UTR Cleavage of Mrna Transcripts Generates Thousands of Stable Uncapped Autonomous RNA Fragments
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Differences Between Human and Chimpanzee Genomes and Their Implications in Gene Expression, Protein Functions and Biochemical Properties of the Two Species Maria V
Suntsova and Buzdin BMC Genomics 2020, 21(Suppl 7):535 https://doi.org/10.1186/s12864-020-06962-8 REVIEW Open Access Differences between human and chimpanzee genomes and their implications in gene expression, protein functions and biochemical properties of the two species Maria V. Suntsova1 and Anton A. Buzdin1,2,3,4* From 11th International Young Scientists School “Systems Biology and Bioinformatics”–SBB-2019 Novosibirsk, Russia. 24-28 June 2019 Abstract Chimpanzees are the closest living relatives of humans. The divergence between human and chimpanzee ancestors dates to approximately 6,5–7,5 million years ago. Genetic features distinguishing us from chimpanzees and making us humans are still of a great interest. After divergence of their ancestor lineages, human and chimpanzee genomes underwent multiple changes including single nucleotide substitutions, deletions and duplications of DNA fragments of different size, insertion of transposable elements and chromosomal rearrangements. Human-specific single nucleotide alterations constituted 1.23% of human DNA, whereas more extended deletions and insertions cover ~ 3% of our genome. Moreover, much higher proportion is made by differential chromosomal inversions and translocations comprising several megabase-long regions or even whole chromosomes. However, despite of extensive knowledge of structural genomic changes accompanying human evolution we still cannot identify with certainty the causative genes of human identity. Most structural gene-influential changes happened at the level of expression regulation, which in turn provoked larger alterations of interactome gene regulation networks. In this review, we summarized the available information about genetic differences between humans and chimpanzees and their potential functional impacts on differential molecular, anatomical, physiological and cognitive peculiarities of these species. -
Download Validation Data
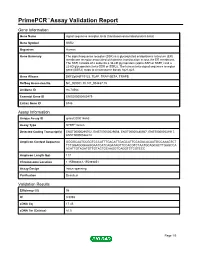
PrimePCR™Assay Validation Report Gene Information Gene Name signal sequence receptor, beta (translocon-associated protein beta) Gene Symbol SSR2 Organism Human Gene Summary The signal sequence receptor (SSR) is a glycosylated endoplasmic reticulum (ER) membrane receptor associated with protein translocation across the ER membrane. The SSR consists of 2 subunits a 34-kD glycoprotein (alpha-SSR or SSR1) and a 22-kD glycoprotein (beta-SSR or SSR2). The human beta-signal sequence receptor gene (SSR2) maps to chromosome bands 1q21-q23. Gene Aliases DKFZp686F19123, TLAP, TRAP-BETA, TRAPB RefSeq Accession No. NC_000001.10, NT_004487.19 UniGene ID Hs.74564 Ensembl Gene ID ENSG00000163479 Entrez Gene ID 6746 Assay Information Unique Assay ID qHsaCID0014663 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000295702, ENST00000529008, ENST00000480567, ENST00000531917, ENST00000526212 Amplicon Context Sequence GGGGCAATCCGGTCCCATTTGACATTGAGCATTCCAGACACAATGCCAAAGTCT TCTGGAGGGAAGGAATCATCAGATAGTTCCACGTCTAATGCAGCACTTGAGCCA ACATTGTAGATGTTGTACTGCAAGGTCAGGTCTCGTCCC Amplicon Length (bp) 117 Chromosome Location 1:155988061-155989851 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 98 R2 0.9998 cDNA Cq 17.45 cDNA Tm (Celsius) 81.5 Page 1/5 PrimePCR™Assay Validation Report gDNA Cq Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report SSR2, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference -

Aneuploidy: Using Genetic Instability to Preserve a Haploid Genome?
Health Science Campus FINAL APPROVAL OF DISSERTATION Doctor of Philosophy in Biomedical Science (Cancer Biology) Aneuploidy: Using genetic instability to preserve a haploid genome? Submitted by: Ramona Ramdath In partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biomedical Science Examination Committee Signature/Date Major Advisor: David Allison, M.D., Ph.D. Academic James Trempe, Ph.D. Advisory Committee: David Giovanucci, Ph.D. Randall Ruch, Ph.D. Ronald Mellgren, Ph.D. Senior Associate Dean College of Graduate Studies Michael S. Bisesi, Ph.D. Date of Defense: April 10, 2009 Aneuploidy: Using genetic instability to preserve a haploid genome? Ramona Ramdath University of Toledo, Health Science Campus 2009 Dedication I dedicate this dissertation to my grandfather who died of lung cancer two years ago, but who always instilled in us the value and importance of education. And to my mom and sister, both of whom have been pillars of support and stimulating conversations. To my sister, Rehanna, especially- I hope this inspires you to achieve all that you want to in life, academically and otherwise. ii Acknowledgements As we go through these academic journeys, there are so many along the way that make an impact not only on our work, but on our lives as well, and I would like to say a heartfelt thank you to all of those people: My Committee members- Dr. James Trempe, Dr. David Giovanucchi, Dr. Ronald Mellgren and Dr. Randall Ruch for their guidance, suggestions, support and confidence in me. My major advisor- Dr. David Allison, for his constructive criticism and positive reinforcement. -

A Crosstalk Between the RNA Binding Protein Smaug and the Hedgehog Pathway Links Cell Signaling to Mrna Regulation in Drosophila Lucía Bruzzone
A crosstalk between the RNA binding protein Smaug and the Hedgehog pathway links cell signaling to mRNA regulation in drosophila Lucía Bruzzone To cite this version: Lucía Bruzzone. A crosstalk between the RNA binding protein Smaug and the Hedgehog pathway links cell signaling to mRNA regulation in drosophila. Cellular Biology. Université Sorbonne Paris Cité, 2018. English. NNT : 2018USPCC234. tel-02899776 HAL Id: tel-02899776 https://tel.archives-ouvertes.fr/tel-02899776 Submitted on 15 Jul 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Thèse de doctorat de l’Université Sorbonne Paris Cité Préparée à l’Université Paris Diderot Ecole doctorale HOB n° 561 Institut Jacques Monod / Equipe Développement, Signalisation et Trafic A crosstalk between the RNA binding protein Smaug and the Hedgehog pathway links cell signaling to mRNA regulation in Drosophila Lucía Bruzzone Thèse de doctorat de Biologie Dirigée par Anne Plessis Présentée et soutenue publiquement à Paris le 19 mars 2018 Président du jury: Alain Zider / Professeur Université Paris Diderot -

Disease-Related Cellular Protein Networks Differentially Affected
www.nature.com/scientificreports OPEN Disease‑related cellular protein networks diferentially afected under diferent EGFR mutations in lung adenocarcinoma Toshihide Nishimura1,8*, Haruhiko Nakamura1,2,8, Ayako Yachie3,8, Takeshi Hase3,8, Kiyonaga Fujii1,8, Hirotaka Koizumi4, Saeko Naruki4, Masayuki Takagi4, Yukiko Matsuoka3, Naoki Furuya5, Harubumi Kato6,7 & Hisashi Saji2 It is unclear how epidermal growth factor receptor EGFR major driver mutations (L858R or Ex19del) afect downstream molecular networks and pathways. This study aimed to provide information on the infuences of these mutations. The study assessed 36 protein expression profles of lung adenocarcinoma (Ex19del, nine; L858R, nine; no Ex19del/L858R, 18). Weighted gene co-expression network analysis together with analysis of variance-based screening identifed 13 co-expressed modules and their eigen proteins. Pathway enrichment analysis for the Ex19del mutation demonstrated involvement of SUMOylation, epithelial and mesenchymal transition, ERK/mitogen- activated protein kinase signalling via phosphorylation and Hippo signalling. Additionally, analysis for the L858R mutation identifed various pathways related to cancer cell survival and death. With regard to the Ex19del mutation, ROCK, RPS6KA1, ARF1, IL2RA and several ErbB pathways were upregulated, whereas AURK and GSKIP were downregulated. With regard to the L858R mutation, RB1, TSC22D3 and DOCK1 were downregulated, whereas various networks, including VEGFA, were moderately upregulated. In all mutation types, CD80/CD86 (B7), MHC, CIITA and IFGN were activated, whereas CD37 and SAFB were inhibited. Costimulatory immune-checkpoint pathways by B7/CD28 were mainly activated, whereas those by PD-1/PD-L1 were inhibited. Our fndings may help identify potential therapeutic targets and develop therapeutic strategies to improve patient outcomes. -

Plasma Cells in Vitro Generation of Long-Lived Human
Downloaded from http://www.jimmunol.org/ by guest on September 24, 2021 is online at: average * The Journal of Immunology , 32 of which you can access for free at: 2012; 189:5773-5785; Prepublished online 16 from submission to initial decision 4 weeks from acceptance to publication November 2012; doi: 10.4049/jimmunol.1103720 http://www.jimmunol.org/content/189/12/5773 In Vitro Generation of Long-lived Human Plasma Cells Mario Cocco, Sophie Stephenson, Matthew A. Care, Darren Newton, Nicholas A. Barnes, Adam Davison, Andy Rawstron, David R. Westhead, Gina M. Doody and Reuben M. Tooze J Immunol cites 65 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription http://www.jimmunol.org/content/suppl/2012/11/16/jimmunol.110372 0.DC1 This article http://www.jimmunol.org/content/189/12/5773.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2012 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 24, 2021. The Journal of Immunology In Vitro Generation of Long-lived Human Plasma Cells Mario Cocco,*,1 Sophie Stephenson,*,1 Matthew A. -

Newly Identified Gon4l/Udu-Interacting Proteins
www.nature.com/scientificreports OPEN Newly identifed Gon4l/ Udu‑interacting proteins implicate novel functions Su‑Mei Tsai1, Kuo‑Chang Chu1 & Yun‑Jin Jiang1,2,3,4,5* Mutations of the Gon4l/udu gene in diferent organisms give rise to diverse phenotypes. Although the efects of Gon4l/Udu in transcriptional regulation have been demonstrated, they cannot solely explain the observed characteristics among species. To further understand the function of Gon4l/Udu, we used yeast two‑hybrid (Y2H) screening to identify interacting proteins in zebrafsh and mouse systems, confrmed the interactions by co‑immunoprecipitation assay, and found four novel Gon4l‑interacting proteins: BRCA1 associated protein‑1 (Bap1), DNA methyltransferase 1 (Dnmt1), Tho complex 1 (Thoc1, also known as Tho1 or HPR1), and Cryptochrome circadian regulator 3a (Cry3a). Furthermore, all known Gon4l/Udu‑interacting proteins—as found in this study, in previous reports, and in online resources—were investigated by Phenotype Enrichment Analysis. The most enriched phenotypes identifed include increased embryonic tissue cell apoptosis, embryonic lethality, increased T cell derived lymphoma incidence, decreased cell proliferation, chromosome instability, and abnormal dopamine level, characteristics that largely resemble those observed in reported Gon4l/udu mutant animals. Similar to the expression pattern of udu, those of bap1, dnmt1, thoc1, and cry3a are also found in the brain region and other tissues. Thus, these fndings indicate novel mechanisms of Gon4l/ Udu in regulating CpG methylation, histone expression/modifcation, DNA repair/genomic stability, and RNA binding/processing/export. Gon4l is a nuclear protein conserved among species. Animal models from invertebrates to vertebrates have shown that the protein Gon4-like (Gon4l) is essential for regulating cell proliferation and diferentiation. -

Investigation of Adiposity Phenotypes in AA Associated with GALNT10 & Related Pathway Genes
Investigation of Adiposity Phenotypes in AA Associated With GALNT10 & Related Pathway Genes By Mary E. Stromberg A Dissertation Submitted to the Graduate Faculty of WAKE FOREST UNIVERSITY GRADUATE SCHOOL OF ARTS AND SCIENCES in Partial Fulfillment of the Requirements for the Degree of DOCTOR OF PHILOSOPHY In Molecular Genetics and Genomics December 2018 Winston-Salem, North Carolina Approved by: Donald W. Bowden, Ph.D., Advisor Maggie C.Y. Ng, Ph.D., Advisor Timothy D. Howard, Ph.D., Chair Swapan Das, Ph.D. John P. Parks, Ph.D. Acknowledgements I would first like to thank my mentors, Dr. Bowden and Dr. Ng, for guiding my learning and growth during my years at Wake Forest University School of Medicine. Thank you Dr. Ng for spending so much time ensuring that I learn every detail of every protocol, and supporting me through personal difficulties over the years. Thank you Dr. Bowden for your guidance in making me a better scientist and person. I would like to thank my committee for their patience and the countless meetings we have had in discussing this project. I would like to say thank you to the members of our lab as well as the Parks lab for their support and friendship as well as their contributions to my project. Special thanks to Dean Godwin for his support and understanding. The umbrella program here at WFU has given me the chance to meet some of the best friends I could have wished for. I would like to also thank those who have taught me along the way and helped me to get to this point of my life, with special thanks to the late Dr. -

Chip-Seq of Transcription Factors Predicts Absolute and Differential Gene Expression in Embryonic Stem Cells
ChIP-Seq of transcription factors predicts absolute and differential gene expression in embryonic stem cells Zhengqing Ouyanga, Qing Zhoub, and Wing Hung Wongc,1 aDepartment of Biology and cDepartments of Statistics, Health Research and Policy, and Biology, Stanford University, Stanford, CA 94305; and bDepartment of Statistics, University of California, Los Angeles, CA 90095 Edited by Terry Speed, University of California, Berkeley, CA, and accepted by the Editorial Board September 25, 2009 (received for review May 5, 2009) Next-generation sequencing has greatly increased the scope and moderate, varying between 9.6% and 36.9% on various datasets the resolution of transcriptional regulation study. RNA sequencing from yeast to human (5, 7, 10, 11), even after considering TF–TF (RNA-Seq) and ChIP-Seq experiments are now generating compre- interaction. The low R2 reported in these studies may be due to hensive data on transcript abundance and on regulator–DNA insufficient data or suboptimal models or both. In any case, interactions. We propose an approach for an integrated analysis of accurate quantitative modeling of expression from binding lo- these data based on feature extraction of ChIP-Seq signals, prin- cation data has not been demonstrated. cipal component analysis, and regression-based component selec- In this paper, we show that accurate quantitative modeling of tion. Compared with traditional methods, our approach not only gene expression data in a mouse cell type is possible provided offers higher power in predicting gene expression from ChIP-Seq that TF-DNA binding locations have been measured by ChIP- data but also provides a way to capture cooperation among Seq experiments for multiple transcriptional regulators. -

Case Report an Adolescent with a Rare De Novo Distal Trisomy 6P and Distal Monosomy 6Q Chromosomal Combination
Hindawi Case Reports in Genetics Volume 2020, Article ID 8857628, 8 pages https://doi.org/10.1155/2020/8857628 Case Report An Adolescent with a Rare De Novo Distal Trisomy 6p and Distal Monosomy 6q Chromosomal Combination Leia A. Peterman , Gail H. Vance, Erin E. Conboy, Katelynn Anderson, and David D. Weaver Department of Medical and Molecular Genetics, Indiana University School of Medicine, Indianapolis, IN, USA Correspondence should be addressed to David D. Weaver; [email protected] Received 15 April 2020; Revised 11 August 2020; Accepted 23 August 2020; Published 31 August 2020 Academic Editor: Silvia Paracchini Copyright © 2020 Leia A. Peterman et al. (is is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. We report on a 12-year-old female with both a partial duplication and deletion involving chromosome 6. (e duplication involves 6p25.3p24.3 (7.585 Mb) while the deletion includes 6q27q27 (6.244 Mb). (is chromosomal abnormality is also described as distal trisomy 6p and distal monosomy 6q. (e patient has a Chiari II malformation, hydrocephalus, agenesis of the corpus callosum, microcephaly, bilateral renal duplicated collecting system, scoliosis, and myelomeningocele associated with a neurogenic bladder and bladder reflux. Additional features have included seizures, feeding dysfunction, failure to thrive, sleep apnea, global de- velopmental delay, intellectual disability, and absent speech. To our knowledge, our report is just the sixth case in the literature with concomitant distal 6p duplication and distal 6q deletion. Although a majority of chromosomal duplication-deletion cases have resulted from a parental pericentric inversion, the parents of our case have normal chromosomes. -

A Concise Review of Human Brain Methylome During Aging and Neurodegenerative Diseases
BMB Rep. 2019; 52(10): 577-588 BMB www.bmbreports.org Reports Invited Mini Review A concise review of human brain methylome during aging and neurodegenerative diseases Renuka Prasad G & Eek-hoon Jho* Department of Life Science, University of Seoul, Seoul 02504, Korea DNA methylation at CpG sites is an essential epigenetic mark position of carbon in the cytosine within CG dinucleotides that regulates gene expression during mammalian development with resultant formation of 5mC. The symmetrical CG and diseases. Methylome refers to the entire set of methylation dinucleotides are also called as CpG, due to the presence of modifications present in the whole genome. Over the last phosphodiester bond between cytosine and guanine. The several years, an increasing number of reports on brain DNA human genome contains short lengths of DNA (∼1,000 bp) in methylome reported the association between aberrant which CpG is commonly located (∼1 per 10 bp) in methylation and the abnormalities in the expression of critical unmethylated form and referred as CpG islands; they genes known to have critical roles during aging and neuro- commonly overlap with the transcription start sites (TSSs) of degenerative diseases. Consequently, the role of methylation genes. In human DNA, 5mC is present in approximately 1.5% in understanding neurodegenerative diseases has been under of the whole genome and CpG base pairs are 5-fold enriched focus. This review outlines the current knowledge of the human in CpG islands than other regions of the genome (3, 4). CpG brain DNA methylomes during aging and neurodegenerative islands have the following salient features. In the human diseases. -

A Regulator of Innate Immune Responses
(19) TZZ ¥_T (11) EP 2 942 357 A1 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 11.11.2015 Bulletin 2015/46 C07K 14/47 (2006.01) A61K 38/00 (2006.01) C12N 15/113 (2010.01) (21) Application number: 15169327.2 (22) Date of filing: 04.08.2009 (84) Designated Contracting States: (72) Inventor: Barber, Glen N. AT BE BG CH CY CZ DE DK EE ES FI FR GB GR Palmetto Bay, FL 33157 (US) HR HU IE IS IT LI LT LU LV MC MK MT NL NO PL PT RO SE SI SK SM TR (74) Representative: Inspicos A/S Kogle Allé 2 (30) Priority: 04.08.2008 US 129975 P P.O. Box 45 2970 Hørsholm (DK) (62) Document number(s) of the earlier application(s) in accordance with Art. 76 EPC: Remarks: 09805473.7 / 2 324 044 This application was filed on 27-05-2015 as a divisional application to the application mentioned (71) Applicant: Barber, Glen N. under INID code 62. Palmetto Bay, FL 33157 (US) (54) STING (STIMULATOR OF INTEFERON GENES), A REGULATOR OF INNATE IMMUNE RESPONSES (57) Novel molecules termed STING which include STING compositions are useful for the treatment of an nucleic acids, polynucleotides, oligonucleotides, pep- immune-related disorder, including treating and prevent- tides, mutants, variants and active fragments thereof, ing infection by modulating immunity. modulate innate and adaptive immunity in a subject. EP 2 942 357 A1 Printed by Jouve, 75001 PARIS (FR) EP 2 942 357 A1 Description RELATED APPLICATIONS 5 [0001] This application claims priority under 35 USC § 119 to U.S.