Structure and Size Selection of Serine Oligopeptidases
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Molecular Markers of Serine Protease Evolution
The EMBO Journal Vol. 20 No. 12 pp. 3036±3045, 2001 Molecular markers of serine protease evolution Maxwell M.Krem and Enrico Di Cera1 ment and specialization of the catalytic architecture should correspond to signi®cant evolutionary transitions in the Department of Biochemistry and Molecular Biophysics, Washington University School of Medicine, Box 8231, St Louis, history of protease clans. Evolutionary markers encoun- MO 63110-1093, USA tered in the sequences contributing to the catalytic apparatus would thus give an account of the history of 1Corresponding author e-mail: [email protected] an enzyme family or clan and provide for comparative analysis with other families and clans. Therefore, the use The evolutionary history of serine proteases can be of sequence markers associated with active site structure accounted for by highly conserved amino acids that generates a model for protease evolution with broad form crucial structural and chemical elements of applicability and potential for extension to other classes of the catalytic apparatus. These residues display non- enzymes. random dichotomies in either amino acid choice or The ®rst report of a sequence marker associated with serine codon usage and serve as discrete markers for active site chemistry was the observation that both AGY tracking changes in the active site environment and and TCN codons were used to encode active site serines in supporting structures. These markers categorize a variety of enzyme families (Brenner, 1988). Since serine proteases of the chymotrypsin-like, subtilisin- AGY®TCN interconversion is an uncommon event, it like and a/b-hydrolase fold clans according to phylo- was reasoned that enzymes within the same family genetic lineages, and indicate the relative ages and utilizing different active site codons belonged to different order of appearance of those lineages. -

Processing of Antigenic Peptides by Aminopeptidases
June 2004 Biol. Pharm. Bull. 27(6) 777—780 (2004) 777 Current Topics Aminopeptidases in Health and Disease Processing of Antigenic Peptides by Aminopeptidases Akira HATTORI* and Masafumi TSUJIMOTO Laboratory of Cellular Biochemistry, RIKEN; 2–1 Hirosawa, Wako, Saitama 351–0198, Japan. Received January 7, 2004 Antigenic peptides presented to major histocompatibility complex (MHC) class I molecules are generated in the cytosol during degradation of cellular proteins by the ubiquitin-proteasome proteolytic pathway. Proteasome can generate N-extended precursors as well as final epitopes, and then the precursors are processed to mature epitopes by aminopeptidases. Both cytosolic peptidases (i.e. puromycin-sensitive aminopeptidase, bleomycin hy- drolase and interferon-g-inducible leucine aminopeptidase) and recently identified metallo-aminopeptidase lo- cated in the endoplasmic reticulum (i.e. adipocyte-derived leucine aminopeptidase/endoplasmic reticulum aminopeptidase 1 and leukocyte-derived arginine aminopeptidase) can generate final epitopes from precursor peptides. Some of these aminopeptidases are also considered to destroy certain antigenic peptides to limit the antigen presentation. Taken together, it is getting evident that aminopeptidases located in the cytosol and the lumen of endoplasmic reticulum play important roles in the generation of antigenic peptides presented to MHC class I molecules. Key words aminopeptidase; antigen processing; major histocompatibility complex (MHC) class I; antigen presentation; protea- some; protein degradation -

Serine Proteases with Altered Sensitivity to Activity-Modulating
(19) & (11) EP 2 045 321 A2 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 08.04.2009 Bulletin 2009/15 C12N 9/00 (2006.01) C12N 15/00 (2006.01) C12Q 1/37 (2006.01) (21) Application number: 09150549.5 (22) Date of filing: 26.05.2006 (84) Designated Contracting States: • Haupts, Ulrich AT BE BG CH CY CZ DE DK EE ES FI FR GB GR 51519 Odenthal (DE) HU IE IS IT LI LT LU LV MC NL PL PT RO SE SI • Coco, Wayne SK TR 50737 Köln (DE) •Tebbe, Jan (30) Priority: 27.05.2005 EP 05104543 50733 Köln (DE) • Votsmeier, Christian (62) Document number(s) of the earlier application(s) in 50259 Pulheim (DE) accordance with Art. 76 EPC: • Scheidig, Andreas 06763303.2 / 1 883 696 50823 Köln (DE) (71) Applicant: Direvo Biotech AG (74) Representative: von Kreisler Selting Werner 50829 Köln (DE) Patentanwälte P.O. Box 10 22 41 (72) Inventors: 50462 Köln (DE) • Koltermann, André 82057 Icking (DE) Remarks: • Kettling, Ulrich This application was filed on 14-01-2009 as a 81477 München (DE) divisional application to the application mentioned under INID code 62. (54) Serine proteases with altered sensitivity to activity-modulating substances (57) The present invention provides variants of ser- screening of the library in the presence of one or several ine proteases of the S1 class with altered sensitivity to activity-modulating substances, selection of variants with one or more activity-modulating substances. A method altered sensitivity to one or several activity-modulating for the generation of such proteases is disclosed, com- substances and isolation of those polynucleotide se- prising the provision of a protease library encoding poly- quences that encode for the selected variants. -

Enzymatic Hydrolysis and Peptide Mapping of Potato Pulp Protein
Enzymatic Hydrolysis and Peptide Mapping of Potato Pulp Protein Von der Naturwissenschaftlichen Fakultät der Universität Hannover zur Erlangung des Grades Doktorin der Naturwissenschaften Dr. rer. nat. genehmigte Dissertation von Chulaporn Kamnerdpetch, M.Sc. geboren in Bangkok, Thailand Hannover 2006 Hauptreferent Prof. Dr. Thomas Scheper Institut für Technische Chemie Universität Hannover Koreferent Prof. Dr. Bernd Hitzmann Institut für Technische Chemie Universität Hannover Tag der Promotion 29. Mai 2006 Erklärung Ich versichere, dass ich diese Dissertation selbstständig und nur unter Verwendung der angegebenen Hilfsmittel und Quellen durchgeführt habe. Diese Arbeit wurde nicht als Diplomarbeit oder ähnliche Prüfungsarbeit verwendet. Chulaporn Kamnerdpetch Hannover, den 29. Mai 2006 ACKNOWLEDGEMENTS This thesis is the result of my four years research work whereby I have been accompanied and supported by many people. It is a pleasant aspect that I have now the opportunity to express my sincere gratitude for all of them who made this thesis possible. The first person I would like to thank is my supervisor Prof. Dr. Thomas Scheper for giving me the opportunity to take part on the doctoral program at the Institut für Technische Chemie der Universität Hannover. I appreciate very much for his enthusiastic and enthusing support. He gave me an encourage independent thinking and the freedom to try out my ways. I would like to thank to Prof. Dr. Bernd Hitzmann for his kindness acceptance as my co-referee. I wish to express my thank to Dr. Cornelia Kasper for preparing my publication and proof reading. It is a great pleasure for me to thank Dr. Pichai Namparkai for proof reading as well. -

JASON MARC GOLDSTEIN the Isolation, Characterization
JASON MARC GOLDSTEIN The Isolation, Characterization and Cloning of Three Novel Peptidases From Streptoccocus gordonii: Their Potential Roles in Subacute Bacterial Endocarditis (Under the Direction of JAMES TRAVIS) Streptococcus gordonii is generally considered a benign inhabitant of the oral microflora yet is a primary etiological agent in the development of subacute bacterial endocarditis (SBE), an inflammatory state that propagates thrombus formation and tissue damage on the surface of heart valves. Colonization and adherence mechanisms have been identified, yet factors necessary to sustain growth remain unidentified. Strain FSS2 produced three extracellular aminopeptidase activities during growth in neutral pH- controlled batch cultures. The first included a serine-class dipeptidyl-aminopeptidase, an x-Pro DPP (Sg-xPDPP) found as an 85 kDa monomer by SDS-PAGE while appearing as a homodimer under native conditions. Kinetic studies indicated a unique and stringent x- Pro specificity comparable to the DPPIV/CD26 and lactococcal x-Pro DPP families. Isolation of the full-length gene uncovered a 759-amino acid polypeptide with a mass of 87,115 Da and theoretical pI of 5.6. Significant homology was found with PepX gene family members from Lactobacillus ssp. and Lactococcus ssp., and putative streptococcal x-Pro DPPs. The second activity was a putative serine-class arginine aminopeptidase (Sg- RAP) with some cysteine-class characteristics. It was found as a protein monomer of 70 kDa under denaturing conditions. Nested PCR cloning enabled the isolation of a 324 bp- long DNA fragment encoding the protein’s 108 amino acid N-terminus. Culture activity profiles and N-terminal sequence analysis indicated the release of this protein from the cell surface. -
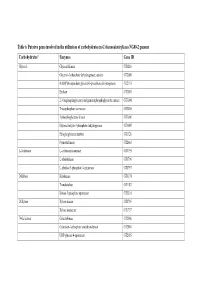
Table 6. Putative Genes Involved in the Utilization of Carbohydrates in G
Table 6. Putative genes involved in the utilization of carbohydrates in G. thermodenitrificans NG80-2 genome Carbohydrates* Enzymes Gene ID Glycerol Glycerol Kinase GT1216 Glycerol-3-phosphate dehydrogenase, aerobic GT2089 NAD(P)H-dependent glycerol-3-phosphate dehydrogenase GT2153 Enolase GT3003 2,3-bisphosphoglycerate-independentphosphoglycerate mutase GT3004 Triosephosphate isomerase GT3005 3-phosphoglycerate kinase GT3006 Glyceraldehyde-3-phosphate dehydrogenase GT3007 Phosphoglycerate mutase GT1326 Pyruvate kinase GT2663 L-Arabinose L-arabinose isomerase GT1795 L-ribulokinase GT1796 L-ribulose 5-phosphate 4-epimerase GT1797 D-Ribose Ribokinase GT3174 Transketolase GT1187 Ribose 5-phosphate epimerase GT3316 D-Xylose Xylose kinase GT1756 Xylose isomerase GT1757 D-Galactose Galactokinase GT2086 Galactose-1-phosphate uridyltransferase GT2084 UDP-glucose 4-epimerase GT2085 Carbohydrates* Enzymes Gene ID D-Fructose 1-phosphofructokinase GT1727 Fructose-1,6-bisphosphate aldolase GT1805 Fructose-1,6-bisphosphate aldolase type II GT3331 Triosephosphate isomerase GT3005 D-Mannose Mannnose-6 phospate isomelase GT3398 6-phospho-1-fructokinase GT2664 D-Mannitol Mannitol-1-phosphate dehydrogenase GT1844 N-Acetylglucosamine N-acetylglucosamine-6-phosphate deacetylase GT2205 N-acetylglucosamine-6-phosphate isomerase GT2204 D-Maltose Alpha-1,4-glucosidase GT0528, GT1643 Sucrose Sucrose phosphorylase GT3215 D-Trehalose Alpha-glucosidase GT1643 Glucose kinase GT2381 Inositol Myo-inositol catabolism protein iolC;5-dehydro-2- GT1807 deoxygluconokinase -

An Investigation on Catalysis of Acylaminoacyl Peptidases
An investigation on catalysis of acylaminoacyl peptidases PhD Thesis András László Kiss Doctorate School of Biology School leader: Prof. Anna Erdei Structural Biochemistry Programme Programme leader: Prof. László Gráf Supervisor: Prof. László Polgár Institute of Enzymology, Biological Research Center, Hungarian Academy of Sciences Budapest 2007 1. Introduction Serine peptidases contain two residues at the active site in addition to the catalytic triad (His, Asp, Ser), which form a cavity called “oxyanion hole” that accommodates the negatively charged oxyanion in the transition state of the catalysis and donate two H- bonds. Enzymes of the prolyl oligopeptidase (POP) family (acylaminoacyl peptidase, prolyl oligopeptidase, dipeptidyl peptidase IV and oligopeptidase B) are extensively studied. These enzymes are larger than classical serine peptidases and are composed of a peptidase domain with an α/β hydrolase fold and a β-propeller domain. POP and oligopeptidase B are monomeric enzymes and endopeptidases, while the exopeptidase acylaminoacyl peptidase and dipeptidyl peptidase IV are tetrameric and dimeric enzymes, respectively. Acylaminoacyl peptidase (AAP) cleaves acylated amino acids from the N-terminus of the N-acylated peptides that plays important role in many biological and disease processes. Human AAP is encoded by the DNF15S2 locus on the short arm of chromosome 3 at the region 21, which suffers deletions in small cell lung carcinomas, and renal carcinomas, resulting in deficiency in the expression of the enzyme. Acylaminoacyl peptidase is also supposed to be involved in the degradation of oxidatively damaged proteins in cells and can be associated with various diseases where damaged proteins aggregate. Its involvement in cataract formation and in the breakdown of immunogenic formylmethionyl-peptides in the digestive tract after bacterial attack was also suggested. -

High-Resolution Mass Spectrometry-Based Approaches for the Detection and Quantification of Peptidase Activity in Plasma
molecules Article High-Resolution Mass Spectrometry-Based Approaches for the Detection and Quantification of Peptidase Activity in Plasma Elisa Maffioli 1,2 , Zhenze Jiang 3, Simona Nonnis 1,2 , Armando Negri 1,2, Valentina Romeo 1, Christopher B. Lietz 3, Vivian Hook 3,4, Giuseppe Ristagno 5, Giuseppe Baselli 6, Erik B. Kistler 7,8 , Federico Aletti 9, Anthony J. O’Donoghue 3,* and Gabriella Tedeschi 1,2,* 1 Department of Veterinary Medicine, University of Milano, 20133 Milano, Italy; elisa.maffi[email protected] (E.M.); [email protected] (S.N.); [email protected] (A.N.); [email protected] (V.R.) 2 Centre for Nanostructured Materials and Interfaces (CIMAINA), University of Milano, 20133 Milano, Italy 3 Skaggs School of Pharmacy and Pharmaceutical Sciences, University of California San Diego, La Jolla, CA 92093, USA; [email protected] (Z.J.); [email protected] (C.B.L.); [email protected] (V.H.) 4 Department of Neurosciences, School of Medicine, University of California San Diego, La Jolla, CA 92093, USA 5 Department of Pathophysiology and Transplantation, University of Milan, 20133 Milan, Italy; [email protected] 6 Dipartimento di Elettronica, Informazione e Bioingegneria, Politecnico di Milano, 20133 Milan, Italy; [email protected] 7 Department of Anesthesiology & Critical Care, University of California San Diego, La Jolla, CA 92093, USA; [email protected] 8 Department of Anesthesiology & Critical Care, VA San Diego HealthCare System, San Diego, CA 92161, USA 9 Department of Bioengineering, University of California San Diego, La Jolla, CA 92093, USA; [email protected] * Correspondence: [email protected] (A.J.O.); [email protected] (G.T.); Tel.: +1-8585345360 (A.J.O.); +39-02-50318127 (G.T.) Academic Editor: Paolo Iadarola Received: 28 July 2020; Accepted: 4 September 2020; Published: 6 September 2020 Abstract: Proteomic technologies have identified 234 peptidases in plasma but little quantitative information about the proteolytic activity has been uncovered. -

(12) Patent Application Publication (10) Pub. No.: US 2006/0110747 A1 Ramseier Et Al
US 200601 10747A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2006/0110747 A1 Ramseier et al. (43) Pub. Date: May 25, 2006 (54) PROCESS FOR IMPROVED PROTEIN (60) Provisional application No. 60/591489, filed on Jul. EXPRESSION BY STRAIN ENGINEERING 26, 2004. (75) Inventors: Thomas M. Ramseier, Poway, CA Publication Classification (US); Hongfan Jin, San Diego, CA (51) Int. Cl. (US); Charles H. Squires, Poway, CA CI2O I/68 (2006.01) (US) GOIN 33/53 (2006.01) CI2N 15/74 (2006.01) Correspondence Address: (52) U.S. Cl. ................................ 435/6: 435/7.1; 435/471 KING & SPALDING LLP 118O PEACHTREE STREET (57) ABSTRACT ATLANTA, GA 30309 (US) This invention is a process for improving the production levels of recombinant proteins or peptides or improving the (73) Assignee: Dow Global Technologies Inc., Midland, level of active recombinant proteins or peptides expressed in MI (US) host cells. The invention is a process of comparing two genetic profiles of a cell that expresses a recombinant (21) Appl. No.: 11/189,375 protein and modifying the cell to change the expression of a gene product that is upregulated in response to the recom (22) Filed: Jul. 26, 2005 binant protein expression. The process can improve protein production or can improve protein quality, for example, by Related U.S. Application Data increasing solubility of a recombinant protein. Patent Application Publication May 25, 2006 Sheet 1 of 15 US 2006/0110747 A1 Figure 1 09 010909070£020\,0 10°0 Patent Application Publication May 25, 2006 Sheet 2 of 15 US 2006/0110747 A1 Figure 2 Ester sers Custer || || || || || HH-I-H 1 H4 s a cisiers TT closers | | | | | | Ya S T RXFO 1961. -

Proteasome Inhibitors: from Research Tools to Drug Candidates
Chemistry & Biology 8 (2001) 739^758 www.elsevier.com/locate/chembiol Review Proteasome inhibitors: from research tools to drug candidates Alexei F. Kisselev*, Alfred L. Goldberg Department of Cell Biology, Harvard Medical School, 240 Longwood Ave., Boston, MA 02115, USA Received 13 December 2000; revisions requested 29 March 2001; revisions received 12 June 2001; accepted 19 June 2001 First published online 12 July 2001 Abstract The 26S proteasome is a 2.4 MDa multifunctional ATP- inhibitors are now in clinical trials for treatment of multiple dependent proteolytic complex, which degrades the majority of cancers and stroke. ß 2001 Elsevier Science Ltd. All rights re- cellular polypeptides by an unusual enzyme mechanism. Several served. groups of proteasome inhibitors have been developed and are now widely used as research tools to study the role of the ubiquitin^ Keywords: Proteasome inhibitor; proteasome pathway in various cellular processes, and two Ubiquitin^proteasome pathway; Drug candidate 1. Introduction of cyclins and inhibitors of cyclin-dependent kinases [7], while degradation of transcriptional regulators, such as The ubiquitin^proteasome pathway is the major proteo- c-Jun, E2F-1 and L-catenin (see [8] for review) is essential lytic system in the cytosol and nucleus of all eukaryotic for the regulation of cell growth and gene expression. cells. This ATP-dependent pathway was discovered more Similarly, degradation by the proteasome of activated pro- than 20 years ago [1,2], but the involvement of the pro- tein kinases, e.g. src and protein kinase C [9,10], is critical teasome particle was demonstrated only in the late 1980s for the termination of certain signal transduction cascades. -

Studies of Structure and Function of Tripeptidyl-Peptidase II
Till familj och vänner List of Papers This thesis is based on the following papers, which are referred to in the text by their Roman numerals. I. Eriksson, S.; Gutiérrez, O.A.; Bjerling, P.; Tomkinson, B. (2009) De- velopment, evaluation and application of tripeptidyl-peptidase II se- quence signatures. Archives of Biochemistry and Biophysics, 484(1):39-45 II. Lindås, A-C.; Eriksson, S.; Josza, E.; Tomkinson, B. (2008) Investiga- tion of a role for Glu-331 and Glu-305 in substrate binding of tripepti- dyl-peptidase II. Biochimica et Biophysica Acta, 1784(12):1899-1907 III. Eklund, S.; Lindås, A-C.; Hamnevik, E.; Widersten, M.; Tomkinson, B. Inter-species variation in the pH dependence of tripeptidyl- peptidase II. Manuscript IV. Eklund, S.; Kalbacher, H.; Tomkinson, B. Characterization of the endopeptidase activity of tripeptidyl-peptidase II. Manuscript Paper I and II were published under maiden name (Eriksson). Reprints were made with permission from the respective publishers. Contents Introduction ..................................................................................................... 9 Enzymes ..................................................................................................... 9 Enzymes and pH dependence .............................................................. 11 Peptidases ................................................................................................. 12 Serine peptidases ................................................................................. 14 Intracellular protein -

The Role of Tricorn Protease and Its Aminopeptidase-Interacting Factors in Cellular Protein Degradation
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Cell, Vol. 95, 637±648, November 25, 1998, Copyright 1998 by Cell Press The Role of Tricorn Protease and Its Aminopeptidase-Interacting Factors in Cellular Protein Degradation Noriko Tamura, Friedrich Lottspeich, complexes that invariably contain ATPase subunits ren- Wolfgang Baumeister,* and Tomohiro Tamura dering protein degradation energy dependent. The oc- Max-Planck-Institut fuÈ r Biochemie currence of self-compartmentalizing proteases in all D-82152 Martinsried three domains of life bears testimony of an old evolution- Germany ary principle. In prokaryotic cells lacking membrane- bounded compartments, ATP-dependent self-compart- mentalizing proteases such as the proteasome, HslV, ClpP, or Lon are responsible for the bulk of the protein Summary turnover. Beyond facilitating the control of proteolysis, the con- Tricorn protease was previously described as the core finement of the proteolytic activity to a nanocompart- enzyme of a modular proteolytic system displaying ment inside these assemblies also provides the struc- multicatalytic activity. Here we elucidate the mode of tural basis for the processive mode of action that is cooperation between Tricorn and its interacting fac- characteristic for these proteases: they do not release tors, and we identify two additional factors, F2 and F3, fragments after a single cleavage, but proceed to make closely related aminopeptidases of 89 kDa. In conjunc- multiple cleavages before finally discharging the degra- tion with these three factors, Tricorn degrades oligo- dation products (Thompson et al., 1994; Akopian et al., peptides in a sequential manner, yielding free amino 1997).