The Contribution of Syntrophic Fatty-Acid Degrading Microbial Communities to Anaerobic Digester Function and Stability
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Crystalline Iron Oxides Stimulate Methanogenic Benzoate Degradation in Marine Sediment-Derived Enrichment Cultures
The ISME Journal https://doi.org/10.1038/s41396-020-00824-7 ARTICLE Crystalline iron oxides stimulate methanogenic benzoate degradation in marine sediment-derived enrichment cultures 1,2 1 3 1,2 1 David A. Aromokeye ● Oluwatobi E. Oni ● Jan Tebben ● Xiuran Yin ● Tim Richter-Heitmann ● 2,4 1 5 5 1 2,3 Jenny Wendt ● Rolf Nimzyk ● Sten Littmann ● Daniela Tienken ● Ajinkya C. Kulkarni ● Susann Henkel ● 2,4 2,4 1,3 2,3,4 1,2 Kai-Uwe Hinrichs ● Marcus Elvert ● Tilmann Harder ● Sabine Kasten ● Michael W. Friedrich Received: 19 May 2020 / Revised: 9 October 2020 / Accepted: 22 October 2020 © The Author(s) 2020. This article is published with open access Abstract Elevated dissolved iron concentrations in the methanic zone are typical geochemical signatures of rapidly accumulating marine sediments. These sediments are often characterized by co-burial of iron oxides with recalcitrant aromatic organic matter of terrigenous origin. Thus far, iron oxides are predicted to either impede organic matter degradation, aiding its preservation, or identified to enhance organic carbon oxidation via direct electron transfer. Here, we investigated the effect of various iron oxide phases with differing crystallinity (magnetite, hematite, and lepidocrocite) during microbial 1234567890();,: 1234567890();,: degradation of the aromatic model compound benzoate in methanic sediments. In slurry incubations with magnetite or hematite, concurrent iron reduction, and methanogenesis were stimulated during accelerated benzoate degradation with methanogenesis as the dominant electron sink. In contrast, with lepidocrocite, benzoate degradation, and methanogenesis were inhibited. These observations were reproducible in sediment-free enrichments, even after five successive transfers. Genes involved in the complete degradation of benzoate were identified in multiple metagenome assembled genomes. -
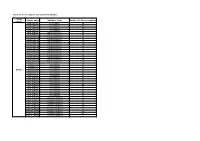
Table S1: List of Samples Included in the Analysis
Table S1: list of samples included in the analysis Study Sample name Inhibitory status Number of days at sampling number DNA.0P2T4 No inhibition 29 DNA.0P2T6 No inhibition 57 DNA.10P2T4 No inhibition 29 DNA.10P2T6 No inhibition 57 DNA.75P2T4 Phenol inhibition 29 DNA.75P2T6 Phenol inhibition 57 DNA.100P2T4 Phenol inhibition 29 DNA.100P2T6 Phenol inhibition 57 DNA.125P1T4 Phenol inhibition 29 DNA.125P1T6 Phenol inhibition 57 DNA.125P2T4 Phenol inhibition 29 DNA.125P2T6 Phenol inhibition 57 DNA.125P3T4 Phenol inhibition 29 DNA.125P3T6 Phenol inhibition 57 DNA.150P2T4 Phenol inhibition 29 DNA.150P2T6 Phenol inhibition 57 DNA.200P2T4 Phenol inhibition 29 DNA.200P2T6 Phenol inhibition 57 DNA.0N2T4 No inhibition 29 DNA.0N2T5 No inhibition 42 DNA.0N2T6 No inhibition 57 Study 1 DNA.5N2T4 No inhibition 29 DNA.5N2T5 No inhibition 42 DNA.5N2T6 No inhibition 57 DNA.10N2T4 No inhibition 29 DNA.10N2T5 No inhibition 42 DNA.10N2T6 No inhibition 57 DNA.15N2T4 No inhibition 29 DNA.15N2T5 No inhibition 42 DNA.15N2T6 No inhibition 57 DNA.25N2T4 No inhibition 29 DNA.25N2T5 No inhibition 42 DNA.25N2T6 No inhibition 57 DNA.75N2T4 Ammonia inhibition 29 DNA.75N2T5 Ammonia inhibition 42 DNA.75N2T6 Ammonia inhibition 57 DNA.100N2T4 Ammonia inhibition 29 DNA.100N2T5 Ammonia inhibition 42 DNA.100N2T6 Ammonia inhibition 57 DNA.250N2T4 Ammonia inhibition 29 DNA.250N2T5 Ammonia inhibition 42 DNA.250N2T6 Ammonia inhibition 57 nono2T3 No inhibition 16 noN2T4 Ammonia inhibition 23 noN2T8 Ammonia inhibition 60 noN2T9 Ammonia inhibition 85 noPhi2T4 Phenol inhibition 23 noPhi2T5 -

Biorremediación De Metales Pesados Por Sulfidogénesis Utilizando Comunidades Y Microorganismos Sulfato-Reductores
FACULTAD DE CIENCIAS EXACTAS DEPARTAMENTO DE QUÍMICA BIORREMEDIACIÓN DE METALES PESADOS POR SULFIDOGÉNESIS UTILIZANDO COMUNIDADES Y MICROORGANISMOS SULFATO-REDUCTORES Trabajo de Tesis Doctoral Graciana Willis Poratti Director: Dr. Edgardo Donati Año 2016 Este Trabajo de Tesis Doctoral es presentado para optar al grado de Doctor de la Facultad de Ciencias Exactas Fue realizado en CINDEFI (CCT La Plata-CONICET, UNLP) A Mamá, Papá y Juan, por acompañarme siempre Reconocimientos A la Facultad de Ciencias Exactas de la Universidad Nacional de La Plata y al Consejo Nacional de Investigaciones Científicas y Técnicas (CONICET) por haber hecho posible la realización del presente Trabajo de Tesis Doctoral. Al CINDEFI (Centro de Investigación en Fermentaciones Industriales) y a toda la gente que lo conforma por haberme brindado el lugar físico para la realización del presente trabajo. Al Prof. Dr. David Barrie Johnson (Bangor Acidophile Research Team) de la Universidad de Bangor, Gales por permitirme realizar parte del trabajo de Tesis Doctoral en su laboratorio. A los Dres. Goh Kian Mau de la Universiti Teknologi, Malaysia y Kok-Gan Chan de la University of Malaysa, por la colaboración en la secuenciación del genoma de la cepa LMa1. Al Ente Provincial de Termas de Neuquén (EPROTEN) por permitir la toma de muestra en el Parque Geotermal Copahue. Al grupo de la Dra. Alejandra Giaveno perteneciente al IDEPA (CONICET, UNCo) y a la Facultad de Ingeniería de la Universidad Nacional del Comahue, por la organización de las campañas de muestreo a la zona de Copahue. Agradecimientos A mi director Dr. Edgardo Donati, por haber depositado su confianza en la realización de este trabajo, por dirigirme y haberme dado la oportunidad de viajar y formarme. -

EXPERIMENTAL STUDIES on FERMENTATIVE FIRMICUTES from ANOXIC ENVIRONMENTS: ISOLATION, EVOLUTION, and THEIR GEOCHEMICAL IMPACTS By
EXPERIMENTAL STUDIES ON FERMENTATIVE FIRMICUTES FROM ANOXIC ENVIRONMENTS: ISOLATION, EVOLUTION, AND THEIR GEOCHEMICAL IMPACTS By JESSICA KEE EUN CHOI A dissertation submitted to the School of Graduate Studies Rutgers, The State University of New Jersey In partial fulfillment of the requirements For the degree of Doctor of Philosophy Graduate Program in Microbial Biology Written under the direction of Nathan Yee And approved by _______________________________________________________ _______________________________________________________ _______________________________________________________ _______________________________________________________ New Brunswick, New Jersey October 2017 ABSTRACT OF THE DISSERTATION Experimental studies on fermentative Firmicutes from anoxic environments: isolation, evolution and their geochemical impacts by JESSICA KEE EUN CHOI Dissertation director: Nathan Yee Fermentative microorganisms from the bacterial phylum Firmicutes are quite ubiquitous in subsurface environments and play an important biogeochemical role. For instance, fermenters have the ability to take complex molecules and break them into simpler compounds that serve as growth substrates for other organisms. The research presented here focuses on two groups of fermentative Firmicutes, one from the genus Clostridium and the other from the class Negativicutes. Clostridium species are well-known fermenters. Laboratory studies done so far have also displayed the capability to reduce Fe(III), yet the mechanism of this activity has not been investigated -

Syntrophic Butyrate and Propionate Oxidation Processes 491
Environmental Microbiology Reports (2010) 2(4), 489–499 doi:10.1111/j.1758-2229.2010.00147.x Minireview Syntrophic butyrate and propionate oxidation processes: from genomes to reaction mechanismsemi4_147 489..499 Nicolai Müller,1† Petra Worm,2† Bernhard Schink,1* a cytoplasmic fumarate reductase to drive energy- Alfons J. M. Stams2 and Caroline M. Plugge2 dependent succinate oxidation. Furthermore, we 1Faculty for Biology, University of Konstanz, D-78457 propose that homologues of the Thermotoga mar- Konstanz, Germany. itima bifurcating [FeFe]-hydrogenase are involved 2Laboratory of Microbiology, Wageningen University, in NADH oxidation by S. wolfei and S. fumaroxidans Dreijenplein 10, 6703 HB Wageningen, the Netherlands. to form hydrogen. Summary Introduction In anoxic environments such as swamps, rice fields In anoxic environments such as swamps, rice paddy fields and sludge digestors, syntrophic microbial communi- and intestines of higher animals, methanogenic commu- ties are important for decomposition of organic nities are important for decomposition of organic matter to matter to CO2 and CH4. The most difficult step is the CO2 and CH4 (Schink and Stams, 2006; Mcinerney et al., fermentative degradation of short-chain fatty acids 2008; Stams and Plugge, 2009). Moreover, they are the such as propionate and butyrate. Conversion of these key biocatalysts in anaerobic bioreactors that are used metabolites to acetate, CO2, formate and hydrogen is worldwide to treat industrial wastewaters and solid endergonic under standard conditions and occurs wastes. Different types of anaerobes have specified only if methanogens keep the concentrations of these metabolic functions in the degradation pathway and intermediate products low. Butyrate and propionate depend on metabolite transfer which is called syntrophy degradation pathways include oxidation steps of (Schink and Stams, 2006). -

Bibliography
Bibliography Abella, C.A., X.P. Cristina, A. Martinez, I. Pibernat and X. Vila. 1998. on moderate concentrations of acetate: production of single cells. Two new motile phototrophic consortia: "Chlorochromatium lunatum" Appl. Microbiol. Biotechnol. 35: 686-689. and "Pelochromatium selenoides". Arch. Microbiol. 169: 452-459. Ahring, B.K, P. Westermann and RA. Mah. 1991b. Hydrogen inhibition Abella, C.A and LJ. Garcia-Gil. 1992. Microbial ecology of planktonic of acetate metabolism and kinetics of hydrogen consumption by Me filamentous phototrophic bacteria in holomictic freshwater lakes. Hy thanosarcina thermophila TM-I. Arch. Microbiol. 157: 38-42. drobiologia 243-244: 79-86. Ainsworth, G.C. and P.H.A Sheath. 1962. Microbial Classification: Ap Acca, M., M. Bocchetta, E. Ceccarelli, R Creti, KO. Stetter and P. Cam pendix I. Symp. Soc. Gen. Microbiol. 12: 456-463. marano. 1994. Updating mass and composition of archaeal and bac Alam, M. and D. Oesterhelt. 1984. Morphology, function and isolation terial ribosomes. Archaeal-like features of ribosomes from the deep of halobacterial flagella. ]. Mol. Biol. 176: 459-476. branching bacterium Aquifex pyrophilus. Syst. Appl. Microbiol. 16: 629- Albertano, P. and L. Kovacik. 1994. Is the genus LeptolynglYya (Cyano 637. phyte) a homogeneous taxon? Arch. Hydrobiol. Suppl. 105: 37-51. Achenbach-Richter, L., R Gupta, KO. Stetter and C.R Woese. 1987. Were Aldrich, H.C., D.B. Beimborn and P. Schönheit. 1987. Creation of arti the original eubacteria thermophiles? Syst. Appl. Microbiol. 9: 34- factual internal membranes during fixation of Methanobacterium ther 39. moautotrophicum. Can.]. Microbiol. 33: 844-849. Adams, D.G., D. Ashworth and B. -

Microbial Communities Mediating Algal Detritus Turnover Under Anaerobic Conditions
Microbial communities mediating algal detritus turnover under anaerobic conditions Jessica M. Morrison1,*, Chelsea L. Murphy1,*, Kristina Baker1, Richard M. Zamor2, Steve J. Nikolai2, Shawn Wilder3, Mostafa S. Elshahed1 and Noha H. Youssef1 1 Department of Microbiology and Molecular Genetics, Oklahoma State University, Stillwater, OK, USA 2 Grand River Dam Authority, Vinita, OK, USA 3 Department of Integrative Biology, Oklahoma State University, Stillwater, OK, USA * These authors contributed equally to this work. ABSTRACT Background. Algae encompass a wide array of photosynthetic organisms that are ubiquitously distributed in aquatic and terrestrial habitats. Algal species often bloom in aquatic ecosystems, providing a significant autochthonous carbon input to the deeper anoxic layers in stratified water bodies. In addition, various algal species have been touted as promising candidates for anaerobic biogas production from biomass. Surprisingly, in spite of its ecological and economic relevance, the microbial community involved in algal detritus turnover under anaerobic conditions remains largely unexplored. Results. Here, we characterized the microbial communities mediating the degradation of Chlorella vulgaris (Chlorophyta), Chara sp. strain IWP1 (Charophyceae), and kelp Ascophyllum nodosum (phylum Phaeophyceae), using sediments from an anaerobic spring (Zodlteone spring, OK; ZDT), sludge from a secondary digester in a local wastewater treatment plant (Stillwater, OK; WWT), and deeper anoxic layers from a seasonally stratified lake -

Und Buttersäure-Bildenden Bakterien Aus Biogasanlagen
Johannes Gutenberg-Universität Mainz Institut für Mikrobiologie und Weinforschung Isolierung von Essigsäure-, Propionsäure- und Buttersäure-bildenden Bakterien aus Biogasanlagen Dissertation Zur Erlangung des Grades Doktor der Naturwissenschaften Am Fachbereich Biologie Der Johannes Gutenberg-Universität Mainz Katharina Gabriela Cibis geboren am 19. Juni 1987 in Worms Mainz, 2015 Dekan: 1. Berichterstatter: 2. Berichterstatter: Tag der mündlichen Prüfung: 3. Dezember 2015 Die Ergebnisse dieser Arbeit sind teilweise in folgenden Publikationen und Posterpräsentatio- nen veröffentlicht oder sind zur Publikation eingereicht: INHALTSVERZEICHNIS Inhaltsverzeichnis 1. Einleitung 1 1.1 Biogasanlagen: Gewinnung von Energie und Forschungsschwerpunkt 1 1.2 Mikrobiologische Prozesse in Biogasanlagen 4 1.3 Aufbau und Betrieb von Biogasanlagen 8 1.4 Substrate für den Betrieb von Biogasanlagen 12 1.5 Mikrobielle Bildung von Essigsäure, Propionsäure und Buttersäure 13 1.6 Ziele der Arbeit 23 2. Material und Methoden 25 2.1 Chemikalien und Gase 25 2.2 Biochemikalien, Enzyme und Kits 27 2.3 Geräte und Hilfsmittel 28 2.4 Verbrauchsmaterialien 30 2.5 Mikrobiologische Methoden 31 2.5.1 Organismen 31 2.5.2 Untersuchte Biogasanlagen 31 2.5.3 Nährmedien zur Isolierung und Kultivierung von Säure-bildenden Bakterien 33 2.5.4 Methoden zur Isolierung von Säure-bildenden Bakterien 37 2.5.5 Physiologische Charakterisierung der Isolate 39 2.6 Analytische Methode: Hochleistungsflüssigkeitschromatographie 41 2.7 Molekularbiologische Methoden 42 2.7.1 DNA-Isolierung aus -

Contents Topic 1. Introduction to Microbiology. the Subject and Tasks
Contents Topic 1. Introduction to microbiology. The subject and tasks of microbiology. A short historical essay………………………………………………………………5 Topic 2. Systematics and nomenclature of microorganisms……………………. 10 Topic 3. General characteristics of prokaryotic cells. Gram’s method ………...45 Topic 4. Principles of health protection and safety rules in the microbiological laboratory. Design, equipment, and working regimen of a microbiological laboratory………………………………………………………………………….162 Topic 5. Physiology of bacteria, fungi, viruses, mycoplasmas, rickettsia……...185 TOPIC 1. INTRODUCTION TO MICROBIOLOGY. THE SUBJECT AND TASKS OF MICROBIOLOGY. A SHORT HISTORICAL ESSAY. Contents 1. Subject, tasks and achievements of modern microbiology. 2. The role of microorganisms in human life. 3. Differentiation of microbiology in the industry. 4. Communication of microbiology with other sciences. 5. Periods in the development of microbiology. 6. The contribution of domestic scientists in the development of microbiology. 7. The value of microbiology in the system of training veterinarians. 8. Methods of studying microorganisms. Microbiology is a science, which study most shallow living creatures - microorganisms. Before inventing of microscope humanity was in dark about their existence. But during the centuries people could make use of processes vital activity of microbes for its needs. They could prepare a koumiss, alcohol, wine, vinegar, bread, and other products. During many centuries the nature of fermentations remained incomprehensible. Microbiology learns morphology, physiology, genetics and microorganisms systematization, their ecology and the other life forms. Specific Classes of Microorganisms Algae Protozoa Fungi (yeasts and molds) Bacteria Rickettsiae Viruses Prions The Microorganisms are extraordinarily widely spread in nature. They literally ubiquitous forward us from birth to our death. Daily, hourly we eat up thousands and thousands of microbes together with air, water, food. -

Pelospora Glutarica Gen. Nov., Sp. Nov., a Glutarate-Fermenting, Strictly Anaerobic, Spore-Forming Bacterium
International Journal of Systematic and Evolutionary Microbiology (2000), 50, 645–648 Printed in Great Britain Pelospora glutarica gen. nov., sp. nov., a glutarate-fermenting, strictly anaerobic, spore-forming bacterium Carola Matthies,1 Nina Springer,2 Wolfgang Ludwig2 and Bernhard Schink3 Author for correspondence: Bernhard Schink. Tel: j49 7531 882140. Fax: j49 7531 882966. e-mail: bernhard.schink!uni-konstanz.de 1 Lehrstuhl fu$ rO$ kologische The strictly anaerobic, Gram-negative, spore-forming bacterium strain WoGl3T $ Mikrobiologie, BITOK, had been enriched and isolated in mineral medium with glutarate as the sole Universita$ t Bayreuth, 95440 Bayreuth, Germany source of energy and organic carbon. Glutarate was fermented to a mixture of butyrate, isobutyrate, CO and small amounts of acetate. Strain WoGl3T grew 2 Lehrstuhl fu$ r 2 Mikrobiologie, Technische only with the dicarboxylates glutarate, methylsuccinate and succinate. 16S Universita$ tMu$ nchen, rDNA sequence analysis revealed an affiliation of strain WoGl3T to the family Arcisstr. 16, 80290 Syntrophomonadaceae. This monophyletic group is comprised of strain WoGl3T Mu$ nchen, Germany and the genera Syntrophomonas, Syntrophospora and Thermosyntropha, 3 Lehrstuhl fu$ r Mikrobielle within the phylum of Gram-positive bacteria with a low DNA GMC content. O$ kologie, Fakulta$ tfu$ r Biologie, Universita$ t Overall intra-group 16S rRNA sequence similarities of 892–939% document a Konstanz, Fach M 654, separate phylogenetic status for strain WoGl3T. Strain WoGl3T (l DSM 6652T)is 78457 Konstanz, Germany described as the type strain of a new species within a new genus, Pelospora glutarica gen. nov., sp. nov. Keywords: Pelospora glutarica sp. nov., anaerobic degradation, dicarboxylic acids, glutarate INTRODUCTION and nitrate-reducing pseudomonads (Anders et al., 1995). -

Characterization of an Anaerobic Population Digesting a Model Substrate for Maize in the Presence of Trace Metals
Chemosphere 80 (2010) 829–836 Contents lists available at ScienceDirect Chemosphere journal homepage: www.elsevier.com/locate/chemosphere Characterization of an anaerobic population digesting a model substrate for maize in the presence of trace metals Herbert Pobeheim, Bernhard Munk, Henry Müller, Gabriele Berg, Georg M. Guebitz * Institute of Environmental Biotechnology, Graz University of Technology, Petersgasse 12, 8010 Graz, Austria article info abstract Article history: The influence of a defined trace metal solution and additionally Ni2+ on anaerobic digestion of biomass Received 1 December 2009 was investigated. A novel synthetic model substrate was designed consisting of cellulose, starch and urea Received in revised form 6 June 2010 as carbon and nitrogen source in a ratio mimicking the basic composition of maize silage. Two indepen- Accepted 7 June 2010 dent batch fermentations were carried out over 21 d with the synthetic model substrate in the presence Available online 7 July 2010 of the trace metal solution. Particularly an increase in nickel concentrations (17 and 34 lM) enhanced methane formation by up to 20%. This increased activity was also corroborated by fluorescence micros- Keywords: copy measurements based on cofactor F . The eubacterial and methanogenic population was character- Anaerobic digestion 420 ized with the single strand conformational polymorphism analysis and the amplified 16S rDNA Biogas Cellulose restriction analysis of 16S rRNA genes amplified by different primer systems. Nearly the half of the ana- Nickel lyzed bacteria were identified as Firmicutes while 70% in this phylum belonged to the class of Clostridiales SSCP and 30% to the class of Bacilli. Bacteroides and uncultured bacteria represented each a quarter of the ana- lyzed community. -

Comparative Genomics of Syntrophic Branched-Chain Fatty Acid Degrading Bacteria
Microbes Environ. Vol. 31, No. 3, 288-292, 2016 https://www.jstage.jst.go.jp/browse/jsme2 doi:10.1264/jsme2.ME16057 Comparative Genomics of Syntrophic Branched-Chain Fatty Acid Degrading Bacteria TAKASHI NARIHIRO1,2†*, MASARU K. NOBU2†, HIDEYUKI TAMAKI1,3, YOICHI KAMAGATA1, YUJI SEKIGUCHI4, and WEN-TSO LIU2 1Bioproduction Research Institute, National Institute of Advanced Industrial Science and Technology (AIST), Central 6, Higashi 1–1–1, Tsukuba, Ibaraki 305–8566, Japan; 2Department of Civil and Environmental Engineering, University of Illinois at Urbana-Champaign, 205 North Mathews Ave, Urbana, IL 61801, USA; 3Biotechnology Research Institute, The University of Tokyo, 1–1–1 Yayoi, Bunkyo-ku, Tokyo 113–8657, Japan; and 4Biomedical Research Institute, National Institute of Advanced Industrial Science and Technology (AIST), Central 6, Higashi 1–1–1, Tsukuba, Ibaraki 305–8566, Japan (Received March 16, 2016—Accepted June 5, 2016—Published online July 16, 2016) The syntrophic degradation of branched-chain fatty acids (BCFAs) such as 2-methylbutyrate and isobutyrate is an essential step in the production of methane from proteins/amino acids in anaerobic ecosystems. While a few syntrophic BCFA-degrading bacteria have been isolated, their metabolic pathways in BCFA and short-chain fatty acid (SCFA) degradation as well as energy conservation systems remain unclear. In an attempt to identify these pathways, we herein performed comparative genomics of three syntrophic bacteria: 2-methylbutyrate-degrading “Syntrophomonas wolfei subsp. methylbutyratica” strain JCM 14075T (=4J5T), isobutyrate-degrading Syntrophothermus lipocalidus strain TGB-C1T, and non-BCFA-metabolizing S. wolfei subsp. wolfei strain GöttingenT. We demonstrated that 4J5 and TGB-C1 both encode multiple genes/gene clusters involved in β-oxidation, as observed in the Göttingen genome, which has multiple copies of genes associated with butyrate degradation.