Identification of Rfk-1, a Meiotic Driver Undergoing RNA Editing in Neurospora
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
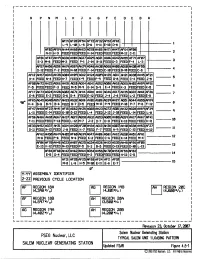
Salem Generating Station, Units 1 & 2, Revision 29 to Updated Final Safety Analysis Report, Chapter 4, Figures 4.5-1 to 4.5
r------------------------------------------- 1 I p M J B I R N L K H G F E D c A I I I I I Af'Jq AF20 AF54 AF72 32 AF52 AF18 I L-q L-10 L-15 D-6 -11 E-10 D-8 l I AF03 Af't;qAH44 AH60 AH63 AG70 AH65 AH7l AH47 AFS4 AF08 I N-ll H-3 FEED FEED FEED H-14 FEED FEED FEED M-12 C-11 2 I AF67 AH4q AH04 AG27 AG2<i' AG21 AG16 AG42 AF71 AF07 AF01 AG36 AH!5!5 3 I E-3 M-6 FEED M-3 FEED P-1 J-14 B-11 FEED D-3 FEED F-4 L-3 I AF67 AH5S AG56 Atflq AGsq AH2<1' AG48 AH30 AG68 AH08 AG60 AH30 AF55 I D-12 FEED F-2 FEED N-11 FEED F-14 FEED C-11 FEED B-11 FEED C-8 4 I AF12 AH57 AG43 AH38 AHtiJq AG12 AH24 AGfR AH25 AGil AG31 AH45 AF21 AGlM AH21 5 I H~4 FEED N-4 FEED H-7 FEED K~q FEED F-q FEED G-8 FEED C-4 FEED J-15 I AF50 AH72 AH22 AGS6 AH15 AGll.lAG64 AG41 AG52 AG88 AH18 AG65 AHIJ2 AH5q AF51 I F-5 FEED FEED F-3 FEED M-5 r+q G-14 o-q E-4 FEED K-3 FEED FEED K-5 6 I f:Fl7 AH73 AG24 AH28 AG82 AG71 AH14 AG18 AHil AG46 AG17 AH35 AG22 AH61 AF26 7 I E-8 FEED E-2 FEED G-6 G-4 FEED E-12 FEED J-4 J-6 FEED L-2 FEED E-5 I Af&q I qeo AF65 AG45 AtM0 AG57 AH33 AG32 AG16 AH01 AGI6 AG3<1' AH27 AG51 AG44 AG55 K-4 B-8 e-q B-6 FEED B-7 P-5 FEEC M-11 P-q FEED P-11 P-7 P-8 F-12 8 I AF47 AH68 AF23 AH41 AF1!5 AG62 AH26 AG03 AH23 AH32 AG28 AHsq AF3<1' q I L-U FEED E-14 FEED G-10 G-12 FEED L-4 FEED FEED L-14 FEED L-8 I ~~ AF66 AH66 AH10 AG67 AH37 AGJq AG68 AG3l AG63 AG05 AH08 AG5q AH17 AH67 AF41 I F-11 FEED FEED F-13 FEED L-12 M-7 J-2 D-7 D-11 FEED K-13 FEED FEED K-11 10 I AE33 AH!52 AG37 AH31 AG14 AH20 AF20 AH34 AG13 AH36 AG07 AH40 AG38 AH!53 AF27 I G-ll FEED N-12 FEED J-8 FEED K-7 FEED -

1St IRF Asia Regional Congress & Exhibition
1st IRF Asia Regional Congress & Exhibition Bali, Indonesia November 17–19 , 2014 For Professionals. By Professionals. "Building the Trans-Asia Highway" Bali’s Mandara toll road Executive Summary International Road Federation Better Roads. Better World. 1 International Road Federation | Washington, D.C. ogether with the Ministry of Public Works Indonesia, we chose the theme “Building the Trans-Asia Highway” to bring new emphasis to a visionary project Tthat traces its roots back to 1959. This Congress brought the region’s stakeholders together to identify new and innovative resources to bridge the current financing gap, while also sharing case studies, best practices and new technologies that can all contribute to making the Trans-Asia Highway a reality. This Congress was a direct result of the IRF’s strategic vision to become the world’s leading industry knowledge platform to help countries everywhere progress towards safer, cleaner, more resilient and better connected transportation systems. The Congress was also a reflection of Indonesia’s rising global stature. Already the largest economy in Southeast Asia, Indonesia aims to be one of world’s leading economies, an achievement that will require the continued development of not just its own transportation network, but also that of its neighbors. Thank you for joining us in Bali for this landmark regional event. H.E. Eng. Abdullah A. Al-Mogbel IRF Chairman Minister of Transport, Kingdom of Saudi Arabia Indonesia Hosts the Region’s Premier Transportation Meeting Indonesia was the proud host to the 1st IRF Asia Regional Congress & Exhibition, a regional gathering of more than 700 transportation professionals from 52 countries — including Ministers, senior national and local government officials, academics, civil society organizations and industry leaders. -

Characterization and Transferability of Microsatellite Markers of the Cultivated Peanut (Arachis Hypogaea)
BMC Plant Biology BioMed Central Research article Open Access Characterization and transferability of microsatellite markers of the cultivated peanut (Arachis hypogaea) Marcos A Gimenes*1,2, Andrea A Hoshino1, Andrea VG Barbosa1, Dario A Palmieri1 and Catalina R Lopes1 Address: 1Laboratório de Biotecnologia e Genética Molecular (BIOGEM), Departamento de Genética, Instituto de Biociências, Universidade Estadual Paulista (UNESP), Botucatu, SP, Brazil and 2Instituto Agronômico de Campinas – RGV, Caixa Postal 28, Campinas, SP, Brazil Email: Marcos A Gimenes* - [email protected]; Andrea A Hoshino - [email protected]; Andrea VG Barbosa - [email protected]; Dario A Palmieri - [email protected]; Catalina R Lopes - [email protected] * Corresponding author Published: 27 February 2007 Received: 3 March 2006 Accepted: 27 February 2007 BMC Plant Biology 2007, 7:9 doi:10.1186/1471-2229-7-9 This article is available from: http://www.biomedcentral.com/1471-2229/7/9 © 2007 Gimenes et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: The genus Arachis includes Arachis hypogaea (cultivated peanut) and wild species that are used in peanut breeding or as forage. Molecular markers have been employed in several studies of this genus, but microsatellite markers have only been used in few investigations. Microsatellites are very informative and are useful to assess genetic variability, analyze mating systems and in genetic mapping. The objectives of this study were to develop A. -

Asian Highway Handbook United Nations
ECONOMIC AND SOCIAL COMMISSION FOR ASIA AND THE PACIFIC ASIAN HIGHWAY HANDBOOK UNITED NATIONS New York, 2003 ST/ESCAP/2303 The Asian Highway Handbook was prepared under the direction of the Transport and Tourism Division of the United Nations Economic and Social Commission for Asia and the Pacific. The team of staff members of the Transport and Tourism Division who prepared the Handbook comprised: Fuyo Jenny Yamamoto, Tetsuo Miyairi, Madan B. Regmi, John R. Moon and Barry Cable. Inputs for the tourism- related parts were provided by an external consultant: Imtiaz Muqbil. The designations employed and the presentation of the material in this publication do not imply the expression of any opinion whatsoever on the part of the Secretariat of the United Nations concerning the legal status of any country, territory, city or area or of its authorities, or concerning the delimitation of its frontiers or boundaries. This publication has been issued without formal editing. CONTENTS I. INTRODUCTION TO THE ASIAN HIGHWAY………………. 1 1. Concept of the Asian Highway Network……………………………… 1 2. Identifying the Network………………………………………………. 2 3. Current status of the Asian Highway………………………………….. 3 4. Formalization of the Asian Highway Network……………………….. 7 5. Promotion of the Asian Highway……………………………………... 9 6. A Vision of the Future………………………………………………… 10 II. ASIAN HIGHWAY ROUTES IN MEMBER COUNTRIES…... 16 1. Afghanistan……………………………………………………………. 16 2. Armenia……………………………………………………………….. 19 3. Azerbaijan……………………………………………………………... 21 4. Bangladesh……………………………………………………………. 23 5. Bhutan…………………………………………………………………. 27 6. Cambodia……………………………………………………………… 29 7. China…………………………………………………………………... 32 8. Democratic People’s Republic of Korea……………………………… 36 9. Georgia………………………………………………………………... 38 10. India…………………………………………………………………… 41 11. Indonesia………………………………………………………………. 45 12. Islamic Republic of Iran………………………………………………. 49 13 Japan………………………………………………………………….. -

Identification of Rfk-1, a Meiotic Driver Undergoing RNA Editing in Neurospora
HIGHLIGHTED ARTICLE | INVESTIGATION Identification of rfk-1, a Meiotic Driver Undergoing RNA Editing in Neurospora Nicholas A. Rhoades,* Austin M. Harvey,*,1 Dilini A. Samarajeewa,*,1 Jesper Svedberg,† Aykhan Yusifov,* Anna Abusharekh,* Pennapa Manitchotpisit,* Daren W. Brown,‡ Kevin J. Sharp,* David G. Rehard,§,** Joshua Peters,* Xavier Ostolaza-Maldonado,* Jackson Stephenson,* Patrick K. T. Shiu,§ Hanna Johannesson,† and Thomas M. Hammond*,2 *School of Biological Sciences, Illinois State University, Normal, Illinois 61790, †Department of Organismal Biology, Uppsala University, Norbyvägen 18D, 752 36 Uppsala, Sweden, ‡Mycotoxin Prevention and Applied Microbiology, National Center for Agricultural Utilization Research, U.S. Department of Agriculture, Agricultural Research Service, Peoria, Illinois 61604, §Division of Biological Sciences, University of Missouri, Columbia, Missouri 65211, and **Department of Biology, University of Iowa, Iowa 52242 ABSTRACT Sk-2 is a meiotic drive element that was discovered in wild populations of Neurospora fungi over 40 years ago. While early studies quickly determined that Sk-2 transmits itself through sexual reproduction in a biased manner via spore killing, the genetic factors responsible for this phenomenon have remained mostly unknown. Here, we identify and characterize rfk-1, a gene required for Sk-2-based spore killing. The rfk-1 gene contains four exons, three introns, and two stop codons, the first of which undergoes RNA editing to a tryptophan codon during sexual development. Translation of an unedited rfk-1 transcript in vegetative tissue is expected to produce a 102-amino acid protein, whereas translation of an edited rfk-1 transcript in sexual tissue is expected to produce a protein with 130 amino acids. These findings indicate that unedited and edited rfk-1 transcripts exist and that these transcripts could have different roles with respect to the mechanism of meiotic drive by spore killing. -

The Effect of Road Upgrading to Overland Trade in Asian Highway Network Ziyodullo PARPIEV ∗ Jamshid SODIKOV **
Eurasian Journal of Business and Economics 2008, 1 (2), 85-101. The Effect of Road Upgrading to Overland Trade in Asian Highway Network Ziyodullo PARPIEV ∗ Jamshid SODIKOV ** Abstract This paper investigates an impact of road upgrading and improvement on overland trade in 18 out of 32 Asian Highway Network member countries. A regression based cost model was developed. The results indicate that approximately 6.5 billion US dollars is required to upgrade and improve surface condition of the selected roads with total length of 15,842 km. The gravity model approach was adopted to quantitatively evaluate overland trade expansion assuming pessimistic and optimistic scenarios: improvements in road quality indices up to 50 and up to 75, respectively. The results suggests that in the first scenario total intra-regional trade will increase by about 20 percent or 48.7 billion US dollars annually, while second scenario predicts that trade will increase by about 35 percent or 89.5 billion US dollars annually. Keywords: Asian Highway Network, road transport, gravity model. Jel Classification: F12, F15, F17. ∗ Advisor-Economist, UNDP Uzbekistan Country Office, Email: [email protected] ** Chief Engineer, Road Research Institute, Tashkent, Uzbekistan The views expressed in this paper are those of the author(s) and do not necessarily represent those of organizations the authors are associated with. Ziyodullo PARPIEV & Jamshid SODIKOV 1. Introduction In 1992, the United Nations Economic and Social Commission for Asia and the Pacific (ESCAP) endorsed the Asian Land Transport Infrastructure Development (ALTID) project comprising of the Asian Highway and the Trans-Asian Railway network. The formalization of the Asian Highway, through the Intergovernmental Agreement on Asian Highway Network (AHN), was adopted in November 2003. -

Growing Together Articulates a Number of Proposals That Can Help the Region Exploit Its Huge Untapped Potential for Regional Economic Integration
i Photo by Warren Field ii FOREWORD For the global economy, these are difficult times. The world is emerging from a crisis whose aftershocks continue to resonate – trapping some of the richest economies in recession and shaking the foundations of one of the world’s major currencies. Here at ESCAP, there are historical echoes. What is now the Economic and Social Commission for Asia and the Pacific was founded more than 60 years ago – also in the aftermath of a global crisis. The countries of Asia and the Pacific established their new Commission partly to assist them in rebuilding their economies as they came out of the yoke of colonialism and the Second World War. The newly established ECAFE, as ESCAP was called then, held a ministerial conference on regional economic cooperation in 1963 that resolved to set up the Asian Development Bank with the aim of assisting the countries in the region in rebuilding their economies. Fifty years later, the Asia-Pacific region is again at a crossroads, on this occasion seeking ways and means to sustain its dynamism in a dramatically changed global context in the aftermath of a global financial and economic crisis. An important change is the fact that, burdened by huge debts and global imbalances, the advanced economies of the West are no longer able to play the role of engines of growth for the Asia-Pacific region that they played in the past. Hence, the Asia-Pacific region has to look for new engines of growth. The secretariat of ESCAP has argued over the past few years that regional developmental challenges, such as poverty and wide disparities in social and physical infrastructure, can be turned into opportunities for sustaining growth in the future. -
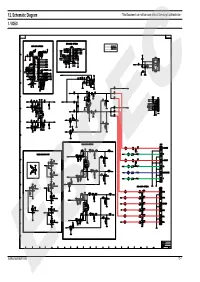
12. Schematic Diagram - This Document Can Not Be Used Without Samsung’S Authorization - 1
12. Schematic Diagram - This Document can not be used without Samsung’s authorization - 1. VIDEO Samsung Electronics ESPEC12-1 2. MPEG - This Document can not be used without Samsung’s authorization - 12-2 ESPECSamsung Electronics 3. FRONT - This Document can not be used without Samsung’s authorization - Samsung Electronics ESPEC12-3 4. SMPS - This Document can not be used without Samsung’s authorization - 12-4 ESPECSamsung Electronics 5. MIC, ECHO - This Document can not be used without Samsung’s authorization - Samsung Electronics ESPEC12-5 6. SUB, SCART - This Document can not be used without Samsung’s authorization - 12-6 ESPECSamsung Electronics 1.Exploded Views and Parts List 1-1 Total Exploded View Samsung Electronics ESPEC1-1 * Parts List of Total Exploded view No. Code No. Description Specification Q’ty Remark 1 AH64-03184B DECO-STAND BY PMMA 1 2 AH64-03185B WINDOW-VFD PMMA 1 3 AH63-00875A FILTER-VFD PC 0.5T 1 4 AH64-03178B CABINET-FRONT PC/ABS 1 5 AH64-03181B DECO-SIDE R B ABS 1 6 AH64-03180B DECO-SIDE L B ABS 1 7 AH67-00201A LENS-POWER PMMA 1 8 AH64-03189B KNOB-POWER ABS 1 9 AH64-03188B KNOB-OPEN ABS 1 10 AH64-03186B KNOB-FUNCTION ABS 1 11 AH63-00892A SHEET-VFD PC 0.5T 1 12 AH61-01766A BRKT-JACK SECC 0.6T 1 13 AH64-03187B KNOB-MIC ABS 2 14 AH64-03190B CABINET-BOTTOM-REAER SECC 0.6T 1 15 AH64-03179B DOOR-CD ABS 1 16 AH61-01765A HOLDER-VFD ABS 1 17 AH63-00856A SHEET-PCB PC 0.5T 1 18 AH63-00876A SHEET-TOP PC 0.5T 1 19 AH64-03183B DECO-SIDE R UP ABS 1 20 AH64-03191A CABINET-TOP SECC 0.5T 1 21 AH64-03182B DECO-SIDE L UP ABS 1 A 6002-000126 TAPTITE SCREW FH + 3*10 B 6003-001327 TAPTITE SCREW RH + 2.6*12 C 6003-001479 TAPTITE SCREW BH + 1.7*4 D 6003-001561 TAPTITE SCREW BH + 3*6 E 6003-000275 TAPTITE SCREW BH + 3*10(BLK) 1-2 ESPECSamsung Electronics 1-2 DVD MECHA Exploded View and Parts list No. -

Asian Highway Handbook
ECONOMIC AND SOCIAL COMMISSION FOR ASIA AND THE PACIFIC ASIAN HIGHWAY HANDBOOK UNITED NATIONS New York, 2003 ST/ESCAP/2303 The Asian Highway Handbook was prepared under the direction of the Transport and Tourism Division of the United Nations Economic and Social Commission for Asia and the Pacific. The team of staff members of the Transport and Tourism Division who prepared the Handbook comprised: Fuyo Jenny Yamamoto, Tetsuo Miyairi, Madan B. Regmi, John R. Moon and Barry Cable. Inputs for the tourism- related parts were provided by an external consultant: Imtiaz Muqbil. The designations employed and the presentation of the material in this publication do not imply the expression of any opinion whatsoever on the part of the Secretariat of the United Nations concerning the legal status of any country, territory, city or area or of its authorities, or concerning the delimitation of its frontiers or boundaries. This publication has been issued without formal editing. CONTENTS I. INTRODUCTION TO THE ASIAN HIGHWAY………………. 1 1. Concept of the Asian Highway Network……………………………… 1 2. Identifying the Network………………………………………………. 2 3. Current status of the Asian Highway………………………………….. 3 4. Formalization of the Asian Highway Network……………………….. 7 5. Promotion of the Asian Highway……………………………………... 9 6. A Vision of the Future………………………………………………… 10 II. ASIAN HIGHWAY ROUTES IN MEMBER COUNTRIES…... 16 1. Afghanistan……………………………………………………………. 16 2. Armenia……………………………………………………………….. 19 3. Azerbaijan……………………………………………………………... 21 4. Bangladesh……………………………………………………………. 23 5. Bhutan…………………………………………………………………. 27 6. Cambodia……………………………………………………………… 29 7. China…………………………………………………………………... 32 8. Democratic People’s Republic of Korea……………………………… 36 9. Georgia………………………………………………………………... 38 10. India…………………………………………………………………… 41 11. Indonesia………………………………………………………………. 45 12. Islamic Republic of Iran………………………………………………. 49 13 Japan………………………………………………………………….. -

ASIAN HIGHWAY ROUTE MAP 500 Km
40° E 60° E 80° E 100° E 120° E 140° E 500 Km. Vyborg Torpynovka 60° N St. Petersburg ASIAN HIGHWAY ROUTE MAP 500 Km. 60° N RUSSIAN FEDERATION AH8 Yekaterinburg AH7 AH6 Moscow AH6 AH6 Krasnoyarsk AH6 Ufa Petuhovo AH6 Chelyabinsk E30 Chistoe Isilkul AH6 Omsk Novosibirsk E30 AH60 Krasnoe Petropavlovsk Karakuga E30 AH4 AH8 Troisk AH6 Samara Cherlak E119 Kaerak AH7 AH62 Pnirtyshskoe E30 Kostanai AH64 Barnaul AH30 E123 Kokshetau AH60 AH30 E125 AH63 E127 Tambov E121 Pavlodar AH64 Irkutsk Ulan-Ude Borysoglebsk Saratov Kurlin AH7 AH64 Chita Kursk AH61 Pogodaevo AH60 AH6 Ozinki AH64 (AH64) AH6 E38 Shiderty (AH67) AH3 AH61 Voronezh Ural'sk AH62 Astana E127 Veseloyarskyj AH4 AH6 Krupets Kamenka AH7 (AH67) Belogorsk E38 AH8 Zhaisan E123 Krasny Aul E119 E125 Kyahta AH61 Arkalyk AH67 Semipalatinsk Heihe AH63 Altanbulag Zabaykalsk Blagoveshchensk E38 Aktobe Karabutak Karaganda AH60 E121 Tashanta (AH67) Ulaanbaishint Darkhan Georgievka AH3 Manzhouli AH30 Volgograd AH61 AH4 AH6 Donetsk AH70 E38 AH67 Khabarovsk AH32 E40 E40 AH60 (AH67) Ulaanbaatar Nalayh AH8 AH7 Hovd AH32 Sumber Arshan E119 Zhezkazgan AH32 Qiqihar AH31 Tongjiang (AH70) Atyrau AH63 E125 Taskesken Choybalsan (AH70) KAZAKHSTAN Uliastay Kotyaevka AH70 E105 Numrug Port of Odessa E40 Aralsk Baketu Ondorhaan AH62 Bakhty AH35 AH33 Astrakhan E121 E123 AH67 Takeshkan Choir Ucharal AH3 AH32 E40 AH68 MONGOLIA Yarantai Harbin AH30 Beyneu E014 Dostyk Bichigt E119 Kyzylorda AH60 AH8 Burubaital Alatawshankou AH4 E40 AH35 Daut-ota Saynshand AH6 AH60 AH5 AH31 Pogranichny AH70 AH5 Port of Constantza 500 -

Pub 2424 Fulltext 0.Pdf
ESCAP is the regional development arm of the United Nations and serves as the main economic and social development centre for the United Nations in Asia and the Pacific. Its mandate is to foster cooperation between its 53 members and 9 associate members. ESCAP provides the strategic link between global and country-level programmes and issues. It supports Governments of the region in consolidating regional positions and advocates regional approaches to meeting the region’s unique socio-economic challenges in a globalizing world. The ESCAP office is located in Bangkok, Thailand. Please visit our website at www.unescap.org for further information. The shaded areas of the map indicate ESCAP members and associate members. ECONOMIC AND SOCIAL COMMISSION FOR ASIA AND THE PACIFIC Priority Investment Needs for the Development of the Asian Highway Network United Nations New York, 2006 ST/ESCAP/2424 This publication was prepared under the direction of the Transport and Tourism Division of the Economic and Social Commission for Asia and the Pacific. Inputs related to priority investment needs and projects were provided by national experts and representatives of member countries at three subregional expert group meetings. The designations employed and the presentation of the material in this publication do not imply the expression of any opinion whatsoever on the part of the Secretariat of the United Nations concerning the legal status of any country, territory, city or area, or of its authorities, or the delineation of its frontiers or boundaries. This publication has been issued without formal editing. ii CONTENTS Page INTRODUCTION ................................................................................................................................... 1 I. STATUS OF THE ASIAN HIGHWAY NETWORK ................................................................. -

Natural Capital Approaches for Sustainable Development
Natural Capital Approaches for Sustainable Development Emily McKenzie Chief Adviser, Economics and Sustainability © Anton Vorauer / WWF Outline 1. Natural capital and the SDGs 2. The case of Myanmar 3. What tools are available? 4. Natural capital in business decisions 2 27-Jun-17 1 Natural Capital and the SDGs © BRANDON COLE/WWW.NATUREPL.COM 2 5 27-Jun-17 © Azote Images for Stockholm Resilience Centre 6 What is natural capital? Natural Capital is the stock of renewable and non-renewable natural resources, (e.g. plants, animals, air, water, soils, minerals) that combine to yield a flow of benefits to people Food, fuel, fiber Climate Pollination regulation Coastal Clean protection water Spiritual Fulfilment 7 27-Jun-17 8 Multiple forms of capital Financial capital Manufactured capital Intellectual Human capital capital Social and relationship capital Natural capital <IR> capitals framework 9 10 11 12 13 Millennium Ecosystem Assessment • 60% of ecosystem services are being degraded or used unsustainably • Degradation of ecosystem services causes significant harm to human well-being 14 27-Jun-17 © Christy Williams / WWF Sustainable Securing Sustainable, Standards for Safe, Resilient Development Freshwater Livable Cities the Private Coastal Planning Sector Communities Working together to account for nature’s values, toward shared outcomes THEORY OF CHANGE Robust evidence of Build and tell Create user-friendly conditions for success stories, approaches & tools success engage leaders Get information about natural capital into decisions Make decisions