Quantitative Proteomics Links the LRRC59 Interactome to Mrna Translation on the ER Membrane
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
![FK506-Binding Protein 12.6/1B, a Negative Regulator of [Ca2+], Rescues Memory and Restores Genomic Regulation in the Hippocampus of Aging Rats](https://docslib.b-cdn.net/cover/6136/fk506-binding-protein-12-6-1b-a-negative-regulator-of-ca2-rescues-memory-and-restores-genomic-regulation-in-the-hippocampus-of-aging-rats-16136.webp)
FK506-Binding Protein 12.6/1B, a Negative Regulator of [Ca2+], Rescues Memory and Restores Genomic Regulation in the Hippocampus of Aging Rats
This Accepted Manuscript has not been copyedited and formatted. The final version may differ from this version. A link to any extended data will be provided when the final version is posted online. Research Articles: Neurobiology of Disease FK506-Binding Protein 12.6/1b, a negative regulator of [Ca2+], rescues memory and restores genomic regulation in the hippocampus of aging rats John C. Gant1, Eric M. Blalock1, Kuey-Chu Chen1, Inga Kadish2, Olivier Thibault1, Nada M. Porter1 and Philip W. Landfield1 1Department of Pharmacology & Nutritional Sciences, University of Kentucky, Lexington, KY 40536 2Department of Cell, Developmental and Integrative Biology, University of Alabama at Birmingham, Birmingham, AL 35294 DOI: 10.1523/JNEUROSCI.2234-17.2017 Received: 7 August 2017 Revised: 10 October 2017 Accepted: 24 November 2017 Published: 18 December 2017 Author contributions: J.C.G. and P.W.L. designed research; J.C.G., E.M.B., K.-c.C., and I.K. performed research; J.C.G., E.M.B., K.-c.C., I.K., and P.W.L. analyzed data; J.C.G., E.M.B., O.T., N.M.P., and P.W.L. wrote the paper. Conflict of Interest: The authors declare no competing financial interests. NIH grants AG004542, AG033649, AG052050, AG037868 and McAlpine Foundation for Neuroscience Research Corresponding author: Philip W. Landfield, [email protected], Department of Pharmacology & Nutritional Sciences, University of Kentucky, 800 Rose Street, UKMC MS 307, Lexington, KY 40536 Cite as: J. Neurosci ; 10.1523/JNEUROSCI.2234-17.2017 Alerts: Sign up at www.jneurosci.org/cgi/alerts to receive customized email alerts when the fully formatted version of this article is published. -

Standard Flow Multiplexed Proteomics (Sflompro) – an Accessible and Cost-Effective Alternative to Nanolc Workflows
bioRxiv preprint doi: https://doi.org/10.1101/2020.02.25.964379; this version posted February 25, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Standard Flow Multiplexed Proteomics (SFloMPro) – An Accessible and Cost-Effective Alternative to NanoLC Workflows Conor Jenkins1 and Ben Orsburn2* 1Hood College Department of Biology, Frederick, MD 2University of Virginia Medical School, Charlottesville, VA *To whom correspondence should be addressed; [email protected] Abstract Multiplexed proteomics using isobaric tagging allows for simultaneously comparing the proteomes of multiple samples. In this technique, digested peptides from each sample are labeled with a chemical tag prior to pooling sample for LC-MS/MS with nanoflow chromatography (NanoLC). The isobaric nature of the tag prevents deconvolution of samples until fragmentation liberates the isotopically labeled reporter ions. To ensure efficient peptide labeling, large concentrations of labeling reagents are included in the reagent kits to allow scientists to use high ratios of chemical label per peptide. The increasing speed and sensitivity of mass spectrometers has reduced the peptide concentration required for analysis, leading to most of the label or labeled sample to be discarded. In conjunction, improvements in the speed of sample loading, reliable pump pressure, and stable gradient construction of analytical flow HPLCs has continued to improve the sample delivery process to the mass spectrometer. In this study we describe a method for performing multiplexed proteomics without the use of NanoLC by using offline fractionation of labeled peptides followed by rapid “standard flow” HPLC gradient LC-MS/MS. -

Mutants in Three Novel Complementation Groups Inhibit
Mutants in Three Novel Complementation Groups Inhibit Membrane Protein Insertion into and Soluble Protein Translocation across the Endoplasmic Reticulum Membrane of Saccharomyces cerevisiae Neil Green,* Hong Fang, $ and Peter Walter* *Department ofBiochemistry and Biophysics, School ofMedicine, University of California, San Francisco, California 94143-0448; and tDepartment of Microbiology and Immunology, School ofMedicine, Vanderbilt University, Nashville, Tennessee 37232-2363 Abstract. We have isolated mutants that inhibit mem- tations in SEC61 and SEC63, which were previously brane protein insertion into the ER membrane of Sac- isolated as mutants inhibiting the translocation of solu- charomyces cerevisiae. The mutants were contained in ble proteins, also affect the insertion of membrane three complementation groups, which we have named proteins into the ER. Taken together our data indicate SEC70, SEC1, and SEC72. The mutants also inhib- that the process of protein translocation across the ER ited the translocation of soluble proteins into the lu- membrane involves a much larger number of gene men of the ER, indicating that they pleiotropically products than previously appreciated . Moreover, differ- affect protein transport across and insertion into the ent translocation substrates appear to have different re- ER membrane. Surprisingly, the mutants inhibited the quirements for components of the cellular targeting translocation and insertion of different proteins to dras- and translocation apparatus . tically different degrees. We have also shown that mu- SCENT genetic studies in Saccharomyces cerevisiae Interestingly, not all proteins passing through the secretory have shown that the SEC61, SEC62, and SEC63 pathway were equally affected by mutations in SEC61, SEC- genes are required for secretory protein translocation 62, and SEC63 ; the translocation of pre-invertase showed into the lumen of the ER (Deshaies and Schekman, 1987; little inhibition in mutant cells (Rothblatt et al., 1989) and Rothblatt et al., 1989). -

SEC62 Rabbit Polyclonal Antibody – TA319720 | Origene
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for TA319720 SEC62 Rabbit Polyclonal Antibody Product data: Product Type: Primary Antibodies Applications: IF, IHC, WB Recommended Dilution: WB: 0.5 - 1 ug/mL, ICC: 5 ug/mL, IF: 20 ug/mL Reactivity: Human, Mouse, Rat Host: Rabbit Isotype: IgG Clonality: Polyclonal Immunogen: SEC62 antibody was raised against an 18 amino acid synthetic peptide near the carboxy terminus of human SEC62. Formulation: SEC62 Antibody is supplied in PBS containing 0.02% sodium azide. Concentration: 1ug/ul Purification: SEC62 Antibody is affinity chromatography purified via peptide column. Conjugation: Unconjugated Storage: Store at -20°C as received. Stability: Stable for 12 months from date of receipt. Gene Name: SEC62 homolog, preprotein translocation factor Database Link: NP_003253 Entrez Gene 69276 MouseEntrez Gene 294912 RatEntrez Gene 7095 Human Q99442 Background: SEC62 Antibody: SEC62 is an integral membrane protein located in the rough endoplasmic reticulum (ER) and is part of the SEC61-SEC62-SEC63 complex that is the central component of the protein translocation apparatus of the ER membrane. It is speculated that SEC61- SEC62-SEC63 may perform post-translational protein translocation into the ER and might also perform the backward transport of ER proteins that are subject to the ubiquitin-proteasome- dependent degradation pathway. Silencing of this gene with RNAi dramatically reduced the migration and invasive potential of numerous tumor cell lines, suggesting that it may be an attractive target for therapy of various tumors. -

Sec62/Ki67 and P16/Ki67 Dual-Staining Immunocytochemistry
Archives of Gynecology and Obstetrics (2019) 299:825–833 https://doi.org/10.1007/s00404-018-5021-0 GYNECOLOGIC ONCOLOGY Sec62/Ki67 and p16/Ki67 dual‑staining immunocytochemistry in vulvar cytology for the identifcation of vulvar intraepithelial neoplasia and vulvar cancer: a pilot study Ferenc Zoltan Takacs1 · Julia Caroline Radosa1 · Florian Bochen2 · Ingolf Juhasz‑Böss1 · Erich‑Franz Solomayer1 · Rainer M. Bohle3 · Georg‑Peter Breitbach1 · Bernard Schick2 · Maximilian Linxweiler2 Received: 22 October 2018 / Accepted: 12 December 2018 / Published online: 4 January 2019 © Springer-Verlag GmbH Germany, part of Springer Nature 2019 Abstract Purpose The aim of this study was to analyze the diagnostic performance of a newly established immunocytochemical dual-staining protocol for the simultaneous expression of SEC62 and Ki67 in vulvar liquid-based cytology specimens for the identifcation of vulvar intraepithelial neoplasia (VIN) and vulvar cancer. In addition, we investigated the p16/Ki67 dual stain, which has already been established in cervical cytology. Materials and methods For this pilot study, residual material from liquid-based cytology was collected retrospectively from 45 women. The presence of one or more double-immunoreactive cells was considered as a positive test result for Sec62/Ki67 and p16/Ki67 dual staining. The test results were correlated with the course of histology. Results All cases of VIN and vulvar cancer were Sec62/Ki67 and p16/Ki67 dual-stain positive, and normal and low-grade squamous intraepithelial lesions were all negative. The sensitivity of cytology for VIN + cases was 100% (22/22), whereas punch biopsy classifed one case of vulvar carcinoma as infammation. All cases with high-intensity (grades 3 and 4) Sec62 staining in Sec62/Ki67-positive cases were carcinomas. -
Targeted Quantitative Proteomics Using Selected Reaction Monitoring
Targeted Quantitative Proteomics Using Selected Reaction Monitoring (SRM)- Mass Spectrometry Coupled with an 18O-labeled Reference as Internal Standards Jong-Seo Kim, Errol Robinson, Boyd L. Champion, Brianne O. Petritis, Thomas L. Fillmore, Ronald J. Moore, Liu Tao, David G. Camp II, Richard D. Smith, and Wei-Jun Qian Biological Sciences Division, Pacific Northwest National Laboratory, Richland, WA Methods Results Effect of Q1 resolution Reproducibility 18O labeling efficiency Overview Patient Samples Conclusions Group A: • To address the need of stable isotope Group B: • The utility of 18O-labeled “universal” O area ratio O area Trypsin 18 Red error bars: label-free peak area ratio labeled internal standards for accurate (= each 16O area / average 18O area) reference as internal standards for targeted O/ digestion 16 18 quantification, we introduce the use of an 16 Green error bars: each pair area ratio of O/ O quantitative proteomics has been Patient peptide 18 Pooled reference ratio O area O-labeled reference as comprehensive samples sample 18 successfully demonstrated 16 18 LC-SRM-MS O/ Concentration ratio ( O/ O) internal standards for accurate SRM-MS- 18O labeling 16 – A linear dynamic range of quantification Concen. based quantification instead of using labeled 18 0.01 0.02 0.04 0.1 0.2 1 5 10 25 50 100 4 O-labeled “universal” Ratio ~ 10 in relative concentration synthetic peptides. reference 16O/18O 6.1 12.5 13.7 13.8 5.5 7.7 16.8 2.8 4.6 4.2 1.6 pairs – Better reproducibility than label-free 18 Label • O-labeling efficiency for most peptides is 39.2 23.4 47.5 26.5 19.3 10.6 31.2 3.2 10.7 4.8 3.5 Fig. -

Quantitative Proteomics Reveals the Selectivity of Ubiquitin-Binding Autophagy Receptors in the Turnover of Damaged Lysosomes by Lysophagy
bioRxiv preprint doi: https://doi.org/10.1101/2021.07.19.452535; this version posted July 19, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Quantitative proteomics reveals the selectivity of ubiquitin-binding autophagy receptors in the turnover of damaged lysosomes by lysophagy Vinay Eapen1,*, Sharan Swarup1,*, Melissa Hoyer1,*, Joao Paolo1 and J. Wade Harper 1 1Department of Cell Biology, Harvard Medical School, Boston MA 02115 *, Equal Contribution Send correspondence to: [email protected] ABSTRACT Removal of damaged organelles via the process of selective autophagy constitutes a major form of cellular quality control. Damaged organelles are recognized by a dedicated surveillance machinery, leading to the assembly of an autophagosome around the damaged organelle, prior to fusion with the degradative lysosomal compartment. Lysosomes themselves are also prone to damage and are degraded through the process of lysophagy. While early steps involve recognition of ruptured lysosomal membranes by glycan-binding Galectins and ubiquitylation of transmembrane lysosomal proteins, many steps in the process, and their inter-relationships, remain poorly understood, including the role and identity of cargo receptors required for completion of lysophagy. Here, we employ quantitative organelle capture and proximity biotinylation proteomics of autophagy adaptors, cargo receptors, and Galectins in response to acute lysosomal damage, thereby revealing the landscape of lysosomal proteome remodeling during lysophagy. Among proteins dynamically recruited to damaged lysosomes were ubiquitin-binding autophagic cargo receptors. -
Download Validation Data
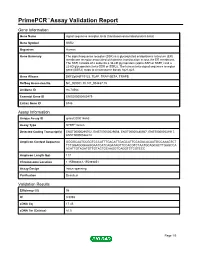
PrimePCR™Assay Validation Report Gene Information Gene Name signal sequence receptor, beta (translocon-associated protein beta) Gene Symbol SSR2 Organism Human Gene Summary The signal sequence receptor (SSR) is a glycosylated endoplasmic reticulum (ER) membrane receptor associated with protein translocation across the ER membrane. The SSR consists of 2 subunits a 34-kD glycoprotein (alpha-SSR or SSR1) and a 22-kD glycoprotein (beta-SSR or SSR2). The human beta-signal sequence receptor gene (SSR2) maps to chromosome bands 1q21-q23. Gene Aliases DKFZp686F19123, TLAP, TRAP-BETA, TRAPB RefSeq Accession No. NC_000001.10, NT_004487.19 UniGene ID Hs.74564 Ensembl Gene ID ENSG00000163479 Entrez Gene ID 6746 Assay Information Unique Assay ID qHsaCID0014663 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000295702, ENST00000529008, ENST00000480567, ENST00000531917, ENST00000526212 Amplicon Context Sequence GGGGCAATCCGGTCCCATTTGACATTGAGCATTCCAGACACAATGCCAAAGTCT TCTGGAGGGAAGGAATCATCAGATAGTTCCACGTCTAATGCAGCACTTGAGCCA ACATTGTAGATGTTGTACTGCAAGGTCAGGTCTCGTCCC Amplicon Length (bp) 117 Chromosome Location 1:155988061-155989851 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 98 R2 0.9998 cDNA Cq 17.45 cDNA Tm (Celsius) 81.5 Page 1/5 PrimePCR™Assay Validation Report gDNA Cq Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report SSR2, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference -

Ma-And-Mayr-Cell-2018.Pdf
Article A Membraneless Organelle Associated with the Endoplasmic Reticulum Enables 30UTR-Mediated Protein-Protein Interactions Graphical Abstract Authors Weirui Ma, Christine Mayr Correspondence [email protected] In Brief ER-associated granules foster selective protein-protein interactions dictated by the nucleotide composition of the encoding mRNA. Highlights d TIS granules are membraneless organelles intertwined with the endoplasmic reticulum d TIS11B forms TIS granules that enrich or exclude specific mRNAs and proteins in vivo d TIS granules enable translation of mRNAs with AU-rich elements at an ER subdomain d Specific protein-protein interactions can only be formed in the TIS granule region Ma & Mayr, 2018, Cell 175, 1–15 November 29, 2018 ª 2018 Elsevier Inc. https://doi.org/10.1016/j.cell.2018.10.007 Please cite this article in press as: Ma and Mayr, A Membraneless Organelle Associated with the Endoplasmic Reticulum Enables 30UTR- Mediated Protein-Protein Interactions, Cell (2018), https://doi.org/10.1016/j.cell.2018.10.007 Article A Membraneless Organelle Associated with the Endoplasmic Reticulum Enables 30UTR-Mediated Protein-Protein Interactions Weirui Ma1 and Christine Mayr1,2,* 1Cancer Biology and Genetics Program, Memorial Sloan Kettering Cancer Center, New York, NY 10065, USA 2Lead Contact *Correspondence: [email protected] https://doi.org/10.1016/j.cell.2018.10.007 SUMMARY cesses, including signaling and the addition of post-translational modifications (Li et al., 2012; Su et al., 2016). Approximately half of human genes generate mRNAs One type of membraneless organelles are RNA granules that with alternative 30 untranslated regions (30UTRs). contain mRNA transcripts packaged into ribonucleoprotein as- Through 30UTR-mediated protein-protein interac- semblies (Han et al., 2012). -

Mass Spectrometry and Proteomics Using R/Bioconductor
Mass spectrometry and proteomics Using R/Bioconductor Laurent Gatto CSAMA - Bressanone - 25 July 2019 1 / 40 Slides available at: http://bit.ly/20190725csama These slides are available under a creative common CC-BY license. You are free to share (copy and redistribute the material in any medium or format) and adapt (remix, transform, and build upon the material) for any purpose, even commercially . 2 / 40 On the menu Morning lecture: 1. Proteomics in R/Bioconductor 2. How does mass spectrometry-based proteomics work? 3. Quantitative proteomics 4. Quantitative proteomics data processing and analysis Afternoon lab: Manipulating MS data (raw and identification data) Manipulating quantitative proteomics data Data processing and DE 3 / 40 4 / 40 1. Proteomics and mass spectrometry packages, questions and workow in Bioconductor. 5 / 40 2. How does mass spectrometry work? (applies to proteomics and metabolomics) 6 / 40 Overview 7 / 40 How does MS work? 1. Digestion of proteins into peptides - as will become clear later, the features we measure in shotgun (or bottom-up) proteomics are peptides, not proteins. 2. On-line liquid chromatography (LC-MS) 3. Mass spectrometry (MS) is a technology that separates charged molecules (ions, peptides) based on their mass to charge ratio (M/Z). 8 / 40 Chromatography MS is generally coupled to chromatography (liquid LC, but can also be gas- based GC). The time an analytes takes to elute from the chromatography column is the retention time. 9 / 40 An mass spectrometer is composed of three components: 1. The source, that ionises the molecules: examples are Matrix-assisted laser desorption/ionisation (MALDI) or electrospray ionisation (ESI). -

Aneuploidy: Using Genetic Instability to Preserve a Haploid Genome?
Health Science Campus FINAL APPROVAL OF DISSERTATION Doctor of Philosophy in Biomedical Science (Cancer Biology) Aneuploidy: Using genetic instability to preserve a haploid genome? Submitted by: Ramona Ramdath In partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biomedical Science Examination Committee Signature/Date Major Advisor: David Allison, M.D., Ph.D. Academic James Trempe, Ph.D. Advisory Committee: David Giovanucci, Ph.D. Randall Ruch, Ph.D. Ronald Mellgren, Ph.D. Senior Associate Dean College of Graduate Studies Michael S. Bisesi, Ph.D. Date of Defense: April 10, 2009 Aneuploidy: Using genetic instability to preserve a haploid genome? Ramona Ramdath University of Toledo, Health Science Campus 2009 Dedication I dedicate this dissertation to my grandfather who died of lung cancer two years ago, but who always instilled in us the value and importance of education. And to my mom and sister, both of whom have been pillars of support and stimulating conversations. To my sister, Rehanna, especially- I hope this inspires you to achieve all that you want to in life, academically and otherwise. ii Acknowledgements As we go through these academic journeys, there are so many along the way that make an impact not only on our work, but on our lives as well, and I would like to say a heartfelt thank you to all of those people: My Committee members- Dr. James Trempe, Dr. David Giovanucchi, Dr. Ronald Mellgren and Dr. Randall Ruch for their guidance, suggestions, support and confidence in me. My major advisor- Dr. David Allison, for his constructive criticism and positive reinforcement. -

A Crosstalk Between the RNA Binding Protein Smaug and the Hedgehog Pathway Links Cell Signaling to Mrna Regulation in Drosophila Lucía Bruzzone
A crosstalk between the RNA binding protein Smaug and the Hedgehog pathway links cell signaling to mRNA regulation in drosophila Lucía Bruzzone To cite this version: Lucía Bruzzone. A crosstalk between the RNA binding protein Smaug and the Hedgehog pathway links cell signaling to mRNA regulation in drosophila. Cellular Biology. Université Sorbonne Paris Cité, 2018. English. NNT : 2018USPCC234. tel-02899776 HAL Id: tel-02899776 https://tel.archives-ouvertes.fr/tel-02899776 Submitted on 15 Jul 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Thèse de doctorat de l’Université Sorbonne Paris Cité Préparée à l’Université Paris Diderot Ecole doctorale HOB n° 561 Institut Jacques Monod / Equipe Développement, Signalisation et Trafic A crosstalk between the RNA binding protein Smaug and the Hedgehog pathway links cell signaling to mRNA regulation in Drosophila Lucía Bruzzone Thèse de doctorat de Biologie Dirigée par Anne Plessis Présentée et soutenue publiquement à Paris le 19 mars 2018 Président du jury: Alain Zider / Professeur Université Paris Diderot