Fusarium Genomics, Molecular and Cellular Biology
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

METACYC ID Description A0AR23 GO:0004842 (Ubiquitin-Protein Ligase
Electronic Supplementary Material (ESI) for Integrative Biology This journal is © The Royal Society of Chemistry 2012 Heat Stress Responsive Zostera marina Genes, Southern Population (α=0. -

Mapping Active Site Residues in Glutamate-5-Kinase. the Substrate Glutamate and the Feed-Back Inhibitor Proline Bind at Overlapping Sites
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector FEBS Letters 580 (2006) 6247–6253 Mapping active site residues in glutamate-5-kinase. The substrate glutamate and the feed-back inhibitor proline bind at overlapping sites Isabel Pe´rez-Arellanoa, Vicente Rubiob, Javier Cerveraa,* a Centro de Investigacio´n Prı´ncipe Felipe, Avda. Autopista del Saler, 16, Valencia 46013, Spain b Instituto de Biomedicina de Valencia (IBV-CSIC), Jaime Roig 11, Valencia 46010, Spain Received 22 August 2006; revised 10 October 2006; accepted 12 October 2006 Available online 20 October 2006 Edited by Stuart Ferguson (a domain named after pseudo uridine synthases and archaeo- Abstract Glutamate-5-kinase (G5K) catalyzes the controlling first step of proline biosynthesis. Substrate binding, catalysis sine-specific transglycosylases) domain [10]. AAK domains and feed-back inhibition by proline are functions of the N-termi- bind ATP and a phosphorylatable acidic substrate, and make nal 260-residue domain of G5K. We study here the impact on a phosphoric anhydride [11]. The function of the PUA domain these functions of 14 site-directed mutations affecting 9 residues (a domain characteristically found in some RNA-modifying of Escherichia coli G5K, chosen on the basis of the structure of enzymes) [12] is not known. By deleting the PUA domain of the bisubstrate complex of the homologous enzyme acetylgluta- E. coli G5K we showed [10] that the isolated AAK domain, mate kinase (NAGK). The results support the predicted roles as the complete enzyme, forms tetramers, catalyzes the reac- of K10, K217 and T169 in catalysis and ATP binding and of tion and is feed-back inhibited by proline. -

Yeast Genome Gazetteer P35-65
gazetteer Metabolism 35 tRNA modification mitochondrial transport amino-acid metabolism other tRNA-transcription activities vesicular transport (Golgi network, etc.) nitrogen and sulphur metabolism mRNA synthesis peroxisomal transport nucleotide metabolism mRNA processing (splicing) vacuolar transport phosphate metabolism mRNA processing (5’-end, 3’-end processing extracellular transport carbohydrate metabolism and mRNA degradation) cellular import lipid, fatty-acid and sterol metabolism other mRNA-transcription activities other intracellular-transport activities biosynthesis of vitamins, cofactors and RNA transport prosthetic groups other transcription activities Cellular organization and biogenesis 54 ionic homeostasis organization and biogenesis of cell wall and Protein synthesis 48 plasma membrane Energy 40 ribosomal proteins organization and biogenesis of glycolysis translation (initiation,elongation and cytoskeleton gluconeogenesis termination) organization and biogenesis of endoplasmic pentose-phosphate pathway translational control reticulum and Golgi tricarboxylic-acid pathway tRNA synthetases organization and biogenesis of chromosome respiration other protein-synthesis activities structure fermentation mitochondrial organization and biogenesis metabolism of energy reserves (glycogen Protein destination 49 peroxisomal organization and biogenesis and trehalose) protein folding and stabilization endosomal organization and biogenesis other energy-generation activities protein targeting, sorting and translocation vacuolar and lysosomal -

ONLINE SUPPLEMENTARY TABLE Table 2. Differentially Expressed
ONLINE SUPPLEMENTARY TABLE Table 2. Differentially Expressed Probe Sets in Livers of GK Rats. A. Immune/Inflammatory (67 probe sets, 63 genes) Age Strain Probe ID Gene Name Symbol Accession Gene Function 5 WKY 1398390_at small inducible cytokine B13 precursor Cxcl13 AA892854 chemokine activity; lymph node development 5 WKY 1389581_at interleukin 33 Il33 BF390510 cytokine activity 5 WKY *1373970_at interleukin 33 Il33 AI716248 cytokine activity 5 WKY 1369171_at macrophage stimulating 1 (hepatocyte growth factor-like) Mst1; E2F2 NM_024352 serine-throenine kinase; tumor suppression 5 WKY 1388071_x_at major histocompatability antigen Mhc M24024 antigen processing and presentation 5 WKY 1385465_at sialic acid binding Ig-like lectin 5 Siglec5 BG379188 sialic acid-recognizing receptor 5 WKY 1393108_at major histocompatability antigen Mhc BM387813 antigen processing and presentation 5 WKY 1388202_at major histocompatability antigen Mhc BI395698 antigen processing and presentation 5 WKY 1371171_at major histocompatability antigen Mhc M10094 antigen processing and presentation 5 WKY 1370382_at major histocompatability antigen Mhc BI279526 antigen processing and presentation 5 WKY 1371033_at major histocompatability antigen Mhc AI715202 antigen processing and presentation 5 WKY 1383991_at leucine rich repeat containing 8 family, member E Lrrc8e BE096426 proliferation and activation of lymphocytes and monocytes. 5 WKY 1383046_at complement component factor H Cfh; Fh AA957258 regulation of complement cascade 4 WKY 1369522_a_at CD244 natural killer -
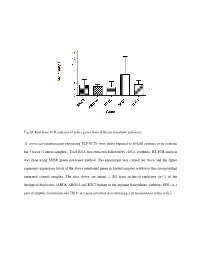
Fig.S1 Real-Time PCR Analysis of Select Genes from Different Metabolic Pathways S. Cerevisiae Transformants Expressing TEF-YCT1
Fig.S1 Real-time PCR analysis of select genes from different metabolic pathways S. cerevisiae transformants expressing TEF-YCT1 were either exposed to 500µM cysteine or no cysteine for 5 hours (Control samples). Total RNA was extracted followed by cDNA synthesis. RT-PCR analysis was done using SYBR green dye-based method. The experiment was carried out twice and the figure represents expression levels of the above mentioned genes in treated samples relative to the corresponding untreated control samples. The data above are means ± SD from technical replicates (n=3) of the biological duplicates. [ARG8, ARG5,6 and RTC2 belong to the arginine biosynthetic pathway, SSU1 is a part of sulphur metabolism and TIS11 is a gene involved in maintaining iron homeostasis in the cells.] Table S1: List of strains used in this study Strain Genotype Source ABC 1738 (yct1) MATα his31 leu20 lys20 ura30 YLL055W:: kanMX4 Euroscarf (Y01543) ABC3612(arg3) MATa his31 leu20 ura30 YJL088w::kanMX4 Euroscarf (Y11335) ABC4812(arg5,6) MATa his31 leu20 ura30 YER069w::kanMX4 Euroscarf (Y10209) ABC4811(arg8) MATa his31 leu20 ura30 YOL140w::kanMX4 Euroscarf (Y17711) ABC4814(spe1) MATa his31 leu20 ura30 YKL184w::kanMX4 Euroscarf (Y15034) ABC4815(spe2) MATa his31 leu20 ura30 YOL052c::kanMX4 Euroscarf (Y11743) ABC4816(spe3) MATa his31 leu20 ura30 YPR069c::kanMX4 Euroscarf (Y15488) ABC4817(spe4) MATa his31 leu20 ura30 YLR146c::kanMX4 Euroscarf (Y16945) ABC3604(ssu1) MATa his31 leu20 ura30 YPL092w::kanMX4 Euroscarf (Y12160) ABC1486(str2) MATa his31 leu20 ura30 YJR130c::kanMX4 -

Relating Metatranscriptomic Profiles to the Micropollutant
1 Relating Metatranscriptomic Profiles to the 2 Micropollutant Biotransformation Potential of 3 Complex Microbial Communities 4 5 Supporting Information 6 7 Stefan Achermann,1,2 Cresten B. Mansfeldt,1 Marcel Müller,1,3 David R. Johnson,1 Kathrin 8 Fenner*,1,2,4 9 1Eawag, Swiss Federal Institute of Aquatic Science and Technology, 8600 Dübendorf, 10 Switzerland. 2Institute of Biogeochemistry and Pollutant Dynamics, ETH Zürich, 8092 11 Zürich, Switzerland. 3Institute of Atmospheric and Climate Science, ETH Zürich, 8092 12 Zürich, Switzerland. 4Department of Chemistry, University of Zürich, 8057 Zürich, 13 Switzerland. 14 *Corresponding author (email: [email protected] ) 15 S.A and C.B.M contributed equally to this work. 16 17 18 19 20 21 This supporting information (SI) is organized in 4 sections (S1-S4) with a total of 10 pages and 22 comprises 7 figures (Figure S1-S7) and 4 tables (Table S1-S4). 23 24 25 S1 26 S1 Data normalization 27 28 29 30 Figure S1. Relative fractions of gene transcripts originating from eukaryotes and bacteria. 31 32 33 Table S1. Relative standard deviation (RSD) for commonly used reference genes across all 34 samples (n=12). EC number mean fraction bacteria (%) RSD (%) RSD bacteria (%) RSD eukaryotes (%) 2.7.7.6 (RNAP) 80 16 6 nda 5.99.1.2 (DNA topoisomerase) 90 11 9 nda 5.99.1.3 (DNA gyrase) 92 16 10 nda 1.2.1.12 (GAPDH) 37 39 6 32 35 and indicates not determined. 36 37 38 39 S2 40 S2 Nitrile hydration 41 42 43 44 Figure S2: Pearson correlation coefficients r for rate constants of bromoxynil and acetamiprid with 45 gene transcripts of ECs describing nucleophilic reactions of water with nitriles. -

Letters to Nature
letters to nature Received 7 July; accepted 21 September 1998. 26. Tronrud, D. E. Conjugate-direction minimization: an improved method for the re®nement of macromolecules. Acta Crystallogr. A 48, 912±916 (1992). 1. Dalbey, R. E., Lively, M. O., Bron, S. & van Dijl, J. M. The chemistry and enzymology of the type 1 27. Wolfe, P. B., Wickner, W. & Goodman, J. M. Sequence of the leader peptidase gene of Escherichia coli signal peptidases. Protein Sci. 6, 1129±1138 (1997). and the orientation of leader peptidase in the bacterial envelope. J. Biol. Chem. 258, 12073±12080 2. Kuo, D. W. et al. Escherichia coli leader peptidase: production of an active form lacking a requirement (1983). for detergent and development of peptide substrates. Arch. Biochem. Biophys. 303, 274±280 (1993). 28. Kraulis, P.G. Molscript: a program to produce both detailed and schematic plots of protein structures. 3. Tschantz, W. R. et al. Characterization of a soluble, catalytically active form of Escherichia coli leader J. Appl. Crystallogr. 24, 946±950 (1991). peptidase: requirement of detergent or phospholipid for optimal activity. Biochemistry 34, 3935±3941 29. Nicholls, A., Sharp, K. A. & Honig, B. Protein folding and association: insights from the interfacial and (1995). the thermodynamic properties of hydrocarbons. Proteins Struct. Funct. Genet. 11, 281±296 (1991). 4. Allsop, A. E. et al.inAnti-Infectives, Recent Advances in Chemistry and Structure-Activity Relationships 30. Meritt, E. A. & Bacon, D. J. Raster3D: photorealistic molecular graphics. Methods Enzymol. 277, 505± (eds Bently, P. H. & O'Hanlon, P. J.) 61±72 (R. Soc. Chem., Cambridge, 1997). -

Saccharomyces Cerevisiae Aspartate Kinase Mechanism and Inhibition
In compliance with the Canadian Privacy Legislation some supporting forms may have been removed from this dissertation. While these forms may be included in the document page count, their removal does not represent any loss of content from the dissertation. Ph.D. Thesis - D. Bareich McMaster University - Department of Biochemistry FUNGAL ASPARTATE KINASE MECHANISM AND INHIBITION By DAVID C. BAREICH, B.Sc. A Thesis Submitted to the School of Graduate Studies in Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy McMaster University © Copyright by David C. Bareich, June 2003 1 Ph.D. Thesis - D. Bareich McMaster University - Department of Biochemistry FUNGAL ASPARTATE KINASE MECHANISM AND INHIBITION Ph.D. Thesis - D. Bareich McMaster University - Department of Biochemistry DOCTOR OF PHILOSOPHY (2003) McMaster University (Biochemistry) Hamilton, Ontario TITLE: Saccharomyces cerevisiae aspartate kinase mechanism and inhibition AUTHOR: David Christopher Bareich B.Sc. (University of Waterloo) SUPERVISOR: Professor Gerard D. Wright NUMBER OF PAGES: xix, 181 11 Ph.D. Thesis - D. Bareich McMaster University - Department of Biochemistry ABSTRACT Aspartate kinase (AK) from Saccharomyces cerevisiae (AKsc) catalyzes the first step in the aspartate pathway responsible for biosynthesis of L-threonine, L-isoleucine, and L-methionine in fungi. Little was known about amino acids important for AKsc substrate binding and catalysis. Hypotheses about important amino acids were tested using site directed mutagenesis to substitute these amino acids with others having different properties. Steady state kinetic parameters and pH titrations of the variant enzymes showed AKsc-K18 and H292 to be important for binding and catalysis. Little was known about how the S. cerevisiae aspartate pathway kinases, AKsc and homoserine kinase (HSKsc), catalyze the transfer of the y-phosphate from adenosine triphosphate (ATP) to L-aspartate or L-homoserine, respectively. -

The Biosynthesis Reaction of Hypotaurine to Taurine
UNIVERSITY OF CENTRAL OKLAHOMA Edmond, Oklahoma Jackson College of Graduate Studies The Biosynthesis Reaction of Hypotaurine to Taurine A THESIS SUBMITTED TO THE GRADUATE FACULTY In partial fulfillment of the requirements For the degree of MASTER OF SCIENCE IN BIOLOGY By Roxanna Q. Grove Edmond, Oklahoma 2018 Acknowledgments Working on this project has been a period of intense learning for me, not only in the scientific arena, but also on a personal level. Writing this thesis has had a significant impact on me. I would like to reflect on people who have been supported and helped me so much throughout this period. First of all, I would like to express my gratitude toward my advisor, Dr. Steven J. Karpowicz, for his devotion, inspiration, and guidance. I am so grateful to have the opportunity to work with such an intelligent, dedicated, and patient professor. I appreciate his vast knowledge and skills in many areas such as biochemistry, genetics, and bioinformatics, and his assistance in writing this thesis. I would like to thank the other members of my committee, Dr. Nikki Seagraves, Dr. Hari Kotturi, and Dr. Lilian Chooback, for their guidance, support, and for providing materials throughout this project. An exceptional thanks go to Dr. John Bowen of the Department of Chemistry for advice in the analytical laboratory and Dr. Susan L. Nimmo from the Department of Chemistry and Biochemistry at the University of Oklahoma for assistance with NMR. This project was supported by funding from the College of Mathematics and Science and a Research, Creative, and Scholarly Activities (RCSA) grant from the Office of High Impact Practices at UCO. -

2018 Annual Meeting Proceedings
Downloaded from orbit.dtu.dk on: Oct 04, 2021 Genomics driven discovery and engineering of fungal polycyclic polyketides Subko, Karolina; Wolff, Peter B.; Theobald, Sebastian; Frisvad, Jens C.; Gotfredsen, Charlotte H.; Andersen, Mikael R.; Mortensen, Uffe H.; Larsen, Thomas O. Publication date: 2018 Document Version Publisher's PDF, also known as Version of record Link back to DTU Orbit Citation (APA): Subko, K., Wolff, P. B., Theobald, S., Frisvad, J. C., Gotfredsen, C. H., Andersen, M. R., Mortensen, U. H., & Larsen, T. O. (2018). Genomics driven discovery and engineering of fungal polycyclic polyketides. 68. Abstract from The American Society of Pharmacognosy Annual Meeting 2018, Lexington, Kentucky, United States. General rights Copyright and moral rights for the publications made accessible in the public portal are retained by the authors and/or other copyright owners and it is a condition of accessing publications that users recognise and abide by the legal requirements associated with these rights. Users may download and print one copy of any publication from the public portal for the purpose of private study or research. You may not further distribute the material or use it for any profit-making activity or commercial gain You may freely distribute the URL identifying the publication in the public portal If you believe that this document breaches copyright please contact us providing details, and we will remove access to the work immediately and investigate your claim. Abstracts | 2018 Annual Meeting of the ASP | July 21-25, 2018 | Lexington, Kentucky, USA 2 The American Society of Pharmacognosy Annual Meeting July 21-25, 2018 Hilton Lexington Downtown Lexington, Kentucky ENGAGE IN THE CONVO #ASP2018 Welcome to 59th Annual Meeting of the American Society of Pharmacognosy (ASP) 2018 in beautiful downtown Lexington, Kentucky! The Local Organizing Committee would like to thank you for joining us for the ASP meeting at the Hilton Lexington Downtown. -

Causal Agent, Biology and Management of the Leaf and Stem
CAUSAL AGENT, BIOLOGY AND MANAGEMENT OF THE LEAF AND STEM DISEASE OF BOXWOOD {BUXUS SPP.) A Thesis Presented to The Faculty of Graduate Studies of The University of Guelph by FANG SHI In partial fulfillment of requirements for the degree of Master of Science May, 2011 ©Fang Shi, 2011 Library and Archives Bibliotheque et 1*1 Canada Archives Canada Published Heritage Direction du Branch Patrimoine de I'edition 395 Wellington Street 395, rue Wellington OttawaONK1A0N4 Ottawa ON K1A 0N4 Canada Canada Your file Votre reference ISBN: 978-0-494-82801-4 Our file Notre reference ISBN: 978-0-494-82801-4 NOTICE: AVIS: The author has granted a non L'auteur a accorde une licence non exclusive exclusive license allowing Library and permettant a la Bibliotheque et Archives Archives Canada to reproduce, Canada de reproduire, publier, archiver, publish, archive, preserve, conserve, sauvegarder, conserver, transmettre au public communicate to the public by par telecommunication ou par I'lnternet, preter, telecommunication or on the Internet, distribuer et vendre des theses partout dans le loan, distribute and sell theses monde, a des fins commerciales ou autres, sur worldwide, for commercial or non support microforme, papier, electronique et/ou commercial purposes, in microform, autres formats. paper, electronic and/or any other formats. The author retains copyright L'auteur conserve la propriete du droit d'auteur ownership and moral rights in this et des droits moraux qui protege cette these. Ni thesis. Neither the thesis nor la these ni des extraits substantiels de celle-ci substantial extracts from it may be ne doivent etre imprimes ou autrement printed or otherwise reproduced reproduits sans son autorisation. -

Inhibition and Cofactor Targeting of Hypoxia-Sensing Proteins
University of Massachusetts Amherst ScholarWorks@UMass Amherst Doctoral Dissertations Dissertations and Theses August 2015 Inhibition and Cofactor Targeting of Hypoxia-Sensing Proteins Cornelius Y. Taabazuing University of Massachusetts Amherst Follow this and additional works at: https://scholarworks.umass.edu/dissertations_2 Part of the Biochemistry Commons, Inorganic Chemistry Commons, and the Structural Biology Commons Recommended Citation Taabazuing, Cornelius Y., "Inhibition and Cofactor Targeting of Hypoxia-Sensing Proteins" (2015). Doctoral Dissertations. 409. https://doi.org/10.7275/6956835.0 https://scholarworks.umass.edu/dissertations_2/409 This Open Access Dissertation is brought to you for free and open access by the Dissertations and Theses at ScholarWorks@UMass Amherst. It has been accepted for inclusion in Doctoral Dissertations by an authorized administrator of ScholarWorks@UMass Amherst. For more information, please contact [email protected]. INHIBITION AND COFACTOR TARGETING OF HYPOXIA-SENSING PROTEINS A Dissertation Presented by CORNELIUS TAABAZUING Submitted to the Graduate School of the University of Massachusetts Amherst in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY May 2015 Department of Chemistry © Copyright by Cornelius Taabazuing 2015 All Rights Reserved INHIBITION AND COFACTOR TARGETING OF HYPOXIA-SENSING PROTEINS A Dissertation Presented by CORNELIUS TAABAZUING Approved as to style and content by: ______________________________ Michael J. Knapp, Chair ______________________________ Michael J. Maroney, Member ______________________________ Nathan A. Schnarr, Member ______________________________ Scott C. Garman, Outside Member ______________________________ Craig T. Martin, Department Head Department of Chemistry DEDICATION To my mom, Barbara Soonyime, thank you for the sacrifices you have made to make it possible to pursue my dreams. To my wife Rachelle Taabazuing, thank you for providing me with your love, motivation, and patience.