Proteases for Biocatalysis
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
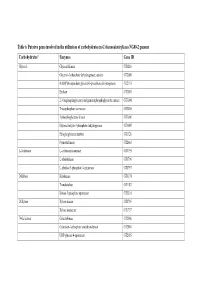
Table 6. Putative Genes Involved in the Utilization of Carbohydrates in G
Table 6. Putative genes involved in the utilization of carbohydrates in G. thermodenitrificans NG80-2 genome Carbohydrates* Enzymes Gene ID Glycerol Glycerol Kinase GT1216 Glycerol-3-phosphate dehydrogenase, aerobic GT2089 NAD(P)H-dependent glycerol-3-phosphate dehydrogenase GT2153 Enolase GT3003 2,3-bisphosphoglycerate-independentphosphoglycerate mutase GT3004 Triosephosphate isomerase GT3005 3-phosphoglycerate kinase GT3006 Glyceraldehyde-3-phosphate dehydrogenase GT3007 Phosphoglycerate mutase GT1326 Pyruvate kinase GT2663 L-Arabinose L-arabinose isomerase GT1795 L-ribulokinase GT1796 L-ribulose 5-phosphate 4-epimerase GT1797 D-Ribose Ribokinase GT3174 Transketolase GT1187 Ribose 5-phosphate epimerase GT3316 D-Xylose Xylose kinase GT1756 Xylose isomerase GT1757 D-Galactose Galactokinase GT2086 Galactose-1-phosphate uridyltransferase GT2084 UDP-glucose 4-epimerase GT2085 Carbohydrates* Enzymes Gene ID D-Fructose 1-phosphofructokinase GT1727 Fructose-1,6-bisphosphate aldolase GT1805 Fructose-1,6-bisphosphate aldolase type II GT3331 Triosephosphate isomerase GT3005 D-Mannose Mannnose-6 phospate isomelase GT3398 6-phospho-1-fructokinase GT2664 D-Mannitol Mannitol-1-phosphate dehydrogenase GT1844 N-Acetylglucosamine N-acetylglucosamine-6-phosphate deacetylase GT2205 N-acetylglucosamine-6-phosphate isomerase GT2204 D-Maltose Alpha-1,4-glucosidase GT0528, GT1643 Sucrose Sucrose phosphorylase GT3215 D-Trehalose Alpha-glucosidase GT1643 Glucose kinase GT2381 Inositol Myo-inositol catabolism protein iolC;5-dehydro-2- GT1807 deoxygluconokinase -

An Investigation on Catalysis of Acylaminoacyl Peptidases
An investigation on catalysis of acylaminoacyl peptidases PhD Thesis András László Kiss Doctorate School of Biology School leader: Prof. Anna Erdei Structural Biochemistry Programme Programme leader: Prof. László Gráf Supervisor: Prof. László Polgár Institute of Enzymology, Biological Research Center, Hungarian Academy of Sciences Budapest 2007 1. Introduction Serine peptidases contain two residues at the active site in addition to the catalytic triad (His, Asp, Ser), which form a cavity called “oxyanion hole” that accommodates the negatively charged oxyanion in the transition state of the catalysis and donate two H- bonds. Enzymes of the prolyl oligopeptidase (POP) family (acylaminoacyl peptidase, prolyl oligopeptidase, dipeptidyl peptidase IV and oligopeptidase B) are extensively studied. These enzymes are larger than classical serine peptidases and are composed of a peptidase domain with an α/β hydrolase fold and a β-propeller domain. POP and oligopeptidase B are monomeric enzymes and endopeptidases, while the exopeptidase acylaminoacyl peptidase and dipeptidyl peptidase IV are tetrameric and dimeric enzymes, respectively. Acylaminoacyl peptidase (AAP) cleaves acylated amino acids from the N-terminus of the N-acylated peptides that plays important role in many biological and disease processes. Human AAP is encoded by the DNF15S2 locus on the short arm of chromosome 3 at the region 21, which suffers deletions in small cell lung carcinomas, and renal carcinomas, resulting in deficiency in the expression of the enzyme. Acylaminoacyl peptidase is also supposed to be involved in the degradation of oxidatively damaged proteins in cells and can be associated with various diseases where damaged proteins aggregate. Its involvement in cataract formation and in the breakdown of immunogenic formylmethionyl-peptides in the digestive tract after bacterial attack was also suggested. -

High-Resolution Mass Spectrometry-Based Approaches for the Detection and Quantification of Peptidase Activity in Plasma
molecules Article High-Resolution Mass Spectrometry-Based Approaches for the Detection and Quantification of Peptidase Activity in Plasma Elisa Maffioli 1,2 , Zhenze Jiang 3, Simona Nonnis 1,2 , Armando Negri 1,2, Valentina Romeo 1, Christopher B. Lietz 3, Vivian Hook 3,4, Giuseppe Ristagno 5, Giuseppe Baselli 6, Erik B. Kistler 7,8 , Federico Aletti 9, Anthony J. O’Donoghue 3,* and Gabriella Tedeschi 1,2,* 1 Department of Veterinary Medicine, University of Milano, 20133 Milano, Italy; elisa.maffi[email protected] (E.M.); [email protected] (S.N.); [email protected] (A.N.); [email protected] (V.R.) 2 Centre for Nanostructured Materials and Interfaces (CIMAINA), University of Milano, 20133 Milano, Italy 3 Skaggs School of Pharmacy and Pharmaceutical Sciences, University of California San Diego, La Jolla, CA 92093, USA; [email protected] (Z.J.); [email protected] (C.B.L.); [email protected] (V.H.) 4 Department of Neurosciences, School of Medicine, University of California San Diego, La Jolla, CA 92093, USA 5 Department of Pathophysiology and Transplantation, University of Milan, 20133 Milan, Italy; [email protected] 6 Dipartimento di Elettronica, Informazione e Bioingegneria, Politecnico di Milano, 20133 Milan, Italy; [email protected] 7 Department of Anesthesiology & Critical Care, University of California San Diego, La Jolla, CA 92093, USA; [email protected] 8 Department of Anesthesiology & Critical Care, VA San Diego HealthCare System, San Diego, CA 92161, USA 9 Department of Bioengineering, University of California San Diego, La Jolla, CA 92093, USA; [email protected] * Correspondence: [email protected] (A.J.O.); [email protected] (G.T.); Tel.: +1-8585345360 (A.J.O.); +39-02-50318127 (G.T.) Academic Editor: Paolo Iadarola Received: 28 July 2020; Accepted: 4 September 2020; Published: 6 September 2020 Abstract: Proteomic technologies have identified 234 peptidases in plasma but little quantitative information about the proteolytic activity has been uncovered. -

And Exopeptidases in a Processing Enzyme System: Activation
Proc. Nail. Acad. Sci. USA Vol. 85, pp. 5468-5472, August 1988 Biochemistry Relationship between endo- and exopeptidases in a processing enzyme system: Activation of an endoprotease by the aminopeptidase B-like activity in somatostatin-28 convertase (brain cortex/basic amino acid pairs/peptide substrates/protease inhibitors/prohormone maturation) SOPHIE GOMEZ, PABLO GLUSCHANKOF, AGNES LEPAGE, AND PAUL COHEN Groupe de Neurobiochimie Cellulaire et Moldculaire, Universitd Pierre et Marie Curie, Unitd Associde 554 au Centre National de la Recherche Scientifique, 96 boulevard Raspail, 75006 Paris, France Communicated by I. Robert Lehman, April 8, 1988 (receivedfor review December 15, 1987) ABSTRACT The somatostatin-28 convertase activity in- somatostatin-14 and the amino-terminal dodecapeptide so- volved in vitro in the processing of somatostatin-28 into the matostatin-28-(1-12) (16). neuropeptides somatostatin-28-(1-12) and somatostatin-14 is We have described an endoprotease that cleaves the composed of an endoprotease and a basic aminopeptidase. We peptide bond on the amino side ofthe Arg-Lys doublet in the report herein on the purification to apparent homogeneity of somatostatin-28 sequence (17, 18), releasing the somato- these two constituents and on their functional interrelationship. statin-28-(1-12) fragment and [Arg-2,Lys-1]somatostatin-14. In particular we observed that after various physicochemical The released [Arg-2,Lys-']somatostatin-14 is further proc- treatments, the 90-kDa endoprotease activity was recovered essed by an aminopeptidase B-like activity (18, 19) that is also both at this molecular mass and as a 45-kDa entity. Moreover, present in the preparation. We report herein the purification the production of [Arg2,LysJllsomatostatin-14 from somato- to apparent homogeneity ofthese two activities. -

Chapter 11 Cysteine Proteases
CHAPTER 11 CYSTEINE PROTEASES ZBIGNIEW GRZONKA, FRANCISZEK KASPRZYKOWSKI AND WIESŁAW WICZK∗ Faculty of Chemistry, University of Gdansk,´ Poland ∗[email protected] 1. INTRODUCTION Cysteine proteases (CPs) are present in all living organisms. More than twenty families of cysteine proteases have been described (Barrett, 1994) many of which (e.g. papain, bromelain, ficain , animal cathepsins) are of industrial impor- tance. Recently, cysteine proteases, in particular lysosomal cathepsins, have attracted the interest of the pharmaceutical industry (Leung-Toung et al., 2002). Cathepsins are promising drug targets for many diseases such as osteoporosis, rheumatoid arthritis, arteriosclerosis, cancer, and inflammatory and autoimmune diseases. Caspases, another group of CPs, are important elements of the apoptotic machinery that regulates programmed cell death (Denault and Salvesen, 2002). Comprehensive information on CPs can be found in many excellent books and reviews (Barrett et al., 1998; Bordusa, 2002; Drauz and Waldmann, 2002; Lecaille et al., 2002; McGrath, 1999; Otto and Schirmeister, 1997). 2. STRUCTURE AND FUNCTION 2.1. Classification and Evolution Cysteine proteases (EC.3.4.22) are proteins of molecular mass about 21-30 kDa. They catalyse the hydrolysis of peptide, amide, ester, thiol ester and thiono ester bonds. The CP family can be subdivided into exopeptidases (e.g. cathepsin X, carboxypeptidase B) and endopeptidases (papain, bromelain, ficain, cathepsins). Exopeptidases cleave the peptide bond proximal to the amino or carboxy termini of the substrate, whereas endopeptidases cleave peptide bonds distant from the N- or C-termini. Cysteine proteases are divided into five clans: CA (papain-like enzymes), 181 J. Polaina and A.P. MacCabe (eds.), Industrial Enzymes, 181–195. -

Proteolytic Cleavage—Mechanisms, Function
Review Cite This: Chem. Rev. 2018, 118, 1137−1168 pubs.acs.org/CR Proteolytic CleavageMechanisms, Function, and “Omic” Approaches for a Near-Ubiquitous Posttranslational Modification Theo Klein,†,⊥ Ulrich Eckhard,†,§ Antoine Dufour,†,¶ Nestor Solis,† and Christopher M. Overall*,†,‡ † ‡ Life Sciences Institute, Department of Oral Biological and Medical Sciences, and Department of Biochemistry and Molecular Biology, University of British Columbia, Vancouver, British Columbia V6T 1Z4, Canada ABSTRACT: Proteases enzymatically hydrolyze peptide bonds in substrate proteins, resulting in a widespread, irreversible posttranslational modification of the protein’s structure and biological function. Often regarded as a mere degradative mechanism in destruction of proteins or turnover in maintaining physiological homeostasis, recent research in the field of degradomics has led to the recognition of two main yet unexpected concepts. First, that targeted, limited proteolytic cleavage events by a wide repertoire of proteases are pivotal regulators of most, if not all, physiological and pathological processes. Second, an unexpected in vivo abundance of stable cleaved proteins revealed pervasive, functionally relevant protein processing in normal and diseased tissuefrom 40 to 70% of proteins also occur in vivo as distinct stable proteoforms with undocumented N- or C- termini, meaning these proteoforms are stable functional cleavage products, most with unknown functional implications. In this Review, we discuss the structural biology aspects and mechanisms -

Intrinsic Evolutionary Constraints on Protease Structure, Enzyme
Intrinsic evolutionary constraints on protease PNAS PLUS structure, enzyme acylation, and the identity of the catalytic triad Andrew R. Buller and Craig A. Townsend1 Departments of Biophysics and Chemistry, The Johns Hopkins University, Baltimore MD 21218 Edited by David Baker, University of Washington, Seattle, WA, and approved January 11, 2013 (received for review December 6, 2012) The study of proteolysis lies at the heart of our understanding of enzyme evolution remain unanswered. Because evolution oper- biocatalysis, enzyme evolution, and drug development. To un- ates through random forces, rationalizing why a particular out- derstand the degree of natural variation in protease active sites, come occurs is a difficult challenge. For example, the hydroxyl we systematically evaluated simple active site features from all nucleophile of a Ser protease was swapped for the thiol of Cys at serine, cysteine and threonine proteases of independent lineage. least twice in evolutionary history (9). However, there is not This convergent evolutionary analysis revealed several interre- a single example of Thr naturally substituting for Ser in the lated and previously unrecognized relationships. The reactive protease catalytic triad, despite its greater chemical similarity rotamer of the nucleophile determines which neighboring amide (9). Instead, the Thr proteases generate their N-terminal nu- can be used in the local oxyanion hole. Each rotamer–oxyanion cleophile through a posttranslational modification: cis-autopro- hole combination limits the location of the moiety facilitating pro- teolysis (10, 11). These facts constitute clear evidence that there ton transfer and, combined together, fixes the stereochemistry of is a strong selective pressure against Thr in the catalytic triad that catalysis. -

Peptide Mapping (Revision 1)
001-1709PDG.pdf 1 B05 - PEPTIDE MAPPING(Revision 1) 2 3 INTRODUCTION 4 Proteins can exist as large complex structures, with some molecules in the population 5 displaying heterogeneity in their primary sequence due to improper assembly, 6 degradation or post-translational modification. The high molecular mass of proteins 7 combined with their complexity makes it particularly challenging to chemically identify an 8 intact protein product using a single analytical method. It is possible to cleave the test 9 protein into smaller fragments which can be identified with sufficient mass resolution to 10 determine the primary sequence of the protein. This process is the basis of the protein 11 identification technique commonly known as peptide mapping. The peptide mapping 12 technique involves a digestion step in which the protein is selectively cleaved at amide 13 bonds between specific amino acid residues to yield a predictable set of peptides. 14 Analytical chromatographic separation, detection, and identification of the peptide 15 mixture reveal information on the amino acid sequence of the protein which can be used 16 to identify the protein. Peptide mapping is a comparative procedure; the results from the 17 test protein are contrasted with the results of the reference standard or material similarly 18 treated to determine the identity of the test protein. This comparative identification 19 confirms that the primary structure of the test protein matches that of the reference 20 protein. 21 Peptide mapping’s ability to detect gross alterations in the primary sequence has 22 resulted in many applications for the determination of protein quality which are outside 23 the scope of this chapter. -

Expression of Proteolytic Enzymes by Small Cell Lung Cancer Circulating Tumor Cell Lines
cancers Article Expression of Proteolytic Enzymes by Small Cell Lung Cancer Circulating Tumor Cell Lines Barbara Rath 1, Lukas Klameth 2, Adelina Plangger 1, Maximilian Hochmair 3, Ernst Ulsperger 4, Ihor Huk 1, Robert Zeillinger 5 and Gerhard Hamilton 1,* 1 Department of Vascular Surgery, Medical University of Vienna, A-1090 Vienna, Austria; [email protected] (B.R.); [email protected] (A.P.); [email protected] (I.H.) 2 Center for Pathophysiology, Infectiology and Immunology, Medical University of Vienna, A-1090 Vienna, Austria; [email protected] 3 Respiratory Oncology Unit, Otto Wagner Hospital, A-1140 Vienna, Austria; [email protected] 4 Hospital Horn, A-3580 Horn, Austria; [email protected] 5 Molecular Oncology Group, Department of Obstetrics and Gynecology, Comprehensive Cancer Center-Gynecological Cancer Unit, Medical University of Vienna, A-1090 Vienna, Austria; [email protected] * Correspondence: [email protected] Received: 28 December 2018; Accepted: 17 January 2019; Published: 19 January 2019 Abstract: Small cell lung cancer (SCLC) is an aggressive type of lung cancer which disseminates vigorously and has a dismal prognosis. Metastasis of SCLC is linked to an extremely high number of circulating tumor cells (CTCs), which form chemoresistant spheroids, termed tumorospheres. Intravasation and extravasation during tumor spread requires the activity of a number of proteases to disintegrate the stroma and vascular tissue. Generation of several permanent SCLC CTC lines allowed us to screen for the expression of 35 proteases using Western blot arrays. Cell culture supernatants of two CTC lines, namely BHGc7 and 10, were analyzed for secreted proteases, including matrix metalloproteinases (MMPs), ADAM/TS, cathepsins, kallikreins, and others, and compared to proteases expressed by SCLC cell lines (GLC14, GLC16, NCI-H526 and SCLC26A). -

Cathepsin B Carboxydipeptidase Specificity Analysis Using Internally
Biochem. J. (2002) 368, 365–369 (Printed in Great Britain) 365 Cathepsin B carboxydipeptidase specificity analysis using internally quenched fluorescent peptides Maria Helena S. CEZARI, Luciano PUZER, Maria Aparecida JULIANO, Adriana K. CARMONA and Luiz JULIANO1 Department of Biophysics, Escola Paulista de Medicina, Universidade Federal de Sa4 o Paulo, Rua Tre# s de Maio, 100, Sa4 o Paulo 04044-020, Brazil We have examined in detail the specificity of the subsites S",S#, analysis of its specificity. The subsite S" accepted preferentially h h S" and S# for the carboxydipeptidase activity of cathepsin B by basic amino acids for hydrolysis; however, substrates with synthesizing and assaying four series of internally quenched phenylalanine and aliphatic side-chain-containing amino acids at #%& fluorescent peptides based on the sequence Dnp-GFRFW-OH, P" had lower Km values. Despite the presence of Glu at S#, where Dnp (2,4-dinitrophenyl) is the quenching group of the this subsite presented clear preference for aromatic amino acid fluorescence of the tryptophan residue. Each position, except the residues, and the substrate with a lysine residue at P# was h glycine, was substituted with 15 different naturally occurring hydrolysed better than that containing an arginine residue. S" is h amino acids. Based on the results we obtained, we also synthesized essentially a hydrophobic subsite, and S# has particular efficient and sensitive substrates that contained o-aminobenzoic preference for phenylalanine or tryptophan residues. acid and 3-Dnp-(2,3-diaminopropionic acid), or ε-amino-Dnp- Lys, as the fluorescence donor–receptor pair. The higher kinetic parameter values for the carboxydipeptidase compared with the Key words: fluorescent peptide, fluorogenic substrate, proteinase, endopeptidase activity of cathepsin B allowed an accurate thiol protease. -

Was the Serine Protease Cathepsin G Discovered by S. G. Hedin in 1903
Vol. 58, No 1/2011 39–44 on-line at: www.actabp.pl Regular paper Was the serine protease cathepsin G discovered by S. G. Hedin in 1903 in bovine spleen? David Palesch1, Marcin Sieńczyk2, Jozef Oleksyszyn2, Michael Reich1, Ewa Wieczerzak3, Bernhard O. Boehm1 and Timo Burster1* 1Division of Endocrinology and Diabetes, Department of Internal Medicine I, University Medical Center Ulm, Ulm, Germany; 2Wrocław University of Technology, Wrocław, Poland; 3Faculty of Chemistry, University of Gdansk, Gdańsk, Poland In the beginning of the 20th century, enzymes with pro- tive under acidic conditions, which he named b-protease teolytic activity were classified as peptidases, Erepsin, (Hedin, 1903a). and proteases. Among these, pepsin, trypsin, and auto- lytic enzymes were of the protease class. Spleen-derived After discovering these two proteases in bovine proteases were poorly characterized until Sven Gustaf spleen, Hedin performed further experiments with bo- Hedin performed several digestion experiments with vine serum and concluded that it contained a weak pro- bovine spleen. He incubated minced bovine spleen un- teolytic enzyme active in neutral medium, the activity of der acidic or neutral conditions and characterized two which could be blocked by heating at 55 °C or neutral- active proteases; the results were published in 1903. ized by antibodies (also named anti-enzymes at the time) The first protease was named α-protease and was ac- present in the serum. Hedin assumed that the α-protease tive under neutral conditions. The second was named in bovine spleen-derived leukocytes was similar to the β-protease and was active under acidic conditions. We protease found in serum. -

Multifunctional Role of Plant Cysteine Proteinases
Vol. 51 No. 3/2004 609–624 QUARTERLY Review Multifunctional role of plant cysteine proteinases Małgorzata Grudkowska1 and Barbara Zagdańska1,2½ 1Plant Physiology and Biochemistry Department, Plant Breeding and Acclimatization Institute, Radzików, Warszawa; 2Biochemistry Department, Warsaw Agricultural University, Warszawa, Poland Received: 22 March, 2004; revised: 09 July, 2004; accepted: 11 July, 2004 Key words: cysteine proteinases, localisation, inhibitors, gene expression, cellular functions Cysteine proteinases also referred to as thiol proteases play an essential role in plant growth and development but also in senescence and programmed cell death, in accumulation of storage proteins such as in seeds, but also in storage protein mobili- zation. Thus, they participate in both anabolic and catabolic processes. In addition, they are involved in signalling pathways and in the response to biotic and abiotic stresses. In this review an attempt was undertaken to illustrate these multiple roles of cysteine proteinases and the mechanisms underlying their action. Proteolysis in plants is a complex process in- they rise to 90% of the total proteolytic activ- volving many enzymes and multifarious ity (Wiśniewski & Zagdańska, 2001). They are proteolytic pathways in various cellular com- involved in protein maturation, degradation, partments, with cysteine proteinases playing and protein rebuilt in response to different an essential role. Their share in total proteol- external stimuli and they also play a ysis depends on the kind of plant and its or- house-keeping function to remove abnormal, gan. It amounts up to 30% of total proteolytic misfolded proteins. In each case, the proteoly- activity in mature non-senescing organs. sis by cysteine proteinases is a highly regu- However, the activities of cysteine protein- lated process.