Biology-Related Parameter Identification in Large Kinetic
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Line 6 POD Go Owner's Manual
® 16C Two–Plus Decades ACTION 1 VIEW Heir Stereo FX Cali Q Apparent Loop Graphic Twin Transistor Particle WAH EXP 1 PAGE PAGE Harmony Tape Verb VOL EXP 2 Time Feedback Wow/Fluttr Scale Spread C D MODE EDIT / EXIT TAP A B TUNER 1.10 OWNER'S MANUAL 40-00-0568 Rev B (For use with POD Go Firmware 1.10) ©2020 Yamaha Guitar Group, Inc. All rights reserved. 0•1 Contents Welcome to POD Go 3 The Blocks 13 Global EQ 31 Common Terminology 3 Input and Output 13 Resetting Global EQ 31 Updating POD Go to the Latest Firmware 3 Amp/Preamp 13 Global Settings 32 Top Panel 4 Cab/IR 15 Rear Panel 6 Effects 17 Restoring All Global Settings 32 Global Settings > Ins/Outs 32 Quick Start 7 Looper 22 Preset EQ 23 Global Settings > Preferences 33 Hooking It All Up 7 Wah/Volume 24 Global Settings > Switches/Pedals 33 Play View 8 FX Loop 24 Global Settings > MIDI/Tempo 34 Edit View 9 U.S. Registered Trademarks 25 USB Audio/MIDI 35 Selecting Blocks/Adjusting Parameters 9 Choosing a Block's Model 10 Snapshots 26 Hardware Monitoring vs. DAW Software Monitoring 35 Moving Blocks 10 Using Snapshots 26 DI Recording and Re-amping 35 Copying/Pasting a Block 10 Saving Snapshots 27 Core Audio Driver Settings (macOS only) 37 Preset List 11 Tips for Creative Snapshot Use 27 ASIO Driver Settings (Windows only) 37 Setlist and Preset Recall via MIDI 38 Saving/Naming a Preset 11 Bypass/Control 28 TAP Tempo 12 Snapshot Recall via MIDI 38 The Tuner 12 Quick Bypass Assign 28 MIDI CC 39 Quick Controller Assign 28 Additional Resources 40 Manual Bypass/Control Assignment 29 Clearing a Block's Assignments 29 Clearing All Assignments 30 Swapping Stomp Footswitches 30 ©2020 Yamaha Guitar Group, Inc. -

EFI Printsmith Vision Four51 Integration Guide
Four51 Integration Guide PrintSmith Vision Version 3.0 June / 2015 2 EFI PrintSmith Vision | Four51 Integration Guide Copyright © 1997 - 2015 by Electronics for Imaging, Inc. All Rights Reserved. EFI PrintSmith Vision | Four51 Integration Guide July 2015 PrintSmith Vision 3.0 This publication is protected by copyright, and all rights are reserved. No part of it may be reproduced or transmitted in any form or by any means for any purpose without express prior written consent from Electronics for Imaging, Inc. Information in this document is subject to change without notice and does not represent a commitment on the part of Electronics for Imaging, Inc. Patents This product may be covered by one or more of the following U.S. Patents: 4,716,978, 4,828,056, 4,917,488, 4,941,038, 5,109,241, 5,170,182, 5,212,546, 5,260,878, 5,276,490, 5,278,599, 5,335,040, 5,343,311, 5,398,107, 5,424,754, 5,442,429, 5,459,560, 5,467,446, 5,506,946, 5,517,334, 5,537,516, 5,543,940, 5,553,200, 5,563,689, 5,565,960, 5,583,623, 5,596,416, 5,615,314, 5,619,624, 5,625,712, 5,640,228, 5,666,436, 5,745,657, 5,760,913, 5,799,232, 5,818,645, 5,835,788, 5,859,711, 5,867,179, 5,940,186, 5,959,867, 5,970,174, 5,982,937, 5,995,724, 6,002,795, 6,025,922, 6,035,103, 6,041,200, 6,065,041, 6,112,665, 6,116,707, 6,122,407, 6,134,018, 6,141,120, 6,166,821, 6,173,286, 6,185,335, 6,201,614, 6,215,562, 6,219,155, 6,219,659, 6,222,641, 6,224,048, 6,225,974, 6,226,419, 6,238,105, 6,239,895, 6,256,108, 6,269,190, 6,271,937, 6,278,901, 6,279,009, 6,289,122, 6,292,270, 6,299,063, 6,310,697, -

Microtemporality: at the Time When Loading-In-Progress
Microtemporality: At The Time When Loading-in-progress Winnie Soon School of Communication and Culture, Aarhus University [email protected] Abstract which data processing and code inter-actions are Loading images and webpages, waiting for social media feeds operated in real-time. The notion of inter-actions mainly and streaming videos and multimedia contents have become a draws references from the notion of "interaction" from mundane activity in contemporary culture. In many situations Computer Science and the notion of "intra-actions" from nowadays, users encounter a distinctive spinning icon during Philosophy. [3][4][5] The term code inter-actions the loading, waiting and streaming of data content. A highlights the operational process of things happen graphically animated logo called throbber tells users something within, and across, machines through different technical is loading-in-progress, but nothing more. This article substrates, and hence produce agency. investigates the process of data buffering that takes place behind a running throbber. Through artistic practice, an experimental project calls The Spinning Wheel of Life explores This article is informed by artistic practice, including the temporal and computational complexity of buffering. The close reading of a throbber and its operational logics of article draws upon Wolfgang Ernst’s concept of data buffering, as well as making and coding of a “microtemporality,” in which microscopic temporality is throbber. These approaches, following the tradition of expressed through operational micro events. [1] artistic research, allow the artist/researcher to think in, Microtemporality relates to the nature of signals and through and with art. [7] Such mode of inquiry questions communications, mathematics, digital computation and the invisibility of computational culture. -

Automated Malware Analysis Report For
ID: 219320 Cookbook: browseurl.jbs Time: 00:06:44 Date: 01/04/2020 Version: 28.0.0 Lapis Lazuli Table of Contents Table of Contents 2 Analysis Report http://ib.adnxs.com 4 Overview 4 General Information 4 Detection 5 Confidence 5 Classification Spiderchart 6 Analysis Advice 6 Mitre Att&ck Matrix 7 Signature Overview 7 Networking: 7 System Summary: 7 Malware Analysis System Evasion: 8 Malware Configuration 8 Behavior Graph 8 Simulations 8 Behavior and APIs 8 Antivirus, Machine Learning and Genetic Malware Detection 8 Initial Sample 8 Dropped Files 9 Unpacked PE Files 9 Domains 9 URLs 9 Yara Overview 10 Initial Sample 10 PCAP (Network Traffic) 10 Dropped Files 10 Memory Dumps 10 Unpacked PEs 10 Sigma Overview 10 Joe Sandbox View / Context 10 IPs 10 Domains 10 ASN 10 JA3 Fingerprints 10 Dropped Files 11 Screenshots 11 Thumbnails 11 Startup 12 Created / dropped Files 12 Domains and IPs 41 Contacted Domains 41 Contacted URLs 41 URLs from Memory and Binaries 42 Contacted IPs 44 Public 44 Static File Info 45 No static file info 45 Network Behavior 45 Network Port Distribution 45 TCP Packets 45 UDP Packets 47 DNS Queries 49 DNS Answers 49 HTTP Request Dependency Graph 53 HTTP Packets 53 HTTPS Packets 54 Copyright Joe Security LLC 2020 Page 2 of 64 Code Manipulations 63 Statistics 63 Behavior 63 System Behavior 63 Analysis Process: iexplore.exe PID: 4776 Parent PID: 696 63 General 63 File Activities 63 Registry Activities 64 Analysis Process: iexplore.exe PID: 2872 Parent PID: 4776 64 General 64 File Activities 64 Registry Activities 64 Disassembly -

Index Images Download 2006 News Crack Serial Warez Full 12 Contact
index images download 2006 news crack serial warez full 12 contact about search spacer privacy 11 logo blog new 10 cgi-bin faq rss home img default 2005 products sitemap archives 1 09 links 01 08 06 2 07 login articles support 05 keygen article 04 03 help events archive 02 register en forum software downloads 3 security 13 category 4 content 14 main 15 press media templates services icons resources info profile 16 2004 18 docs contactus files features html 20 21 5 22 page 6 misc 19 partners 24 terms 2007 23 17 i 27 top 26 9 legal 30 banners xml 29 28 7 tools projects 25 0 user feed themes linux forums jobs business 8 video email books banner reviews view graphics research feedback pdf print ads modules 2003 company blank pub games copyright common site comments people aboutus product sports logos buttons english story image uploads 31 subscribe blogs atom gallery newsletter stats careers music pages publications technology calendar stories photos papers community data history arrow submit www s web library wiki header education go internet b in advertise spam a nav mail users Images members topics disclaimer store clear feeds c awards 2002 Default general pics dir signup solutions map News public doc de weblog index2 shop contacts fr homepage travel button pixel list viewtopic documents overview tips adclick contact_us movies wp-content catalog us p staff hardware wireless global screenshots apps online version directory mobile other advertising tech welcome admin t policy faqs link 2001 training releases space member static join health -

CSC245 - Web Technologies and Programming Homework Assignment 5: Baby Names
CSC245 - Web Technologies and Programming Homework Assignment 5: Baby Names This assignment tests your understanding of fetching data from files and web services using Ajax requests. You must match in appearance and behavior the following web page: Background Information: Every 10 years, the Social Security Administration provides data about the 1000 most popular boy and girl names for children born in the US, at http://www.ssa.gov/OACT/babynames/ . Your task for this assignment is to write the JavaScript code for a web page to display the baby names, popularity rankings, and meanings. You do not need to submit any HTML or CSS file. You will be provided with the HTML (names.html) and CSS (names.css) code to use. Turn in only the following file: ◦ names.Js, the JavaScript code for your Baby Names web page Data: Your program will get its data from a web service located at the following URL: https://cs.berry.edu/webdev/scripts/labs/babynames/babynames.php This web service has three “modes,” each of which provides a different type of data. Specify which type of data you want from it by changing a query parameter named type. Each query returns either plain text or JSON as its output. If you make a malformed request, such as one missing a necessary parameter, the service will respond with an HTTP error code of 400 rather than the default 200. If you request data for a name the server doesn't have data for, the service will respond with an error code of 404. (Hint: You can test queries by typing the URL of the web service, along with the appropriate query string parameters, in your web browser's address bar and seeing the result. -

HIPE Owner's Guide 1
Build 15.0.3262 HIPE Owner's Guide 1. Welcome Welcome to HIPE, the Herschel Interactive Processing Environment! This guide contains everything you need to know about the main software tool for the analysis of Herschel data. Here is what the next pages have in store for you: • Section 2 – Installing HIPE and performing basic configuration. • Section 3 – Basic concepts about HIPE. • Section 4 – Fine-tuning every aspect of HIPE with the Preferences dialogue window. • Section 5 – The Help System and its browsing and search features. • Section 6 – Extending HIPE with plug-ins. • Section 7 – Managing memory. • Section 8 – Managing running jobs. • Section 9 – Opening and running scripts. • Section 10 – Running scripts outside HIPE. • Section 11 – Editing scripts and configuring the editor. • Section 12 – Managing log messages. • Section 13 – Saving and restoring variables and sessions. • Section 14 – Managing calibration sources. • Section 15 – Viewing data products and observations. • Section 16 – Giving feedback about HIPE. • Section 17 – HIPE views. • Section 18 – HIPE perspectives. • Section 19 – Customising views and perspectives. • Section 20 – Mouse click behaviours and keyboard shortcuts. • Section 21 – Managing monospaced fonts. • Section 22 – Comparison tables to translate the most used IDL commands into HIPE. There is much more to HIPE than is contained in this manual: • If you want to learn more about graphical tools to download, visualise and analyse your data, check the Data Analysis Guide. • If you want to learn the scripting language used in HIPE and master the command line interface, have a look at the Scripting Guide. 1 HIPE Owner's Guide Build 15.0.3262 • See the Welcome page of the HIPE Help System for links to our YouTube and Twitter channels. -

Release Notes – Experience 2.35.4.En
ReDat eXperience v 2.35.4 Release notes ATC-ATM ReDat eXperience 2.35.4 Release notes Issued: 11/2019 v 2.35.4 rev. 2 Producer: RETIA, a.s. Pražská 341 Zelené Předměstí 530 02 Pardubice Czech Republic with certified system of quality control by ISO 9001 and member of AOBP The manual employs the following fonts for distinction of meaning of the text: Bold • names of programs, files, services, modules, functions, parameters, icons, database tables, formats, numbers and names of chapters in the text, paths, IP addresses. Bold, italics • names of selection items (options of combo boxes, degrees of authorization), user names, role names. LINK, REFERENCE . in an electronic form it is a functional link to the chapter. Courier, bold . source code, text from log files, text from config files. Example, demonstration. 2019 Note, hint. Warning, alert. © Copyright RETIA, a.s. a.s. RETIA, Copyright © 2 ReDat eXperience 2.35.4 Release notes Content 1. KNOWN INCOMPATIBILITIES........................................................................................................... 4 2. NEW FEATURES ................................................................................................................................. 5 2.1 INSTALLATION ........................................................................................................................................... 5 2.2 SYSTEM .................................................................................................................................................... 5 2.2.1 SMTP -

Research on the Perception of Progress Bar Distortions
Research on the Perception of Progress Bar Distortions Bachelor’s Thesis Marek Augustin Masaryk University Faculty of Informatics Brno, Fall 2018 These pages are where the Statement of an Author and the official signed assignment are located in the printed version of the document. These pages are where the Statement of an Author and the official signed assignment are located in the printed version of the document. Declaration Hereby I declare that this thesis is my original authorial work, which I have worked out on my own. All sources, references, and literature used or excerpted during elaboration of this work are properly cited and listed in complete reference to the due source. Marek Augustin Advisor: Mgr. Marek Žuži Aknowledgement I would like to thank my supervisor Mgr. Marek Žuži for all the advices, friendliness and sup- port he gave me during the whole semester I was preparing this thesis. I would also like to thank Mgr. Vojtěch Juřík for all the time he dedicated to giving me an insight to the basics of psychol- ogy research. Abstract In today’s software, the usage of progress phenomenon of “second shown progress bar” bars (of any type) is very high and users come has been discovered. across them everywhere, from their personal computers to public places, like ATMs. Some of the processes that are being visualized by the progress bars consist of multiple subpro- cesses, of which some might have a variable speed that may cause decelerations or stops of the progress bar. This work aims to broaden the research made by Harrison et al. -

L23-Animation.Pdf
1 Today’s hall-of-fame-and-shame discussion concerns the recent trend in web browsers to replace the window’s title bar with a row of tabs (as in Google Chrome, top, and the beta release of Apple Safari 4, bottom). Let’s talk about this idea from a few perspectives: - Simplicity - Visibility - Graphic design - Consistency - Affordances (what is the little triangle on the right edge of the Safari tab?) - Errors 2 Today we’re going to talk about using animation in graphical user interfaces. Some might say, based on bad experiences with the Web, that animation has no place in a usable interface. Indeed, the <blink> tag originally introduced by the Netscape browser was an abomination. And many advertisements on the Web use animation to grab your attention, which distracts you from what you’re really trying to do and makes you annoyed. So animation has gotten a bad rap in UI. Others complain that animation is just eye candy – it makes interfaces prettier, but that’s all. But neither of those viewpoints is completely fair. Used judiciously, animation can make an important contribution to the usability of an interface. And it’s not as hard to implement as you might think. We’ll talk about both design issues and implementation today. 3 Why would we want to use animation in a GUI? First, it might be an essential part of the application itself. Games and educational simulations generally have to use animation to be realistic and engaging, because they’re simulating a virtual world in which time passes and things move. -
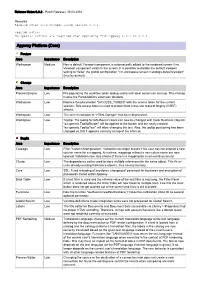
Appway 6.3.2 Release Notes
Release Notes 6.3.2 - Patch Release | 30.03.2016 Remarks Release notes with changes since version 6.3.1. Upgrade Notes: No special actions are required when upgrading from Appway 6.3.1 to 6.3.2. Appway Platform (Core) Feature Area Importance Description Workspace Medium Now a default Viewport component is automatically added to the rendered screen if no Viewport component exists in the screen. It is possible to disable the default viewport setting to "false" the global configuration "nm.workspace.screen.metatags.defaultviewport" (true by default). Change Area Importance Description Process Engine Low Pre-populating the workflow token lookup cache with local content on start-up. This change makes the PortalAddOns extension obsolete. Workspace Low Replace the placeholder "{ACCESS_TOKEN}" with the access token for the current session. This access token is used to protect from cross-site request forgery (CSRF) attacks. Workspace Low The screen component "HTML Doctype" has been deprecated. Workspace Low Tooltip: The tooltip for Info Boxes' colors can now be changed with Color Business Objects: "aw.generic.TooltipBorder" will be applied to the border and the newly created "aw.generic.TooltipText" will allow changing the text. Also, the tooltip positioning has been changed so that it appears correctly on top of the info icon. Bugfix Area Importance Description Catalogs Low Filter "Column Manipulation": Validation no longer breaks if the user has not entered a new column name for a mapping. At runtime, mappings without a new colum name are now ignored. Validation now also checks if there is a mapping for a non-existing column. -

Portal Agency Training Manual Advanced Editors Guide Drupal 7 – Georgiagov Platform
Portal Agency Training Manual Advanced Editors Guide Drupal 7 – GeorgiaGov Platform Prepared By: GeorgiaGov Interactive Support: For further assistance, fill out a Support Request at http://portal.georgia.gov/support Revised 11/03/2016 pg. 1 Table of Contents 1.0 Review 2.0 Webforms..............................................................................................................................4 2.1 Building a Webform .................................................................................................................. 4 2.2 Completing a Webform ........................................................................................................... 17 2.3 Setting Up E-mail Recipients for a Webform .......................................................................... 18 2.4 Setting Up E-mail Confirmation .............................................................................................. 19 2.4 Viewing & Exporting Results ...................................................................................................... 19 2.5 Editing a Webform .................................................................................................................. 21 2.6 Closing or Deleting a Webform ............................................................................................... 21 2.7 Convert a Webform to PDF ..................................................................................................... 22 Note: If you need a custom template for your Webform please submit a ticket