IP6K Structure and the Molecular Determinants of Catalytic Specificity
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Weiterführende Literatur Zum Lehrbuch „Biochemie“ (Kapitel 4 – 50)
Weiterführende Literatur zum Lehrbuch „Biochemie“ (Kapitel 4 – 50) Im Buch ist aus Platzgründen auf die Angabe weiterführender Literatur verzichtet worden. Auskünfte zu den thematischen Schwerpunkten der vier Hauptteile II bis V geben die im Folgenden aufgeführten knapp 1000 einschlägigen Artikel aus wissenschaftlichen Fachzeitschriften. Die Zusammenstellung folgt der Gliederung des Buchs. Jeder Link führt direkt zur Literaturliste des entsprechenden Kapitels, aber es ist stets auch die gesamte Literatur hinterlegt (z.B. für einen kompletten Ausdruck). Die Aufstellung enthält überwiegend Review-Artikel der letzten 4 Jahre, in denen der aktuelle Kenntnisstand zu einem ausgewählten Thema von führenden Wissenschaftlern präsentiert wird. Beim Einführungsteil (Kapitel 1 bis 3) haben wir dagegen bewusst auf Spezialliteratur verzichtet; hier sei auf einschlägige Lehrbücher der Biochemie, Molekularbiologie, Genetik und Zellbiologie, die diesen fundamentalen Aspekten breiten Raum widmen, verwiesen. Falls Sie als Leser interessante Artikel finden, die unsere Literatursammlung ergänzen und bereichern könnten, bitten wir um kurze Mitteilung an den Verlag. Werner Müller-Esterl Ulrich Brandt Oliver Anderka Stefan Kerscher 1 Teil II Struktur und Funktion von Proteinen 4 Proteine – Werkzeuge der Zelle 4.1 Liganden binden an Proteine und verändern deren Konformation Wilson MA, Brunger AT (2000) The 1.0 Å crystal structure of Ca2+-bound calmodulin: an analysis of disorder and implications for functionally relevant plasticity, J Mol Biol 301, 1237-1256 O'Day DH & Myre MA (2004) Calmodulin-binding domains in Alzheimer's disease proteins: extending the calcium hypothesis. Biochemical and Biophysical Research Communications, 320, 1051-1054. 4.2 Enzyme binden Substrate und setzen sie zu Produkten um Showalter AK & Tsai MD (2002) A reexamination of the nucleotide incorporation fidelity of DNA polymerases. -

Part I Principles of Enzyme Catalysis
j1 Part I Principles of Enzyme Catalysis Enzyme Catalysis in Organic Synthesis, Third Edition. Edited by Karlheinz Drauz, Harald Groger,€ and Oliver May. Ó 2012 Wiley-VCH Verlag GmbH & Co. KGaA. Published 2012 by Wiley-VCH Verlag GmbH & Co. KGaA. j3 1 Introduction – Principles and Historical Landmarks of Enzyme Catalysis in Organic Synthesis Harald Gr€oger and Yasuhisa Asano 1.1 General Remarks Enzyme catalysis in organic synthesis – behind this term stands a technology that today is widely recognized as a first choice opportunity in the preparation of a wide range of chemical compounds. Notably, this is true not only for academic syntheses but also for industrial-scale applications [1]. For numerous molecules the synthetic routes based on enzyme catalysis have turned out to be competitive (and often superior!) compared with classic chemicalaswellaschemocatalyticsynthetic approaches. Thus, enzymatic catalysis is increasingly recognized by organic chemists in both academia and industry as an attractive synthetic tool besides the traditional organic disciplines such as classic synthesis, metal catalysis, and organocatalysis [2]. By means of enzymes a broad range of transformations relevant in organic chemistry can be catalyzed, including, for example, redox reactions, carbon–carbon bond forming reactions, and hydrolytic reactions. Nonetheless, for a long time enzyme catalysis was not realized as a first choice option in organic synthesis. Organic chemists did not use enzymes as catalysts for their envisioned syntheses because of observed (or assumed) disadvantages such as narrow substrate range, limited stability of enzymes under organic reaction conditions, low efficiency when using wild-type strains, and diluted substrate and product solutions, thus leading to non-satisfactory volumetric productivities. -
Snapshot: Inositol Phosphates Ace J
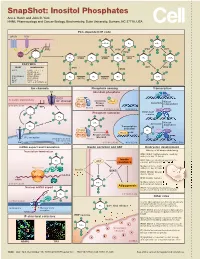
SnapShot: Inositol Phosphates Ace J. Hatch and John D. York HHMI, Pharmacology and Cancer Biology, Biochemistry, Duke University, Durham, NC 27710, USA PLC-dependent IP code GPCR RTK O O O O O O 5-PP-IP4 IP4 5-IP7 O O O O O O PIP2 O IP6K O IP6K O VIP1 O O O 2 O ITPK1 O 13 O PLC 2 O O O O O O O O 4 6 13 O 5 IP3 IPMK IP4 IPMK IP5 IPK1 IP6 1,5-IP8 4 6 O O 5 O O O O O O O O ENZYMES O O O O O O YEAST MAMMALIAN IP3K VIP1 IP6K IPMK PLC1 PLCβ, γ, δ, ε, ζ, η O - IP3KA, B, C - ITPK1 (IP56K) O O O O O O O O IPK2(ARG82) IPMK (IPK2) IP4 IP3 IP4 1-IP7 IPK1 IPK1 (IP5K) INPP5 ITPK1 KCS1 IP6K1, 2, 3 O O O O O O VIP1 VIP1, 2 (PPIP5K1, 2) O O Ion channels Phosphate sensing Transcription Cl- Abundant phosphate MCM1 ARG80 CIC3 P PLASMA MEMBRANE - Pho80 Cl channel Pho4 Kinase Kinase Assembly Pho85 independent CYTOPLASM activity 2 O PIP2 Pho81 13 CYTOPLASM NUCLEUS IPK2 ARG81 4 6 Phosphate starvation MCM1-ArgR O 5 O complex O O IP4 O O O O O O O 1-IP7 Kinase Activation dependent IP3 O O Transcription O O O activated Pho80 IP4 O X Pho4 O O Pho85 Kinase activity IP receptor blocked O 3 ENDOPLASMIC Pho81 RETICULUM Ca2+ CYTOPLASM NUCLEUS NUCLEUS mRNA export and translation Insulin secretion and AKT Embryonic development Translation termination Effects of IP kinase deficiency O IPMK (IPK2): Multiple defects, death by embryonic day 10 (mice) O O Insulin IPK1: Cillia are shortened and immotile IP6 AKT resistance causing patterning defects (zebrash) O O Multiple defects, death by Ribosome O embryonic day 8.5 (mice) GleI eRF1 Insulin GSK3β Dbp5 ITPK1 (IP56K): Neural tube -

Enzyme DHRS7
Toward the identification of a function of the “orphan” enzyme DHRS7 Inauguraldissertation zur Erlangung der Würde eines Doktors der Philosophie vorgelegt der Philosophisch-Naturwissenschaftlichen Fakultät der Universität Basel von Selene Araya, aus Lugano, Tessin Basel, 2018 Originaldokument gespeichert auf dem Dokumentenserver der Universität Basel edoc.unibas.ch Genehmigt von der Philosophisch-Naturwissenschaftlichen Fakultät auf Antrag von Prof. Dr. Alex Odermatt (Fakultätsverantwortlicher) und Prof. Dr. Michael Arand (Korreferent) Basel, den 26.6.2018 ________________________ Dekan Prof. Dr. Martin Spiess I. List of Abbreviations 3α/βAdiol 3α/β-Androstanediol (5α-Androstane-3α/β,17β-diol) 3α/βHSD 3α/β-hydroxysteroid dehydrogenase 17β-HSD 17β-Hydroxysteroid Dehydrogenase 17αOHProg 17α-Hydroxyprogesterone 20α/βOHProg 20α/β-Hydroxyprogesterone 17α,20α/βdiOHProg 20α/βdihydroxyprogesterone ADT Androgen deprivation therapy ANOVA Analysis of variance AR Androgen Receptor AKR Aldo-Keto Reductase ATCC American Type Culture Collection CAM Cell Adhesion Molecule CYP Cytochrome P450 CBR1 Carbonyl reductase 1 CRPC Castration resistant prostate cancer Ct-value Cycle threshold-value DHRS7 (B/C) Dehydrogenase/Reductase Short Chain Dehydrogenase Family Member 7 (B/C) DHEA Dehydroepiandrosterone DHP Dehydroprogesterone DHT 5α-Dihydrotestosterone DMEM Dulbecco's Modified Eagle's Medium DMSO Dimethyl Sulfoxide DTT Dithiothreitol E1 Estrone E2 Estradiol ECM Extracellular Membrane EDTA Ethylenediaminetetraacetic acid EMT Epithelial-mesenchymal transition ER Endoplasmic Reticulum ERα/β Estrogen Receptor α/β FBS Fetal Bovine Serum 3 FDR False discovery rate FGF Fibroblast growth factor HEPES 4-(2-Hydroxyethyl)-1-Piperazineethanesulfonic Acid HMDB Human Metabolome Database HPLC High Performance Liquid Chromatography HSD Hydroxysteroid Dehydrogenase IC50 Half-Maximal Inhibitory Concentration LNCaP Lymph node carcinoma of the prostate mRNA Messenger Ribonucleic Acid n.d. -

Enzyme Promiscuity Prediction Using Hierarchy-Informed Multi-Label Classification Gian Marco Visani1, Michael C
Bioinformatics doi: 10.1093/bioinformatics/xxxxx Advance Access Publication Date: 22 Jan 2021 Manuscript Category Systems Biology Enzyme promiscuity prediction using hierarchy-informed multi-label classification Gian Marco Visani1, Michael C. Hughes1 and Soha Hassoun1,2,* 1Department of Computer Science, Tufts University, 161 College Ave, Medford, MA, 02155, USA, 2Department of Chemical and Biological Engineering, Tufts University, 4 Colby St, Medford, MA, 02155, USA *To whom correspondence should be addressed. Associate Editor: XXXXXXX Received on XXXXX; revised on XXXXX; accepted on XXXXX Abstract Motivation: As experimental efforts are costly and time consuming, computational characterization of enzyme capabilities is an attractive alternative. We present and evaluate several machine-learning models to predict which of 983 distinct enzymes, as defined via the Enzyme Commission (EC) numbers, are likely to interact with a given query molecule. Our data consists of enzyme-substrate interactions from the BRENDA database. Some interactions are attributed to natural selection and involve the enzyme’s natural substrates. The majority of the interactions however involve non-natural substrates, thus reflecting promiscuous enzymatic activities. Results: We frame this “enzyme promiscuity prediction” problem as a multi-label classification task. We maximally utilize inhibitor and unlabelled data to train prediction models that can take advantage of known hierarchical relationships between enzyme classes. We report that a hierarchical multi-label neural network, EPP-HMCNF, is the best model for solving this problem, outperforming k-nearest neighbours similarity-based and other machine learning models. We show that inhibitor information during training consistently improves predictive power, particularly for EPP-HMCNF. We also show that all promiscuity prediction models perform worse under a realistic data split when compared to a random data split, and when evaluating performance on non-natural substrates compared to natural substrates. -

Biosynthesis of New Alpha-Bisabolol Derivatives Through a Synthetic Biology Approach Arthur Sarrade-Loucheur
Biosynthesis of new alpha-bisabolol derivatives through a synthetic biology approach Arthur Sarrade-Loucheur To cite this version: Arthur Sarrade-Loucheur. Biosynthesis of new alpha-bisabolol derivatives through a synthetic biology approach. Biochemistry, Molecular Biology. INSA de Toulouse, 2020. English. NNT : 2020ISAT0003. tel-02976811 HAL Id: tel-02976811 https://tel.archives-ouvertes.fr/tel-02976811 Submitted on 23 Oct 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. THÈSE En vue de l’obtention du DOCTORAT DE L’UNIVERSITÉ DE TOULOUSE Délivré par l'Institut National des Sciences Appliquées de Toulouse Présentée et soutenue par Arthur SARRADE-LOUCHEUR Le 30 juin 2020 Biosynthèse de nouveaux dérivés de l'α-bisabolol par une approche de biologie synthèse Ecole doctorale : SEVAB - Sciences Ecologiques, Vétérinaires, Agronomiques et Bioingenieries Spécialité : Ingénieries microbienne et enzymatique Unité de recherche : TBI - Toulouse Biotechnology Institute, Bio & Chemical Engineering Thèse dirigée par Gilles TRUAN et Magali REMAUD-SIMEON Jury -

Ceramide Kinase Is Upregulated in Metastatic Breast Cancer Cells and Contributes to Migration and Invasion by Activation of PI 3-Kinase and Akt
International Journal of Molecular Sciences Article Ceramide Kinase Is Upregulated in Metastatic Breast Cancer Cells and Contributes to Migration and Invasion by Activation of PI 3-Kinase and Akt 1,2, 1, 2 2 Stephanie Schwalm y, Martin Erhardt y, Isolde Römer , Josef Pfeilschifter , Uwe Zangemeister-Wittke 1,* and Andrea Huwiler 1,* 1 Institute of Pharmacology, University of Bern, Inselspital, INO-F, CH-3010 Bern, Switzerland; [email protected] (S.S.); [email protected] (M.E.) 2 Institute of General Pharmacology and Toxicology, University Hospital Frankfurt am Main, Goethe-University, Theodor-Stern Kai 7, D-60590 Frankfurt am Main, Germany; [email protected] (I.R.); [email protected] (J.P.) * Correspondence: [email protected] (U.Z.-W.); [email protected] (A.H.); Tel.: +41-31-6323214 (A.H.) These authors contributed equally to this work. y Received: 29 November 2019; Accepted: 12 February 2020; Published: 19 February 2020 Abstract: Ceramide kinase (CerK) is a lipid kinase that converts the proapoptotic ceramide to ceramide 1-phosphate, which has been proposed to have pro-malignant properties and regulate cell responses such as proliferation, migration, and inflammation. We used the parental human breast cancer cell line MDA-MB-231 and two single cell progenies derived from lung and bone metastasis upon injection of the parental cells into immuno-deficient mice. The lung and the bone metastatic cell lines showed a marked upregulation of CerK mRNA and activity when compared to the parental cell line. The metastatic cells also had increased migratory and invasive activity, which was dose-dependently reduced by the selective CerK inhibitor NVP-231. -

Inhibition of Spinach Phosphoribulokinase by DL-Glyceraldehyde by ANTONI R
Biochem. J. (1976) 153, 613-619 613 Printed in Great Britain Inhibition of Spinach Phosphoribulokinase by DL-Glyceraldehyde By ANTONI R. SLABAS and DAVID A. WALKER Department ofBotany, University ofSheffield, Sheffield S1O 2TN, U.K. (Received 10 September 1975) Spinach chloroplast phosphoribulokinase is inhibited by DL-glyceraldehyde. The in- hibition is non-competitive with respect to ribulose 5-phosphate (Ki 19mM) and ATP (Ki 20mM). The inhibition is discussed in relation to a previously reported inhibition of CO2 assimilation in intact and envelope-free chloroplasts by DL-glyceraldehyde. It is concluded that the inhibition of phosphoribulokinase is insufficient to account for the inhibition, by DL-glyceraldehyde, of 02 evolution with ribose 5-phosphate as substrate and that a further site of inhibition is also present in this system. DL-Glyceraldehyde inhibits photosynthetic carbon glycylglycine buffer, pH7.4, in a total volume of assiniilation by intact chloroplasts and by the re- 13 ml. The reaction was terminated by the addition constituted chloroplast system (Stokes & Walker, of 2ml of 50 % (w/v) trichloroacetic acid and the pH 1972). The inhibition is most pronounced with intact was adjusted to pH8.0 with KOH. After the addition chloroplasts, a concentration of 1OmM being sufficient of Sml of 1.OM-barium acetate the solution was to suppress 02 evolution completely. It is important centrifuged and the precipitate discarded after because DL-glyceraldehyde is the only known inhibi- washing with Sml of water. Cold ethanol (4vol.) was tor of photosynthesis that is entirely without detect- added to the combined supernatants (total volume able effect on photophosphorylation or photo- 25.4ml); the new precipitate was recovered by synthetic electron transport. -

Enzyme Promiscuity of Carbohydrate Active Enzymes and Their Applications in Biocatalysis Pallister E, Gray CJ and Flitsch SL
Enzyme Promiscuity of Carbohydrate Active Enzymes and their applications in Biocatalysis Pallister E, Gray CJ and Flitsch SL School of Chemistry & Manchester Institute of Biotechnology, The University of Manchester, 131 Princess Street, Manchester M1 7DN, UK Corresponding Author: Sabine Flitsch ([email protected]) Abstract The application of biocatalysis for the synthesis of glycans and glycoconjugates is a well-established and successful strategy, both for small and large scale synthesis. Compared to chemical synthesis, is has the advantage of high selectivity, but biocatalysis had been largely limited to natural glycans both in terms of reactivity and substrates. This review describes recent advances in exploiting enzyme promiscuity to expand the range of substrates and reactions that carbohydrate active enzymes (CAZymes) can catalyse. The main focus is on formation and hydrolysis of glycosidic linkages, including sugar kinases, reactions that are central to glycobiotechnology. In addition, biocatalysts that generate sugar analogues and modify carbohydrates, such as oxidases, transaminases and acylases are reviewed. As carbohydrate active enzymes become more accessible and protein engineering strategies become faster, the application of biocatalysis in the generation of a wide range of glycoconjugates, beyond natural structures is expected to expand. Introduction Carbohydrate active enzymes (CAZymes) are responsible for the biosynthesis and metabolism of all sugars and in Nature they often have to be highly selective in a biochemically complex environment. However, as observed with other enzymes1,2, CAZymes do display ‘promiscuity’, a term that describes the capability of catalysing non-physiological secondary reactions. With the advent of high-throughput screening approaches, many examples of enzyme promiscuity have been identified and discussed in the context of evolution. -

Swiveling Domain Mechanism in Pyruvate Phosphate Dikinase†,‡ Kap Lim,§ Randy J
Biochemistry 2007, 46, 14845-14853 14845 Swiveling Domain Mechanism in Pyruvate Phosphate Dikinase†,‡ Kap Lim,§ Randy J. Read,| Celia C. H. Chen,§ Aleksandra Tempczyk,§ Min Wei,⊥ Dongmei Ye,⊥ Chun Wu,⊥ Debra Dunaway-Mariano,⊥ and Osnat Herzberg*,§ Center for AdVanced Research in Biotechnology, UniVersity of Maryland Biotechnology Institute, RockVille, Maryland 20850, Department of Haematology, Cambridge Institute for Medical Research, UniVersity of Cambridge, Cambridge, United Kingdom, and Department of Chemistry, UniVersity of New Mexico, Albuquerque, New Mexico ReceiVed September 10, 2007; ReVised Manuscript ReceiVed October 17, 2007 ABSTRACT: Pyruvate phosphate dikinase (PPDK) catalyzes the reversible conversion of phosphoenolpyruvate (PEP), AMP, and Pi to pyruvate and ATP. The enzyme contains two remotely located reaction centers: the nucleotide partial reaction takes place at the N-terminal domain, and the PEP/pyruvate partial reaction takes place at the C-terminal domain. A central domain, tethered to the N- and C-terminal domains by two closely associated linkers, contains a phosphorylatable histidine residue (His455). The molecular architecture suggests a swiveling domain mechanism that shuttles a phosphoryl group between the two reaction centers. In an early structure of PPDK from Clostridium symbiosum, the His445-containing domain (His domain) was positioned close to the nucleotide binding domain and did not contact the PEP/pyruvate- binding domain. Here, we present the crystal structure of a second conformational state of C. symbiosum PPDK with the His domain adjacent to the PEP-binding domain. The structure was obtained by producing a three-residue mutant protein (R219E/E271R/S262D) that introduces repulsion between the His and nucleotide-binding domains but preserves viable interactions with the PEP/pyruvate-binding domain. -

Ceramides: Nutrient Signals That Drive Hepatosteatosis
J Lipid Atheroscler. 2020 Jan;9(1):50-65 Journal of https://doi.org/10.12997/jla.2020.9.1.50 Lipid and pISSN 2287-2892·eISSN 2288-2561 Atherosclerosis Review Ceramides: Nutrient Signals that Drive Hepatosteatosis Scott A. Summers Department of Nutrition and Integrative Physiology, University of Utah, Salt Lake City, UT, USA Received: Sep 24, 2019 ABSTRACT Revised: Nov 4, 2019 Accepted: Nov 10, 2019 Ceramides are minor components of the hepatic lipidome that have major effects on liver Correspondence to function. These products of lipid and protein metabolism accumulate when the energy needs Scott A. Summers of the hepatocyte have been met and its storage capacity is full, such that free fatty acids start Department of Nutrition and Integrative to couple to the sphingoid backbone rather than the glycerol moiety that is the scaffold for Physiology, University of Utah, 15N 2030E, Salt Lake City, UT 84112, USA. glycerolipids (e.g., triglycerides) or the carnitine moiety that shunts them into mitochondria. E-mail: [email protected] As ceramides accrue, they initiate actions that protect cells from acute increases in detergent- like fatty acids; for example, they alter cellular substrate preference from glucose to lipids Copyright © 2020 The Korean Society of Lipid and they enhance triglyceride storage. When prolonged, these ceramide actions cause insulin and Atherosclerosis. This is an Open Access article distributed resistance and hepatic steatosis, 2 of the underlying drivers of cardiometabolic diseases. under the terms of the Creative Commons Herein the author discusses the mechanisms linking ceramides to the development of insulin Attribution Non-Commercial License (https:// resistance, hepatosteatosis and resultant cardiometabolic disorders. -

Glucokinase September 15, 2005
Glucokinase September 15, 2005 Summary Glucokinase phosphorylates glucose into glucose 6-phosphate using ATP as the phosphate group donor, according to the reaction glucose + ATP Æ glucose 6-phosphate + ADP This protocol describes an indirect assay to determine the activity of glucokinase. Glucose 6- phosphate formed by glucokinase is measured by the formation of NADPH in presence of glucose 6-phosphate dehydrogenase. Solutions Required 1. 100 mM tris HCl pH 7.4 Stock solution could be used 2. 100 mM MgSO4·7H2O Stock solution could be used 3. 10 mM glucose Must be prepared fresh 4. 20 mM ATP Must be prepared fresh 5. 10 mM NADP Must be prepared fresh 6. a solution containing 10 U of glucose 6-phosphate dehydrogenase per 50 μL in tris HCl pH 7.4 buffer. This solution is prepared by mixing 40 μL of a 5000 U/mL solution (Sigma G8529) with 960 μL tris buffer. Preparation of Cell Extract Follow general protocol described in Preparation of Cell Extract. The cell extract should be suspended in tris buffer after pelletization. Spectrophotometer Turn on the ultraviolet bulb on the spectrophotometer (Beckman DU50) and wait 10 minutes for warm-up. Select the kinetics-time window on the instrument. Load the method "A:/nadh" ". These methods each have a run-time of 120 s, a temperature of 37°C, a wavelength of 340 nm and use 2 autosamplers. Procedure 1. For each assay, prepare the two cocktails shown in the following table into two separate quartz cuvettes. Keep them on ice. Volume (μL) added to: Solution Control Experimental tris 500 500 DI H2O 200 100 MgSO4 100 100 glucose 0 100 ATP 50 50 NADP 50 50 G6P dehydrogenase 50 50 2.