was synthesized by Shioiri et several steps involving the synthesis of NCA (a a!. in 1972.6 Recently, we reported that this reagent amino acid N-carboxyanhydride) by the rather can also be used successfully for the polymerization classical Leuchs method/ have been used for the of amino acids or peptides. -

Characterization of the Scavenger Cell Proteome in Mouse and Rat Liver
Biol. Chem. 2021; 402(9): 1073–1085 Martha Paluschinski, Cheng Jun Jin, Natalia Qvartskhava, Boris Görg, Marianne Wammers, Judith Lang, Karl Lang, Gereon Poschmann, Kai Stühler and Dieter Häussinger* Characterization of the scavenger cell proteome in mouse and rat liver + https://doi.org/10.1515/hsz-2021-0123 The data suggest that the population of perivenous GS Received January 25, 2021; accepted July 4, 2021; scavenger cells is heterogeneous and not uniform as previ- published online July 30, 2021 ously suggested which may reflect a functional heterogeneity, possibly relevant for liver regeneration. Abstract: The structural-functional organization of ammonia and glutamine metabolism in the liver acinus involves highly Keywords: glutaminase; glutamine synthetase; liver specialized hepatocyte subpopulations like glutamine syn- zonation; proteomics; scavenger cells. thetase (GS) expressing perivenous hepatocytes (scavenger cells). However, this cell population has not yet been char- acterized extensively regarding expression of other genes and Introduction potential subpopulations. This was investigated in the present study by proteome profiling of periportal GS-negative and There is a sophisticated structural-functional organization in perivenous GS-expressing hepatocytes from mouse and rat. the liver acinus with regard to ammonium and glutamine Apart from established markers of GS+ hepatocytes such as metabolism (Frieg et al. 2021; Gebhardt and Mecke 1983; glutamate/aspartate transporter II (GLT1) or ammonium Häussinger 1983, 1990). Periportal hepatocytes express en- transporter Rh type B (RhBG), we identified novel scavenger zymes required for urea synthesis such as the rate-controlling cell-specific proteins like basal transcription factor 3 (BTF3) enzyme carbamoylphosphate synthetase 1 (CPS1) and liver- and heat-shock protein 25 (HSP25). -

Solutions to 7.012 Problem Set 1
MIT Biology Department 7.012: Introductory Biology - Fall 2004 Instructors: Professor Eric Lander, Professor Robert A. Weinberg, Dr. Claudette Gardel Solutions to 7.012 Problem Set 1 Question 1 Bob, a student taking 7.012, looks at a long-standing puddle outside his dorm window. Curious as to what was growing in the cloudy water, he takes a sample to his TA, Brad Student. He wanted to know whether the organisms in the sample were prokaryotic or eukaryotic. a) Give an example of a prokaryotic and a eukaryotic organism. Prokaryotic: Eukaryotic: All bacteria Yeast, fungi, any animial or plant b) Using a light microscope, how could he tell the difference between a prokaryotic organism and a eukaryotic one? The resolution of the light microscope would allow you to see if the cell had a true nucleus or organelles. A cell with a true nucleus and organelles would be eukaryotic. You could also determine size, but that may not be sufficient to establish whether a cell is prokaryotic or eukaryotic. c) What additional differences exist between prokaryotic and eukaryotic organisms? Any answer from above also fine here. In addition, prokaryotic and eukaryotic organisms differ at the DNA level. Eukaryotes have more complex genomes than prokaryotes do. Question 2 A new startup company hires you to help with their product development. Your task is to find a protein that interacts with a polysaccharide. a) You find a large protein that has a single binding site for the polysaccharide cellulose. Which amino acids might you expect to find in the binding pocket of the protein? What is the strongest type of interaction possible between these amino acids and the cellulose? Cellulose is a polymer of glucose and as such has many free hydroxyl groups. -

Generate Metabolic Map Poster
Authors: Pallavi Subhraveti Ron Caspi Peter Midford Peter D Karp An online version of this diagram is available at BioCyc.org. Biosynthetic pathways are positioned in the left of the cytoplasm, degradative pathways on the right, and reactions not assigned to any pathway are in the far right of the cytoplasm. Transporters and membrane proteins are shown on the membrane. Ingrid Keseler Periplasmic (where appropriate) and extracellular reactions and proteins may also be shown. Pathways are colored according to their cellular function. Gcf_001591825Cyc: Bacillus vietnamensis NBRC 101237 Cellular Overview Connections between pathways are omitted for legibility. Anamika Kothari sn-glycerol phosphate phosphate pro phosphate phosphate phosphate thiamine molybdate D-xylose D-ribose glutathione 3-phosphate D-mannitol L-cystine L-djenkolate lanthionine α,β-trehalose phosphate phosphate [+ 3 more] α,α-trehalose predicted predicted ABC ABC FliY ThiT XylF RbsB RS10935 UgpC TreP PutP RS10200 PstB PstB RS10385 RS03335 RS20030 RS19075 transporter transporter of molybdate of phosphate α,β-trehalose 6-phosphate L-cystine D-xylose D-ribose sn-glycerol D-mannitol phosphate phosphate thiamine glutathione α α phosphate phosphate phosphate phosphate L-djenkolate 3-phosphate , -trehalose 6-phosphate pro 1-phosphate lanthionine molybdate phosphate [+ 3 more] Metabolic Regulator Amino Acid Degradation Amine and Polyamine Biosynthesis Macromolecule Modification tRNA-uridine 2-thiolation Degradation ATP biosynthesis a mature peptidoglycan a nascent β an N-terminal- -

The Microbiota-Produced N-Formyl Peptide Fmlf Promotes Obesity-Induced Glucose
Page 1 of 230 Diabetes Title: The microbiota-produced N-formyl peptide fMLF promotes obesity-induced glucose intolerance Joshua Wollam1, Matthew Riopel1, Yong-Jiang Xu1,2, Andrew M. F. Johnson1, Jachelle M. Ofrecio1, Wei Ying1, Dalila El Ouarrat1, Luisa S. Chan3, Andrew W. Han3, Nadir A. Mahmood3, Caitlin N. Ryan3, Yun Sok Lee1, Jeramie D. Watrous1,2, Mahendra D. Chordia4, Dongfeng Pan4, Mohit Jain1,2, Jerrold M. Olefsky1 * Affiliations: 1 Division of Endocrinology & Metabolism, Department of Medicine, University of California, San Diego, La Jolla, California, USA. 2 Department of Pharmacology, University of California, San Diego, La Jolla, California, USA. 3 Second Genome, Inc., South San Francisco, California, USA. 4 Department of Radiology and Medical Imaging, University of Virginia, Charlottesville, VA, USA. * Correspondence to: 858-534-2230, [email protected] Word Count: 4749 Figures: 6 Supplemental Figures: 11 Supplemental Tables: 5 1 Diabetes Publish Ahead of Print, published online April 22, 2019 Diabetes Page 2 of 230 ABSTRACT The composition of the gastrointestinal (GI) microbiota and associated metabolites changes dramatically with diet and the development of obesity. Although many correlations have been described, specific mechanistic links between these changes and glucose homeostasis remain to be defined. Here we show that blood and intestinal levels of the microbiota-produced N-formyl peptide, formyl-methionyl-leucyl-phenylalanine (fMLF), are elevated in high fat diet (HFD)- induced obese mice. Genetic or pharmacological inhibition of the N-formyl peptide receptor Fpr1 leads to increased insulin levels and improved glucose tolerance, dependent upon glucagon- like peptide-1 (GLP-1). Obese Fpr1-knockout (Fpr1-KO) mice also display an altered microbiome, exemplifying the dynamic relationship between host metabolism and microbiota. -

Impaired Amino Acid and TCA Metabolism and Cardiovascular Autonomic Neuropathy Progression in Type 1 Diabetes
Diabetes Volume 68, October 2019 2035 Impaired Amino Acid and TCA Metabolism and Cardiovascular Autonomic Neuropathy Progression in Type 1 Diabetes Anna V. Mathew,1 Mamta Jaiswal,2 Lynn Ang,2 George Michailidis,3 Subramaniam Pennathur,1,4 and Rodica Pop-Busui2 Diabetes 2019;68:2035–2044 | https://doi.org/10.2337/db19-0145 GENETICS/GENOMES/PROTEOMICS/METABOLOMICS While diabetes is characterized by hyperglycemia, nutri- Cardiovascular autonomic neuropathy (CAN) is a widely ent metabolic pathways like amino acid and tricarboxylic prevalent chronic diabetes complication that is character- acid (TCA) cycle are also profoundly perturbed. As gly- ized by impaired autonomic control of the cardiovascular cemic control alone does not prevent complications, we system (1). Although the initial prevalence of CAN in hypothesized that these metabolic disruptions are re- patients newly diagnosed with type 1 diabetes is low, later sponsible for the development and progression of di- prevalence after 15 years of diabetes increases to 35% in abetic cardiovascular autonomic neuropathy (CAN). We patients with type 1 diabetes and 60% in patients with performed standardized cardiovascular autonomic re- type 2 diabetes (1–4). CAN is an independent predictor of fl ex tests and targeted fasting plasma metabolomic chronic kidney disease progression and of cardiovascular analysis of amino acids and TCA cycle intermediates disease morbidity and mortality in patients with diabetes in subjects with type 1 diabetes and healthy control (5–8). CAN is also associated with an increased risk of subjects followed for 3 years. Forty-seven participants cardiac arrhythmias, silent myocardial ischemia, myocar- with type 1 diabetes (60% female and mean 6 SD age dial dysfunction, and sudden death (1,2,4,5,8). -

Supplementary Data TMAO Microbiome R1
BMJ Publishing Group Limited (BMJ) disclaims all liability and responsibility arising from any reliance Supplemental material placed on this supplemental material which has been supplied by the author(s) Gut Content for Supplementary Data Tables Tables Page Annotation guide for taxonomic features. 1 Supplementary Data 1a. Microbial associations with TMAO, and intakes 7 of choline and red meat. Supplementary Data 1b. Microbial associations with TMAO in 8 15 sensitivity analysis models. Supplementary Data 2a. Associations of intakes of choline and red meat, 21 or other TMAO precursors with TMAO levels, stratified by TMAO- associated abundant species, in full models and additionally adjusted for other TMAO precursors or TMAO predicting species. Supplementary Data 2b. Associations of intakes of choline and red meat, 34 or other TMAO precursors with TMAO levels, stratified by TMAO- producer status, additionally adjusted for other TMAO precursors. Supplementary Data 2c. Associations of intakes of choline and red meat, 37 or other TMAO precursors with TMAO levels, stratified by TMAO- producer status, in 6 different sensitivity analysis models. Supplementary Data 2d. Associations of intakes of red meat with HDLC 39 and HBA1c levels, stratified by TMAO-producer status or TMAO- associated abundant species. Supplementary Data 3a. Association of DNA gene clusters within 40 TMAO-associated species with plasma TMAO levels, and intakes of choline and red meat. Supplementary Data 3b. Association of transcriptions of gene clusters 46 (RNA/DNA ratio) within TMAO-associated species with plasma TMAO levels, and intakes of choline and red meat. Supplementary Data 4a. Association of DNA enzyme within TMAO- 62 associated species with plasma TMAO levels, and intakes of choline and red meat. -

Product Sheet Info
Master Clone List for NR-19279 ® Vibrio cholerae Gateway Clone Set, Recombinant in Escherichia coli, Plates 1-46 Catalog No. NR-19279 Table 1: Vibrio cholerae Gateway® Clones, Plate 1 (NR-19679) Clone ID Well ORF Locus ID Symbol Product Accession Position Length Number 174071 A02 367 VC2271 ribD riboflavin-specific deaminase NP_231902.1 174346 A03 336 VC1877 lpxK tetraacyldisaccharide 4`-kinase NP_231511.1 174354 A04 342 VC0953 holA DNA polymerase III, delta subunit NP_230600.1 174115 A05 388 VC2085 sucC succinyl-CoA synthase, beta subunit NP_231717.1 174310 A06 506 VC2400 murC UDP-N-acetylmuramate--alanine ligase NP_232030.1 174523 A07 132 VC0644 rbfA ribosome-binding factor A NP_230293.2 174632 A08 322 VC0681 ribF riboflavin kinase-FMN adenylyltransferase NP_230330.1 174930 A09 433 VC0720 phoR histidine protein kinase PhoR NP_230369.1 174953 A10 206 VC1178 conserved hypothetical protein NP_230823.1 174976 A11 213 VC2358 hypothetical protein NP_231988.1 174898 A12 369 VC0154 trmA tRNA (uracil-5-)-methyltransferase NP_229811.1 174059 B01 73 VC2098 hypothetical protein NP_231730.1 174075 B02 82 VC0561 rpsP ribosomal protein S16 NP_230212.1 174087 B03 378 VC1843 cydB-1 cytochrome d ubiquinol oxidase, subunit II NP_231477.1 174099 B04 383 VC1798 eha eha protein NP_231433.1 174294 B05 494 VC0763 GTP-binding protein NP_230412.1 174311 B06 314 VC2183 prsA ribose-phosphate pyrophosphokinase NP_231814.1 174603 B07 108 VC0675 thyA thymidylate synthase NP_230324.1 174474 B08 466 VC1297 asnS asparaginyl-tRNA synthetase NP_230942.2 174933 B09 198 -
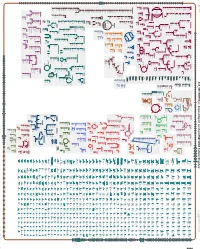
Generate Metabolic Map Poster
Authors: Pallavi Subhraveti Ron Caspi Quang Ong Peter D Karp An online version of this diagram is available at BioCyc.org. Biosynthetic pathways are positioned in the left of the cytoplasm, degradative pathways on the right, and reactions not assigned to any pathway are in the far right of the cytoplasm. Transporters and membrane proteins are shown on the membrane. Ingrid Keseler Periplasmic (where appropriate) and extracellular reactions and proteins may also be shown. Pathways are colored according to their cellular function. Gcf_900114035Cyc: Amycolatopsis sacchari DSM 44468 Cellular Overview Connections between pathways are omitted for legibility. -

Amino Acid Requirements of the Free-Living Nematode Caenorhabditis Briggsae
AMINO ACID REQUIREMENTS OF THE FREE-LIVING NEMATODE CAENORHABDITIS BRIGGSAE BY J. R. VANFLETEREN Instituut voor Dierkunde, Laboratoria voor Morfologie en Systematiek, RijksuniversiteitGent, Belgium Washed yeast ribosomes promote growth and reproduction of C. briggsae, even when supple- mented to the basal medium at dosages too low to provide the organisms with sufficient amounts of essential amino acids. Hence, a re-investigation of the amino acid requirements of C. briggsae by single and multiple omission of amino acids from the basal medium revealed unambiguously that arginine, histidine, lysine, tryptophan, phenylalanine, methionine, threonine, leucine, isoleucine and valine are not synthetized at levels to permit reproduction; they are called essential amino acids. The requirement for arginine and isoleucinehowever appears to be less clear-cut. On the contrary, evidence is presented that alanine, asparagine, cysteine, glutamate, glutamine, glycine, proline, serine and tryosine can be synthetized at adequate levels; they are called non- essential amino acids. In addition it was shown that multiple omission of the non-essential amino acids is not deleterious. This is believed to be an important step towards the development of a minimum essential medium (MEM) for growth and reproduction of C. briggsae. Sustained growth of the free-living nematode Caenorhabditis brigg.rae can be obtained on a chemically defined medium, supplemented with adequate levels of a proteinaceous growth factor. The most satisfactory, chemically defined medium hitherto reported (Buecher, Hansen & Yarwood, 1966), has been called C. brigg.iae Maintenance Medium (CbMM) and is now commercially available. CbMM is an extremely rich medium, being composed of 53 components, all present at high concentrations.