1 Regulation of Neuronal Mrna Splicing and Tau Isoform Ratio by ATXN3
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
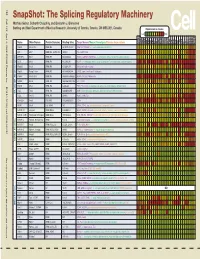
Snapshot: the Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J
192 Cell SnapShot: The Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J. Blencowe 133 Banting and Best Department of Medical Research, University of Toronto, Toronto, ON M5S 3E1, Canada Expression in mouse , April4, 2008©2008Elsevier Inc. Low High Name Other Names Protein Domains Binding Sites Target Genes/Mouse Phenotypes/Disease Associations Amy Ceb Hip Hyp OB Eye SC BM Bo Ht SM Epd Kd Liv Lu Pan Pla Pro Sto Spl Thy Thd Te Ut Ov E6.5 E8.5 E10.5 SRp20 Sfrs3, X16 RRM, RS GCUCCUCUUC SRp20, CT/CGRP; −/− early embryonic lethal E3.5 9G8 Sfrs7 RRM, RS, C2HC Znf (GAC)n Tau, GnRH, 9G8 ASF/SF2 Sfrs1 RRM, RS RGAAGAAC HipK3, CaMKIIδ, HIV RNAs; −/− embryonic lethal, cond. KO cardiomyopathy SC35 Sfrs2 RRM, RS UGCUGUU AChE; −/− embryonic lethal, cond. KO deficient T-cell maturation, cardiomyopathy; LS SRp30c Sfrs9 RRM, RS CUGGAUU Glucocorticoid receptor SRp38 Fusip1, Nssr RRM, RS ACAAAGACAA CREB, type II and type XI collagens SRp40 Sfrs5, HRS RRM, RS AGGAGAAGGGA HipK3, PKCβ-II, Fibronectin SRp55 Sfrs6 RRM, RS GGCAGCACCUG cTnT, CD44 DOI 10.1016/j.cell.2008.03.010 SRp75 Sfrs4 RRM, RS GAAGGA FN1, E1A, CD45; overexpression enhances chondrogenic differentiation Tra2α Tra2a RRM, RS GAAARGARR GnRH; overexpression promotes RA-induced neural differentiation SR and SR-Related Proteins Tra2β Sfrs10 RRM, RS (GAA)n HipK3, SMN, Tau SRm160 Srrm1 RS, PWI AUGAAGAGGA CD44 SWAP Sfrs8 RS, SWAP ND SWAP, CD45, Tau; possible asthma susceptibility gene hnRNP A1 Hnrnpa1 RRM, RGG UAGGGA/U HipK3, SMN2, c-H-ras; rheumatoid arthritis, systemic lupus -

Aneuploidy: Using Genetic Instability to Preserve a Haploid Genome?
Health Science Campus FINAL APPROVAL OF DISSERTATION Doctor of Philosophy in Biomedical Science (Cancer Biology) Aneuploidy: Using genetic instability to preserve a haploid genome? Submitted by: Ramona Ramdath In partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biomedical Science Examination Committee Signature/Date Major Advisor: David Allison, M.D., Ph.D. Academic James Trempe, Ph.D. Advisory Committee: David Giovanucci, Ph.D. Randall Ruch, Ph.D. Ronald Mellgren, Ph.D. Senior Associate Dean College of Graduate Studies Michael S. Bisesi, Ph.D. Date of Defense: April 10, 2009 Aneuploidy: Using genetic instability to preserve a haploid genome? Ramona Ramdath University of Toledo, Health Science Campus 2009 Dedication I dedicate this dissertation to my grandfather who died of lung cancer two years ago, but who always instilled in us the value and importance of education. And to my mom and sister, both of whom have been pillars of support and stimulating conversations. To my sister, Rehanna, especially- I hope this inspires you to achieve all that you want to in life, academically and otherwise. ii Acknowledgements As we go through these academic journeys, there are so many along the way that make an impact not only on our work, but on our lives as well, and I would like to say a heartfelt thank you to all of those people: My Committee members- Dr. James Trempe, Dr. David Giovanucchi, Dr. Ronald Mellgren and Dr. Randall Ruch for their guidance, suggestions, support and confidence in me. My major advisor- Dr. David Allison, for his constructive criticism and positive reinforcement. -

A Novel Resveratrol Analog: Its Cell Cycle Inhibitory, Pro-Apoptotic and Anti-Inflammatory Activities on Human Tumor Cells
A NOVEL RESVERATROL ANALOG : ITS CELL CYCLE INHIBITORY, PRO-APOPTOTIC AND ANTI-INFLAMMATORY ACTIVITIES ON HUMAN TUMOR CELLS A dissertation submitted to Kent State University in partial fulfillment of the requirements for the degree of Doctor of Philosophy by Boren Lin May 2006 Dissertation written by Boren Lin B.S., Tunghai University, 1996 M.S., Kent State University, 2003 Ph. D., Kent State University, 2006 Approved by Dr. Chun-che Tsai , Chair, Doctoral Dissertation Committee Dr. Bryan R. G. Williams , Co-chair, Doctoral Dissertation Committee Dr. Johnnie W. Baker , Members, Doctoral Dissertation Committee Dr. James L. Blank , Dr. Bansidhar Datta , Dr. Gail C. Fraizer , Accepted by Dr. Robert V. Dorman , Director, School of Biomedical Sciences Dr. John R. Stalvey , Dean, College of Arts and Sciences ii TABLE OF CONTENTS LIST OF FIGURES……………………………………………………………….………v LIST OF TABLES……………………………………………………………………….vii ACKNOWLEDGEMENTS….………………………………………………………….viii I INTRODUCTION….………………………………………………….1 Background and Significance……………………………………………………..1 Specific Aims………………………………………………………………………12 II MATERIALS AND METHODS.…………………………………………….16 Cell Culture and Compounds…….……………….…………………………….….16 MTT Cell Viability Assay………………………………………………………….16 Trypan Blue Exclusive Assay……………………………………………………...18 Flow Cytometry for Cell Cycle Analysis……………..……………....……………19 DNA Fragmentation Assay……………………………………………...…………23 Caspase-3 Activity Assay………………………………...……….….…….………24 Annexin V-FITC Staining Assay…………………………………..…...….………28 NF-kappa B p65 Activity Assay……………………………………..………….…29 -

Gene Expression During Normal and FSHD Myogenesis Tsumagari Et Al
Gene expression during normal and FSHD myogenesis Tsumagari et al. Tsumagari et al. BMC Medical Genomics 2011, 4:67 http://www.biomedcentral.com/1755-8794/4/67 (27 September 2011) Tsumagari et al. BMC Medical Genomics 2011, 4:67 http://www.biomedcentral.com/1755-8794/4/67 RESEARCHARTICLE Open Access Gene expression during normal and FSHD myogenesis Koji Tsumagari1, Shao-Chi Chang1, Michelle Lacey2,3, Carl Baribault2,3, Sridar V Chittur4, Janet Sowden5, Rabi Tawil5, Gregory E Crawford6 and Melanie Ehrlich1,3* Abstract Background: Facioscapulohumeral muscular dystrophy (FSHD) is a dominant disease linked to contraction of an array of tandem 3.3-kb repeats (D4Z4) at 4q35. Within each repeat unit is a gene, DUX4, that can encode a protein containing two homeodomains. A DUX4 transcript derived from the last repeat unit in a contracted array is associated with pathogenesis but it is unclear how. Methods: Using exon-based microarrays, the expression profiles of myogenic precursor cells were determined. Both undifferentiated myoblasts and myoblasts differentiated to myotubes derived from FSHD patients and controls were studied after immunocytochemical verification of the quality of the cultures. To further our understanding of FSHD and normal myogenesis, the expression profiles obtained were compared to those of 19 non-muscle cell types analyzed by identical methods. Results: Many of the ~17,000 examined genes were differentially expressed (> 2-fold, p < 0.01) in control myoblasts or myotubes vs. non-muscle cells (2185 and 3006, respectively) or in FSHD vs. control myoblasts or myotubes (295 and 797, respectively). Surprisingly, despite the morphologically normal differentiation of FSHD myoblasts to myotubes, most of the disease-related dysregulation was seen as dampening of normal myogenesis- specific expression changes, including in genes for muscle structure, mitochondrial function, stress responses, and signal transduction. -

Identification of Differentially Expressed Genes in Human Bladder Cancer Through Genome-Wide Gene Expression Profiling
521-531 24/7/06 18:28 Page 521 ONCOLOGY REPORTS 16: 521-531, 2006 521 Identification of differentially expressed genes in human bladder cancer through genome-wide gene expression profiling KAZUMORI KAWAKAMI1,3, HIDEKI ENOKIDA1, TOKUSHI TACHIWADA1, TAKENARI GOTANDA1, KENGO TSUNEYOSHI1, HIROYUKI KUBO1, KENRYU NISHIYAMA1, MASAKI TAKIGUCHI2, MASAYUKI NAKAGAWA1 and NAOHIKO SEKI3 1Department of Urology, Graduate School of Medical and Dental Sciences, Kagoshima University, 8-35-1 Sakuragaoka, Kagoshima 890-8520; Departments of 2Biochemistry and Genetics, and 3Functional Genomics, Graduate School of Medicine, Chiba University, 1-8-1 Inohana, Chuo-ku, Chiba 260-8670, Japan Received February 15, 2006; Accepted April 27, 2006 Abstract. Large-scale gene expression profiling is an effective CKS2 gene not only as a potential biomarker for diagnosing, strategy for understanding the progression of bladder cancer but also for staging human BC. This is the first report (BC). The aim of this study was to identify genes that are demonstrating that CKS2 expression is strongly correlated expressed differently in the course of BC progression and to with the progression of human BC. establish new biomarkers for BC. Specimens from 21 patients with pathologically confirmed superficial (n=10) or Introduction invasive (n=11) BC and 4 normal bladder samples were studied; samples from 14 of the 21 BC samples were subjected Bladder cancer (BC) is among the 5 most common to microarray analysis. The validity of the microarray results malignancies worldwide, and the 2nd most common tumor of was verified by real-time RT-PCR. Of the 136 up-regulated the genitourinary tract and the 2nd most common cause of genes we detected, 21 were present in all 14 BCs examined death in patients with cancer of the urinary tract (1-7). -

Mouse Srsf7 Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Srsf7 Knockout Project (CRISPR/Cas9) Objective: To create a Srsf7 knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Srsf7 gene (NCBI Reference Sequence: NM_146083.2 ; Ensembl: ENSMUSG00000024097 ) is located on Mouse chromosome 17. 8 exons are identified, with the ATG start codon in exon 1 and the TGA stop codon in exon 8 (Transcript: ENSMUST00000063417). Exon 3~8 will be selected as target site. Cas9 and gRNA will be co-injected into fertilized eggs for KO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Exon 3 starts from about 29.41% of the coding region. Exon 3~8 covers 70.73% of the coding region. The size of effective KO region: ~3889 bp. The KO region does not have any other known gene. Page 1 of 9 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele 5' gRNA region gRNA region 3' 1 2 3 4 5 6 7 8 Legends Exon of mouse Srsf7 Knockout region Page 2 of 9 https://www.alphaknockout.com Overview of the Dot Plot (up) Window size: 15 bp Forward Reverse Complement Sequence 12 Note: The 2000 bp section upstream of Exon 3 is aligned with itself to determine if there are tandem repeats. No significant tandem repeat is found in the dot plot matrix. So this region is suitable for PCR screening or sequencing analysis. Overview of the Dot Plot (down) Window size: 15 bp Forward Reverse Complement Sequence 12 Note: The 2000 bp section downstream of stop codon is aligned with itself to determine if there are tandem repeats. -

Ultraconserved Elements Are Associated with Homeostatic Control of Splicing Regulators by Alternative Splicing and Nonsense-Mediated Decay
Downloaded from genesdev.cshlp.org on September 24, 2021 - Published by Cold Spring Harbor Laboratory Press Ultraconserved elements are associated with homeostatic control of splicing regulators by alternative splicing and nonsense-mediated decay Julie Z. Ni,1 Leslie Grate,1 John Paul Donohue,1 Christine Preston,2 Naomi Nobida,2 Georgeann O’Brien,2 Lily Shiue,1 Tyson A. Clark,3 John E. Blume,3 and Manuel Ares Jr.1,2,4 1Center for Molecular Biology of RNA and Department of Molecular, Cell, and Developmental Biology, University of California at Santa Cruz, Santa Cruz, California 95064, USA; 2Hughes Undergraduate Research Laboratory, University of California at Santa Cruz, Santa Cruz, California 95064, USA; 3Affymetrix, Inc., Santa Clara, California 95051, USA Many alternative splicing events create RNAs with premature stop codons, suggesting that alternative splicing coupled with nonsense-mediated decay (AS-NMD) may regulate gene expression post-transcriptionally. We tested this idea in mice by blocking NMD and measuring changes in isoform representation using splicing-sensitive microarrays. We found a striking class of highly conserved stop codon-containing exons whose inclusion renders the transcript sensitive to NMD. A genomic search for additional examples identified >50 such exons in genes with a variety of functions. These exons are unusually frequent in genes that encode splicing activators and are unexpectedly enriched in the so-called “ultraconserved” elements in the mammalian lineage. Further analysis show that NMD of mRNAs for splicing activators such as SR proteins is triggered by splicing activation events, whereas NMD of the mRNAs for negatively acting hnRNP proteins is triggered by splicing repression, a polarity consistent with widespread homeostatic control of splicing regulator gene expression. -

Insertional Mutagenesis Identifies Multiple Networks of Cooperating Genes Driving Intestinal Tumorigenesis
Edinburgh Research Explorer Insertional mutagenesis identifies multiple networks of cooperating genes driving intestinal tumorigenesis Citation for published version: March, HN, Rust, AG, Wright, NA, ten Hoeve, J, de Ridder, J, Eldridge, M, van der Weyden, L, Berns, A, Gadiot, J, Uren, A, Kemp, R, Arends, MJ, Wessels, LFA, Winton, DJ & Adams, DJ 2011, 'Insertional mutagenesis identifies multiple networks of cooperating genes driving intestinal tumorigenesis', Nature Genetics, vol. 43, no. 12, pp. 1202-9. https://doi.org/10.1038/ng.990 Digital Object Identifier (DOI): 10.1038/ng.990 Link: Link to publication record in Edinburgh Research Explorer Document Version: Peer reviewed version Published In: Nature Genetics General rights Copyright for the publications made accessible via the Edinburgh Research Explorer is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The University of Edinburgh has made every reasonable effort to ensure that Edinburgh Research Explorer content complies with UK legislation. If you believe that the public display of this file breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 07. Oct. 2021 Europe PMC Funders Group Author Manuscript Nat Genet. Author manuscript; available in PMC 2012 June 01. Published in final edited form as: Nat Genet. ; 43(12): 1202–1209. doi:10.1038/ng.990. Europe PMC Funders Author Manuscripts Insertional mutagenesis identifies multiple networks of co- operating genes driving intestinal tumorigenesis H. -

SFRS7-Mediated Splicing of Tau Exon 10 Is Directly Regulated by STOX1A in Glial Cells
SFRS7-Mediated Splicing of Tau Exon 10 Is Directly Regulated by STOX1A in Glial Cells Daan van Abel1, Dennis R. Ho¨ lzel2, Shushant Jain2, Fiona M. F. Lun3, Yama W. L. Zheng3, Eric Z. Chen3, Hao Sun3, Rossa W. K. Chiu3, Y. M. Dennis Lo3, Marie van Dijk1, Cees B. M. Oudejans1* 1 Department of Clinical Chemistry, VU University Medical Center, Amsterdam, The Netherlands, 2 Department of Clinical Genetics, Section Medical Genomics, VU University Medical Center, Amsterdam, The Netherlands, 3 Department of Chemical Pathology, Prince of Wales Hospital, Shatin, New Territories, Hong Kong SAR, China Abstract Background: In this study, we performed a genome-wide search for effector genes bound by STOX1A, a winged helix transcription factor recently demonstrated to be involved in late onset Alzheimer’s disease and affecting the amyloid processing pathway. Methodology/Principal Findings: Our results show that out of 218 genes bound by STOX1A as identified by chromatin- immunoprecipitation followed by sequencing (ChIP-Seq), the serine/arginine-rich splicing factor 7 (SFRS7) was found to be induced, both at the mRNA and protein levels, by STOX1A after stable transfection in glial cells. The increase in SFRS7 was followed by an increase in the 4R/3R ratios of the microtubule-associated protein tau (MAPT) by differential exon 10 splicing. Secondly, STOX1A also induced expression of total tau both at the mRNA and protein levels. Upregulation of total tau expression (SFRS7-independent) and tau exon 10 splicing (SFRS7-dependent), as shown in this study to be both affected by STOX1A, is known to have implications in neurodegeneration. Conclusions: Our data further supports the functional importance and central role of STOX1A in neurodegeneration. -

Renoprotective Effect of Combined Inhibition of Angiotensin-Converting Enzyme and Histone Deacetylase
BASIC RESEARCH www.jasn.org Renoprotective Effect of Combined Inhibition of Angiotensin-Converting Enzyme and Histone Deacetylase † ‡ Yifei Zhong,* Edward Y. Chen, § Ruijie Liu,*¶ Peter Y. Chuang,* Sandeep K. Mallipattu,* ‡ ‡ † | ‡ Christopher M. Tan, § Neil R. Clark, § Yueyi Deng, Paul E. Klotman, Avi Ma’ayan, § and ‡ John Cijiang He* ¶ *Department of Medicine, Mount Sinai School of Medicine, New York, New York; †Department of Nephrology, Longhua Hospital, Shanghai University of Traditional Chinese Medicine, Shanghai, China; ‡Department of Pharmacology and Systems Therapeutics and §Systems Biology Center New York, Mount Sinai School of Medicine, New York, New York; |Baylor College of Medicine, Houston, Texas; and ¶Renal Section, James J. Peters Veterans Affairs Medical Center, New York, New York ABSTRACT The Connectivity Map database contains microarray signatures of gene expression derived from approximately 6000 experiments that examined the effects of approximately 1300 single drugs on several human cancer cell lines. We used these data to prioritize pairs of drugs expected to reverse the changes in gene expression observed in the kidneys of a mouse model of HIV-associated nephropathy (Tg26 mice). We predicted that the combination of an angiotensin-converting enzyme (ACE) inhibitor and a histone deacetylase inhibitor would maximally reverse the disease-associated expression of genes in the kidneys of these mice. Testing the combination of these inhibitors in Tg26 mice revealed an additive renoprotective effect, as suggested by reduction of proteinuria, improvement of renal function, and attenuation of kidney injury. Furthermore, we observed the predicted treatment-associated changes in the expression of selected genes and pathway components. In summary, these data suggest that the combination of an ACE inhibitor and a histone deacetylase inhibitor could have therapeutic potential for various kidney diseases. -

Report a Recessive Contiguous Gene Deletion of Chromosome 2P16
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Am. J. Hum. Genet. 69:869–875, 2001 Report A Recessive Contiguous Gene Deletion of Chromosome 2p16 Associated with Cystinuria and a Mitochondrial Disease Ruti Parvari,1 Irena Brodyansky,2 Orly Elpeleg,3 Shimon Moses,2 Daniel Landau,2 and Eli Hershkovitz2 1Department of Developmental Molecular Genetics, Faculty of Health Sciences, and 2Department of Pediatrics, Soroka Medical Center and Faculty of Health Sciences, Ben Gurion University of the Negev, Beer Sheva, Israel; and 3Metabolic Disease Unit, Shaare Zedek Medical Center, Jerusalem Deletions ranging from 100 Kb to 1 Mb—too small to be detected under the microscope—may still involve dozens of genes, thus causing microdeletion syndromes. The vast majority of these syndromes are caused by haploinsuf- ficiency of one or several genes and are transmitted as dominant traits. We identified seven patients originating from an extended family and presenting with a unique syndrome, inherited in a recessive mode, consisting of cystinuria, neonatal seizures, hypotonia, severe somatic and developmental delay, facial dysmorphism, and lactic acidemia. Reduced activity of all the respiratory chain enzymatic complexes that are encoded in the mitochondria was found in muscle biopsy specimens of the patients examined. The molecular basis of this disorder is a homozygous deletion of 179,311 bp on chromosome 2p16, which includes the type I cystinuria gene (SLC3A1), the protein phosphatase 2Cb gene (PP2Cb), an unidentified gene (KIAA0436), and several expressed sequence tags. The extent of the deletion suggests that this unique syndrome is related to the complete absence of these genes’ products, one of which may be essential for the synthesis of mitochondrial encoded proteins. -

Understanding the Alternative Splicing-Regulated Transcriptomic Changes
University of Connecticut OpenCommons@UConn Doctoral Dissertations University of Connecticut Graduate School 10-6-2015 Understanding the Alternative Splicing-regulated Transcriptomic Changes During Mouse Retinal Development and Disease Through Global and Gene-centric Approaches Devi Krishna Priya Karunakaran University of Connecticut - Storrs, [email protected] Follow this and additional works at: https://opencommons.uconn.edu/dissertations Recommended Citation Karunakaran, Devi Krishna Priya, "Understanding the Alternative Splicing-regulated Transcriptomic Changes During Mouse Retinal Development and Disease Through Global and Gene-centric Approaches" (2015). Doctoral Dissertations. 930. https://opencommons.uconn.edu/dissertations/930 ABSTRACT Understanding the Alternative Splicing-regulated Transcriptomic Changes During Mouse Retinal Development and Disease Through Global and Gene-centric Approaches Devi Krishna Priya Karunakaran, Ph.D University of Connecticut, 2015 Alternative splicing (AS) is an important layer of gene regulation and has been shown to control various cellular processes including splicing, mRNA export, translation and cell cycle. Mis-regulation in AS has been implicated in many diseases. But, the role of AS in the retinal development and diseases remains unexplored. To this end, I employed a two- pronged approach, (i.e), gene-centric approach and an en-mass transcriptome analysis approach to address this question. For the first approach, I studied the role of an alternative splicing factor, Sfrs10 and a kinase, Citron Kinase (CitK) and its spliced isoforms in murine retinal development and diseases. Expression analysis of Sfrs10 in mouse and human retinae showed that unlike mouse, it was not expressed in normal human retina but was observed only in AMD retina, suggesting a specific role in response to oxidative stress.