Open ALA Thesis.Pdf
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Cellular and Molecular Signatures in the Disease Tissue of Early
Cellular and Molecular Signatures in the Disease Tissue of Early Rheumatoid Arthritis Stratify Clinical Response to csDMARD-Therapy and Predict Radiographic Progression Frances Humby1,* Myles Lewis1,* Nandhini Ramamoorthi2, Jason Hackney3, Michael Barnes1, Michele Bombardieri1, Francesca Setiadi2, Stephen Kelly1, Fabiola Bene1, Maria di Cicco1, Sudeh Riahi1, Vidalba Rocher-Ros1, Nora Ng1, Ilias Lazorou1, Rebecca E. Hands1, Desiree van der Heijde4, Robert Landewé5, Annette van der Helm-van Mil4, Alberto Cauli6, Iain B. McInnes7, Christopher D. Buckley8, Ernest Choy9, Peter Taylor10, Michael J. Townsend2 & Costantino Pitzalis1 1Centre for Experimental Medicine and Rheumatology, William Harvey Research Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, Charterhouse Square, London EC1M 6BQ, UK. Departments of 2Biomarker Discovery OMNI, 3Bioinformatics and Computational Biology, Genentech Research and Early Development, South San Francisco, California 94080 USA 4Department of Rheumatology, Leiden University Medical Center, The Netherlands 5Department of Clinical Immunology & Rheumatology, Amsterdam Rheumatology & Immunology Center, Amsterdam, The Netherlands 6Rheumatology Unit, Department of Medical Sciences, Policlinico of the University of Cagliari, Cagliari, Italy 7Institute of Infection, Immunity and Inflammation, University of Glasgow, Glasgow G12 8TA, UK 8Rheumatology Research Group, Institute of Inflammation and Ageing (IIA), University of Birmingham, Birmingham B15 2WB, UK 9Institute of -

Supplementary Tables Supplemental Table S1: Comparison of the Coverage of Reference Panels Used for SNP Imputation. the Markers
Supplementary Tables Supplemental Table S1: Comparison of the coverage of reference panels used for SNP imputation. The markers on the Consortium on Asthma among African-ancestry Populations in the Americas (CAAPA) reference panel and Haplotype Reference Consortium (HRG) used to impute SNPs from our AA and CAU participants were compared to one thousand genomes (1kG) dataset. Loci information CAAPA vs 1kG HRC vs 1kG Total loci 45,639,158 90,558,388 Overlapping loci 24,880,301 49,826,569 Percent overlap 54.52% 55.02% 1kG-only loci 9,363,544 15,148,191 Ref-only loci 8,461,186 22,649,501 Supplemental Table S2: SNP imputation results. The total number of SNPs imputed for the AA and CAU participants either using the Michigan imputation server (Minimac) or Beagle. Targets prepared for: AA CAU Beagle 730,616 726,165 Minimac 698,343 660,733 Supplemental Table S3: Allele frequencies of TA2R38 SNPs by each ancestral group and time point. The rs number of each SNP, the location of SNP buy chromosome (CHR) and base pair position (POS) is provided along with the allele frequency (ALLELE:FREQ) for each SNP. Baseline 6-month Baseline 6-month AA (N = 297) AA (N = 234) CAU (N = 198) CAU (N = 151) SNP CHR POS ALLELE:FREQ ALLELE:FREQ ALLELE:FREQ ALLELE:FREQ rs10246939 7 141672604 T:0.49 C:0.51 T:0.50 C:0.50 T:0.54 C:0.46 T:0.54 C:0.46 rs1726866 7 141672705 G:0.68 A:0.32 G:0.68 A:0.32 G:0.46 A:0.54 G:0.46 A:0.54 rs713598 7 141673345 C:0.50 G:0.50 C:0.50 G:0.50 C:0.58 G:0.42 C:0.58 G:0.42 Supplemental Table S4: Linkage disequilibrium analysis of TAS2R38 SNPs at each time point of the intervention. -

G Protein-Coupled Receptors
S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2015/16: G protein-coupled receptors. British Journal of Pharmacology (2015) 172, 5744–5869 THE CONCISE GUIDE TO PHARMACOLOGY 2015/16: G protein-coupled receptors Stephen PH Alexander1, Anthony P Davenport2, Eamonn Kelly3, Neil Marrion3, John A Peters4, Helen E Benson5, Elena Faccenda5, Adam J Pawson5, Joanna L Sharman5, Christopher Southan5, Jamie A Davies5 and CGTP Collaborators 1School of Biomedical Sciences, University of Nottingham Medical School, Nottingham, NG7 2UH, UK, 2Clinical Pharmacology Unit, University of Cambridge, Cambridge, CB2 0QQ, UK, 3School of Physiology and Pharmacology, University of Bristol, Bristol, BS8 1TD, UK, 4Neuroscience Division, Medical Education Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, DD1 9SY, UK, 5Centre for Integrative Physiology, University of Edinburgh, Edinburgh, EH8 9XD, UK Abstract The Concise Guide to PHARMACOLOGY 2015/16 provides concise overviews of the key properties of over 1750 human drug targets with their pharmacology, plus links to an open access knowledgebase of drug targets and their ligands (www.guidetopharmacology.org), which provides more detailed views of target and ligand properties. The full contents can be found at http://onlinelibrary.wiley.com/doi/ 10.1111/bph.13348/full. G protein-coupled receptors are one of the eight major pharmacological targets into which the Guide is divided, with the others being: ligand-gated ion channels, voltage-gated ion channels, other ion channels, nuclear hormone receptors, catalytic receptors, enzymes and transporters. These are presented with nomenclature guidance and summary information on the best available pharmacological tools, alongside key references and suggestions for further reading. -

G Protein‐Coupled Receptors
S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2019/20: G protein-coupled receptors. British Journal of Pharmacology (2019) 176, S21–S141 THE CONCISE GUIDE TO PHARMACOLOGY 2019/20: G protein-coupled receptors Stephen PH Alexander1 , Arthur Christopoulos2 , Anthony P Davenport3 , Eamonn Kelly4, Alistair Mathie5 , John A Peters6 , Emma L Veale5 ,JaneFArmstrong7 , Elena Faccenda7 ,SimonDHarding7 ,AdamJPawson7 , Joanna L Sharman7 , Christopher Southan7 , Jamie A Davies7 and CGTP Collaborators 1School of Life Sciences, University of Nottingham Medical School, Nottingham, NG7 2UH, UK 2Monash Institute of Pharmaceutical Sciences and Department of Pharmacology, Monash University, Parkville, Victoria 3052, Australia 3Clinical Pharmacology Unit, University of Cambridge, Cambridge, CB2 0QQ, UK 4School of Physiology, Pharmacology and Neuroscience, University of Bristol, Bristol, BS8 1TD, UK 5Medway School of Pharmacy, The Universities of Greenwich and Kent at Medway, Anson Building, Central Avenue, Chatham Maritime, Chatham, Kent, ME4 4TB, UK 6Neuroscience Division, Medical Education Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, DD1 9SY, UK 7Centre for Discovery Brain Sciences, University of Edinburgh, Edinburgh, EH8 9XD, UK Abstract The Concise Guide to PHARMACOLOGY 2019/20 is the fourth in this series of biennial publications. The Concise Guide provides concise overviews of the key properties of nearly 1800 human drug targets with an emphasis on selective pharmacology (where available), plus links to the open access knowledgebase source of drug targets and their ligands (www.guidetopharmacology.org), which provides more detailed views of target and ligand properties. Although the Concise Guide represents approximately 400 pages, the material presented is substantially reduced compared to information and links presented on the website. -

Bivariate Genome-Wide Association Analysis Strengthens the Role of Bitter Receptor Clusters on Chromosomes 7 and 12 in Human Bitter Taste
bioRxiv preprint doi: https://doi.org/10.1101/296269; this version posted April 6, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Bivariate genome-wide association analysis strengthens the role of bitter receptor clusters on chromosomes 7 and 12 in human bitter taste Liang-Dar Hwang1,2,3,4, Puya Gharahkhani1, Paul A. S. Breslin5,6, Scott D. Gordon1, Gu Zhu1, Nicholas G. Martin1, Danielle R. Reed5, and Margaret J. Wright2,7 1 QIMR Berghofer Medical Research Institute, Herston, Queensland 4006, Australia 2 Queensland Brain Institute, University of Queensland, St Lucia, Queensland 4072, Australia 3 Faculty of Medicine, University of Queensland, Herston, Queensland 4006, Australia 4 University of Queensland Diamantina Institute, University of Queensland, Translational Research Institute, Woolloongabba, Queensland 4102, Australia 5 Monell Chemical Senses Center, Philadelphia, Pennsylvania 19104, USA 6 Department of Nutritional Sciences, School of Environmental and Biological Sciences, Rutgers University, New Brunswick NJ, 08901 USA 7 Centre for Advanced Imaging, University of Queensland, St Lucia, Queensland 4072, Australia Correspondence to be sent to: Liang-Dar Hwang University of Queensland Diamantina Institute Wolloongabba QLD 4102, Australia Email: [email protected] Telephone: +61 7 3443 7976 Fax: +61 7 3443 6966 1 bioRxiv preprint doi: https://doi.org/10.1101/296269; this version posted April 6, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

Transcriptome Analysis Reveals Upregulation of Bitter Taste Receptors in Severe Asthmatics
ORIGINAL ARTICLE ASTHMA Transcriptome analysis reveals upregulation of bitter taste receptors in severe asthmatics Christina Orsmark-Pietras1,2, Anna James2,3, Jon R. Konradsen2,4,5, Bjo¨rn Nordlund2,4,5, Cilla So¨derha¨ll1,2, Ville Pulkkinen2,3,6,7, Christophe Pedroletti2,5, Kameran Daham2,8,9, Maciek Kupczyk2,3, Barbro Dahle´n2,8,9, Juha Kere1,2,10, Sven-Erik Dahle´n2,3, Gunilla Hedlin2,4,5,11 and Erik Mele´n2,3,4,11 Affiliations: 1Dept of Biosciences and Nutrition at Novum, Karolinska Institute, Stockholm, 2The Centre for Allergy Research at Karolinska Institute, Stockholm, 3Institute of Environmental Medicine, Karolinska Institute, Stockholm, 4Astrid Lindgren Children’s Hospital, Karolinska University Hospital, Stockholm, 5Dept of Women’s and Children’s Health, Karolinska Institute, Stockholm, 8Lung/Allergy Clinic, Karolinska University Hospital Huddinge, Stockholm, 9Dept of Internal Medicine, Karolinska University Hospital Huddinge, Stockholm, and 10Science for Life Laboratory, Karolinska Institute, Stockholm, Sweden. 6Dept of Medical Genetics, Biomedicum Helsinki, University of Helsinki, Helsinki, and 7Folkha¨lsan Institute of Genetics, Helsinki, Finland. 11Both authors contributed equally. Correspondence: S-E. Dahle´n, The National Institute of Environmental Medicine, Karolinska Institutet, SE- 17177 Stockholm, Sweden. E-mail: [email protected] ABSTRACT The causes of severe childhood asthma are poorly understood. Our aim was to define global patterns of gene expression in children with severe therapy-resistant and controlled asthma. White blood cells were isolated and the global transcriptome profile was characterised using the Affymetrix Human Gene ST 1.0 chip in children with severe, therapy-resistant asthma (n517), controlled asthma (n519) and healthy controls (n518). -

The Bitter Taste Receptor Tas2r14 Is Expressed in Ovarian Cancer and Mediates Apoptotic Signalling
THE BITTER TASTE RECEPTOR TAS2R14 IS EXPRESSED IN OVARIAN CANCER AND MEDIATES APOPTOTIC SIGNALLING by Louis T. P. Martin Submitted in partial fulfilment of the requirements for the degree of Master of Science at Dalhousie University Halifax, Nova Scotia June 2017 © Copyright by Louis T. P. Martin, 2017 DEDICATION PAGE To my grandparents, Christina, Frank, Brenda and Bernie, and my parents, Angela and Tom – for teaching me the value of hard work. ii TABLE OF CONTENTS LIST OF TABLES ............................................................................................................. vi LIST OF FIGURES .......................................................................................................... vii ABSTRACT ....................................................................................................................... ix LIST OF ABBREVIATIONS AND SYMBOLS USED .................................................... x ACKNOWLEDGEMENTS .............................................................................................. xii CHAPTER 1 INTRODUCTION ........................................................................................ 1 1.1 G-PROTEIN COUPLED RECEPTORS ................................................................ 1 1.2 GPCR CLASSES .................................................................................................... 4 1.3 GPCR SIGNALING THROUGH G PROTEINS ................................................... 6 1.4 BITTER TASTE RECEPTORS (TAS2RS) ........................................................... -

Spezifische Seroreaktivitätsmuster Zur Minimal-Invasiven Detektion
Aus dem Institut für Humangenetik Theoretische Medizin und Biowissenschaften der Medizinischen Fakultät der Universität des Saarlandes, Homburg/Saar Spezifische Seroreaktivitätsmuster zur minimal-invasiven Detektion humaner Hirntumoren Dissertation zur Erlangung des Grades eines Doktors der Naturwissenschaften der Medizinischen Fakultät der UNIVERSITÄT DES SAARLANDES 2009 vorgelegt von Nicole Ludwig geb. am 08.03.1981 in Püttlingen Zusammenfassung i Zusammenfassung Mithilfe eines serologischen Spotassays bestehend aus über 60 bereits identifizierten Meningeom- bzw. Gliom-assoziierten Antigenen wurden Seroreaktivitätsmuster von über 400 Patienten aufgenommen und mit statistischen Methoden differenziert. Das Expressionsprofil von 24 Meningeomen wurde mithilfe von cDNA-Microarrays ermittelt, welche mit ca. 50.000 Transkripten den Großteil des bekannten menschlichen Transkriptoms repräsentieren. Es wurden über 1800 weitere Tumor-assoziierte Antigene identifiziert und zu einem Proteinmakroarray zusammengeführt. Basierend auf dieser Plattform wurden 110 Patientenseren und 60 Kontrollen auf Autoantikörper hin getestet. • Es wurden komplexe Seroreaktivitätsmuster für Meningeom- und Gliompatienten etabliert. • Unter Verwendung von statistischen Lernverfahren war eine Unterscheidung der Meningeom- bzw. Gliompatienten von Gesunden anhand ihres Seroreaktivitäts- musters in Bezug auf über 60 Hirntumor-assoziierte Antigene mit einer Klassifikationsgenauigkeit von über 90 % möglich. • Auch mit dem vergrößerten Set von über 1800 Tumor-assoziierten Antigenen -

The Hypothalamus As a Hub for SARS-Cov-2 Brain Infection and Pathogenesis
bioRxiv preprint doi: https://doi.org/10.1101/2020.06.08.139329; this version posted June 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. The hypothalamus as a hub for SARS-CoV-2 brain infection and pathogenesis Sreekala Nampoothiri1,2#, Florent Sauve1,2#, Gaëtan Ternier1,2ƒ, Daniela Fernandois1,2 ƒ, Caio Coelho1,2, Monica ImBernon1,2, Eleonora Deligia1,2, Romain PerBet1, Vincent Florent1,2,3, Marc Baroncini1,2, Florence Pasquier1,4, François Trottein5, Claude-Alain Maurage1,2, Virginie Mattot1,2‡, Paolo GiacoBini1,2‡, S. Rasika1,2‡*, Vincent Prevot1,2‡* 1 Univ. Lille, Inserm, CHU Lille, Lille Neuroscience & Cognition, DistAlz, UMR-S 1172, Lille, France 2 LaBoratorY of Development and PlasticitY of the Neuroendocrine Brain, FHU 1000 daYs for health, EGID, School of Medicine, Lille, France 3 Nutrition, Arras General Hospital, Arras, France 4 Centre mémoire ressources et recherche, CHU Lille, LiCEND, Lille, France 5 Univ. Lille, CNRS, INSERM, CHU Lille, Institut Pasteur de Lille, U1019 - UMR 8204 - CIIL - Center for Infection and ImmunitY of Lille (CIIL), Lille, France. # and ƒ These authors contriButed equallY to this work. ‡ These authors directed this work *Correspondence to: [email protected] and [email protected] Short title: Covid-19: the hypothalamic hypothesis 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.06.08.139329; this version posted June 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -
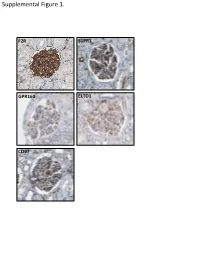
Supplemental Data
Supplemental Figure 1. F2R S1PR1 GPR160 ELTD1 CD97 Supplemental Figure 2. A B brain heart lung liver kidney spleen testis skm Expression in mouse ssues glom rok Gprc5a Gapdh Human Protein Atlas RNAseq database Supplemental Figure 3. A Gprc5a Pdgfrb Merged CL CL Gprc5a CD31 Merged CL CL B C 1.4 * 1.2 mc 1 matrix 0.8 0.6 0.4 matrix 0.2 0 pod end 1 2 mes3 D Vector Gprc5a E 40 kD - - Gprc5a - acn Supplemental Figure 4. Control 12 month-old KO 12 month-old A B C D E F 1 2 0.8 * 1.5 score 0.6 m µ 1 0.4 0.2 Slits/ 0.5 Mesangial 0 0 Ctrl KO Ctrl KO Supplemental Figure 5. A B 1.2 Vector Gprc5a 1 * 0.8 0.6 * 0.4 * Normalized Density 0.2 0 pEGFR/tEGR pSmad/tSmad TGF-β1 C 1.8 siCON 1.6 siGprc5a * 1.4 * * 1.2 1 0.8 0.6 NormalizedDensity 0.4 0.2 0 pEGFR/tEGR pSmad/tSmad TGF-β1 Supplemental table 1. List of glomerulus-expressed GPCRs as detected by qPCR. Data shown as mean ± standard deviation (Glom=glomerulus, Rok=rest of kidney). GPCR Glom Rok Glom/Rok LPAR6 41892,11 ± 38478,89 1040,06 ± 1370,12 39,28 ELTD1 30275,64 ± 14085,26 33,23 ± 46,99 910,13 GPR116 24020,06 ± 7789,84 55,1 ± 47,75 434,96 PTH1R 15402,81 ± 17644,32 7521,01 ± 3264,57 1,05 CALCRL 14096,09 ± 3854,84 199,06 ± 222,52 69,81 HPRT1 13342,04 ± 10677,69 1824,77 ± 1767,23 6,31 S1PR5 9474,7 ± 10124,3 110,84 ± 29,54 84,48 LPHN2 8645,89 ± 914,74 256,04 ± 293,87 32,77 FZD1 8176,2 ± 4947,45 1321,97 ± 1311,91 5,18 CXCR4 7097,31 ± 4388,91 535,98 ± 640,28 12,24 GPR160 6446,59 ± 1550,07 4816,3 ± 5918,99 0,34 NPY1R 6177,74 ± 6282,76 209,64 ± 296,48 28,47 PTGER4 5323,66 ± 3789,51 179,13 ± 210 28,72 RXFP1 -

Anhang Tabelle D1: Differenziell Exprimierte Gene in Meningeomen
Anhang Tabelle D1: Differenziell exprimierte Gene in Meningeomen im Vergleich zu Dura-Kontrollen Expression in Meningeomen im Vergleich zu Spot ID Gene IDSymbol Genname Chrom. LokalisationDura Faktor GE61423 3347 HTN3 histatin 3 4q13 überexprimiert 62,6162314 GE546433 119395 FAM26A family with sequence similarity 26, member A 10q24.33 überexprimiert 31,6272538 GE57732 999 CDH1 cadherin 1, type 1, E-cadherin (epithelial) 16q22.1 überexprimiert 30,2402589 GE53152 9167 COX7A2L cytochrome c oxidase subunit VIIa polypeptide 2 like 2p21 überexprimiert 29,9416437 GE79185 10551 AGR2 anterior gradient 2 homolog (Xenopus laevis) 7p21.3 überexprimiert 28,7583579 collagen, type II, alpha 1 (primary osteoarthritis, GE57600 1280 COL2A1 spondyloepiphyseal dysplasia, congenital) 12q13.11-q13.2 überexprimiert 28,577144 GE81679 10804 GJB6 gap junction protein, beta 6 (connexin 30) 13q11-q12.1 überexprimiert 28,1127885 GE88765 51200 CPA4 carboxypeptidase A4 7q32 überexprimiert 27,9914458 GE79214 195814 9q22.32 überexprimiert 23,1302115 GE53364 23308 ICOSLG inducible T-cell co-stimulator ligand 21q22.3 überexprimiert 21,3201887 solute carrier family 26 (sulfate transporter), member GE661107 1836 SLC26A2 2 5q31-q34 überexprimiert 20,9918216 GE82475 56245 C21orf62 chromosome 21 open reading frame 62 21q22.1 überexprimiert 20,7144411 GE80625 23632 CA14 carbonic anhydrase XIV 1q21 überexprimiert 19,6290117 GE88282 10804 GJB6 gap junction protein, beta 6 (connexin 30) 13q11-q12.1 überexprimiert 18,6854418 GE80611 5145 PDE6A phosphodiesterase 6A, cGMP-specific, -

G Protein‐Coupled Receptors
S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2019/20: G protein-coupled receptors. British Journal of Pharmacology (2019) 176, S21–S141 THE CONCISE GUIDE TO PHARMACOLOGY 2019/20: G protein-coupled receptors Stephen PH Alexander1 , Arthur Christopoulos2 , Anthony P Davenport3 , Eamonn Kelly4, Alistair Mathie5 , John A Peters6 , Emma L Veale5 ,JaneFArmstrong7 , Elena Faccenda7 ,SimonDHarding7 ,AdamJPawson7 , Joanna L Sharman7 , Christopher Southan7 , Jamie A Davies7 and CGTP Collaborators 1School of Life Sciences, University of Nottingham Medical School, Nottingham, NG7 2UH, UK 2Monash Institute of Pharmaceutical Sciences and Department of Pharmacology, Monash University, Parkville, Victoria 3052, Australia 3Clinical Pharmacology Unit, University of Cambridge, Cambridge, CB2 0QQ, UK 4School of Physiology, Pharmacology and Neuroscience, University of Bristol, Bristol, BS8 1TD, UK 5Medway School of Pharmacy, The Universities of Greenwich and Kent at Medway, Anson Building, Central Avenue, Chatham Maritime, Chatham, Kent, ME4 4TB, UK 6Neuroscience Division, Medical Education Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, DD1 9SY, UK 7Centre for Discovery Brain Sciences, University of Edinburgh, Edinburgh, EH8 9XD, UK Abstract The Concise Guide to PHARMACOLOGY 2019/20 is the fourth in this series of biennial publications. The Concise Guide provides concise overviews of the key properties of nearly 1800 human drug targets with an emphasis on selective pharmacology (where available), plus links to the open access knowledgebase source of drug targets and their ligands (www.guidetopharmacology.org), which provides more detailed views of target and ligand properties. Although the Concise Guide represents approximately 400 pages, the material presented is substantially reduced compared to information and links presented on the website.