Altered Gene Expression in Circulating Immune Cells Following a 24-Hour Passive Dehydration
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Metallothionein-Protein Interactions
DOI 10.1515/bmc-2012-0049 BioMol Concepts 2013; 4(2): 143–160 Review S í lvia Atrian * and Merc è Capdevila Metallothionein-protein interactions Abstract: Metallothioneins (MTs) are a family of univer- Introduction sal, small proteins, sharing a high cysteine content and an optimal capacity for metal ion coordination. They take Metallothioneins (MTs) are a family of small ( < 10 kDa), part in a plethora of metal ion-related events (from detoxi- extremely heterogeneous proteins, sharing a high cysteine fication to homeostasis, storage, and delivery), in a wide content (15 – 30 % ) that confers them an optimal capacity range of stress responses, and in different pathological for metal ion coordination. After their discovery in horse processes (tumorigenesis, neurodegeneration, and inflam- kidneys by Bert Vallee in 1957 (1) , MTs have been identi- mation). The information on both intracellular and extra- fied and characterized in most prokaryotic and all eukary- cellular interactions of MTs with other proteins is here otic organisms. Besides metal ion detoxification, they comprehensively reviewed. In mammalian kidney, MT1/ have been related to a plethora of physiological events, MT2 interact with megalin and related receptors, and with from the homeostasis, storage, and delivery of physiologi- the transporter transthyretin. Most of the mammalian MT cal metals, to the defense against a wide range of stresses partners identified concern interactions with central nerv- and pathological processes (tumor genesis, neurodegen- ous system (mainly brain) proteins, both through physical eration, inflammation, etc.). It is now a common agree- contact or metal exchange reactions. Physical interactions ment among MT researchers that the ambiguity when mainly involve neuronal secretion multimers. -

Diseasespecific and Inflammationindependent Stromal
Full Length Arthritis & Rheumatism DOI 10.1002/art.37704 Disease-specific and inflammation-independent stromal alterations in spondyloarthritis synovitis Nataliya Yeremenko1,2, Troy Noordenbos1,2, Tineke Cantaert1,3, Melissa van Tok1,2, Marleen van de Sande1, Juan D. Cañete4, Paul P. Tak1,5*, Dominique Baeten1,2 1Department of Clinical Immunology and Rheumatology and 2Department of Experimental Immunology, Academic Medical Center/University of Amsterdam, the Netherlands. 3Department of Immunobiology, Yale University School of Medicine, New Haven, CT, USA. 4Department of Rheumatology, Hospital Clinic de Barcelona and IDIBAPS, Spain. 5Arthrogen B.V., Amsterdam, the Netherlands. *Currently also: GlaxoSmithKline, Stevenage, U.K. Corresponding author: Dominique Baeten, MD, PhD, Department of Clinical Immunology and Rheumatology, F4-105, Academic Medical Center/University of Amsterdam, Meibergdreef 9, 1105 AZ Amsterdam, The Netherlands. E-mail: [email protected] This article has been accepted for publication and undergone full peer review but has not been through the copyediting, typesetting, pagination and proofreading process which may lead to differences between this version and the Version of Record. Please cite this article as an ‘Accepted Article’, doi: 10.1002/art.37704 © 2012 American College of Rheumatology Received: Apr 11, 2012; Revised: Jul 25, 2012; Accepted: Sep 06, 2012 Arthritis & Rheumatism Page 2 of 36 Abstract Objective: The molecular processes driving the distinct patterns of synovial inflammation and tissue remodelling in spondyloarthritis (SpA) versus rheumatoid arthritis (RA) remain largely unknown. Therefore, we aimed to identify novel and unsuspected disease- specific pathways in SpA by a systematic and unbiased synovial gene expression analysis. Methods: Differentially expressed genes were identified by pan-genomic microarray and confirmed by quantitative PCR and immunohistochemistry using synovial tissue biopsies of SpA (n=63), RA (n=28) and gout (n=9) patients. -

Identification of the Binding Partners for Hspb2 and Cryab Reveals
Brigham Young University BYU ScholarsArchive Theses and Dissertations 2013-12-12 Identification of the Binding arP tners for HspB2 and CryAB Reveals Myofibril and Mitochondrial Protein Interactions and Non- Redundant Roles for Small Heat Shock Proteins Kelsey Murphey Langston Brigham Young University - Provo Follow this and additional works at: https://scholarsarchive.byu.edu/etd Part of the Microbiology Commons BYU ScholarsArchive Citation Langston, Kelsey Murphey, "Identification of the Binding Partners for HspB2 and CryAB Reveals Myofibril and Mitochondrial Protein Interactions and Non-Redundant Roles for Small Heat Shock Proteins" (2013). Theses and Dissertations. 3822. https://scholarsarchive.byu.edu/etd/3822 This Thesis is brought to you for free and open access by BYU ScholarsArchive. It has been accepted for inclusion in Theses and Dissertations by an authorized administrator of BYU ScholarsArchive. For more information, please contact [email protected], [email protected]. Identification of the Binding Partners for HspB2 and CryAB Reveals Myofibril and Mitochondrial Protein Interactions and Non-Redundant Roles for Small Heat Shock Proteins Kelsey Langston A thesis submitted to the faculty of Brigham Young University in partial fulfillment of the requirements for the degree of Master of Science Julianne H. Grose, Chair William R. McCleary Brian Poole Department of Microbiology and Molecular Biology Brigham Young University December 2013 Copyright © 2013 Kelsey Langston All Rights Reserved ABSTRACT Identification of the Binding Partners for HspB2 and CryAB Reveals Myofibril and Mitochondrial Protein Interactors and Non-Redundant Roles for Small Heat Shock Proteins Kelsey Langston Department of Microbiology and Molecular Biology, BYU Master of Science Small Heat Shock Proteins (sHSP) are molecular chaperones that play protective roles in cell survival and have been shown to possess chaperone activity. -

Number 2 February 2017 Atlas of Genetics and Cytogenetics in Oncology and Haematology
Volume 1 - Number 1 May - September 1997 Volume 21 - Number 2 February 2017 Atlas of Genetics and Cytogenetics in Oncology and Haematology OPEN ACCESS JOURNAL INIST-CNRS Scope The Atlas of Genetics and Cytogenetics in Oncology and Haematology is a peer reviewed on-line journal in open access, devoted to genes, cytogenetics, and clinical entities in cancer, and cancer-prone diseases. It is made for and by: clinicians and researchers in cytogenetics, molecular biology, oncology, haematology, and pathology. One main scope of the Atlas is to conjugate the scientific information provided by cytogenetics/molecular genetics to the clinical setting (diagnostics, prognostics and therapeutic design), another is to provide an encyclopedic knowledge in cancer genetics. The Atlas deals with cancer research and genomics. It is at the crossroads of research, virtual medical university (university and post-university e-learning), and telemedicine. It contributes to "meta-medicine", this mediation, using information technology, between the increasing amount of knowledge and the individual, having to use the information. Towards a personalized medicine of cancer. It presents structured review articles ("cards") on: 1- Genes, 2- Leukemias, 3- Solid tumors, 4- Cancer-prone diseases, and also 5- "Deep insights": more traditional review articles on the above subjects and on surrounding topics. It also present 6- Case reports in hematology and 7- Educational items in the various related topics for students in Medicine and in Sciences. The Atlas of Genetics and Cytogenetics in Oncology and Haematology does not publish research articles. See also: http://documents.irevues.inist.fr/bitstream/handle/2042/56067/Scope.pdf Editorial correspondance Jean-Loup Huret, MD, PhD, Genetics, Department of Medical Information, University Hospital F-86021 Poitiers, France phone +33 5 49 44 45 46 [email protected] or [email protected] . -

Supplementary Table 3 Complete List of RNA-Sequencing Analysis of Gene Expression Changed by ≥ Tenfold Between Xenograft and Cells Cultured in 10%O2
Supplementary Table 3 Complete list of RNA-Sequencing analysis of gene expression changed by ≥ tenfold between xenograft and cells cultured in 10%O2 Expr Log2 Ratio Symbol Entrez Gene Name (culture/xenograft) -7.182 PGM5 phosphoglucomutase 5 -6.883 GPBAR1 G protein-coupled bile acid receptor 1 -6.683 CPVL carboxypeptidase, vitellogenic like -6.398 MTMR9LP myotubularin related protein 9-like, pseudogene -6.131 SCN7A sodium voltage-gated channel alpha subunit 7 -6.115 POPDC2 popeye domain containing 2 -6.014 LGI1 leucine rich glioma inactivated 1 -5.86 SCN1A sodium voltage-gated channel alpha subunit 1 -5.713 C6 complement C6 -5.365 ANGPTL1 angiopoietin like 1 -5.327 TNN tenascin N -5.228 DHRS2 dehydrogenase/reductase 2 leucine rich repeat and fibronectin type III domain -5.115 LRFN2 containing 2 -5.076 FOXO6 forkhead box O6 -5.035 ETNPPL ethanolamine-phosphate phospho-lyase -4.993 MYO15A myosin XVA -4.972 IGF1 insulin like growth factor 1 -4.956 DLG2 discs large MAGUK scaffold protein 2 -4.86 SCML4 sex comb on midleg like 4 (Drosophila) Src homology 2 domain containing transforming -4.816 SHD protein D -4.764 PLP1 proteolipid protein 1 -4.764 TSPAN32 tetraspanin 32 -4.713 N4BP3 NEDD4 binding protein 3 -4.705 MYOC myocilin -4.646 CLEC3B C-type lectin domain family 3 member B -4.646 C7 complement C7 -4.62 TGM2 transglutaminase 2 -4.562 COL9A1 collagen type IX alpha 1 chain -4.55 SOSTDC1 sclerostin domain containing 1 -4.55 OGN osteoglycin -4.505 DAPL1 death associated protein like 1 -4.491 C10orf105 chromosome 10 open reading frame 105 -4.491 -
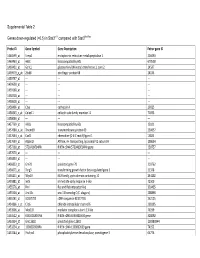
0.5) in Stat3∆/∆ Compared with Stat3flox/Flox
Supplemental Table 2 Genes down-regulated (<0.5) in Stat3∆/∆ compared with Stat3flox/flox Probe ID Gene Symbol Gene Description Entrez gene ID 1460599_at Ermp1 endoplasmic reticulum metallopeptidase 1 226090 1460463_at H60c histocompatibility 60c 670558 1460431_at Gcnt1 glucosaminyl (N-acetyl) transferase 1, core 2 14537 1459979_x_at Zfp68 zinc finger protein 68 24135 1459747_at --- --- --- 1459608_at --- --- --- 1459168_at --- --- --- 1458718_at --- --- --- 1458618_at --- --- --- 1458466_at Ctsa cathepsin A 19025 1458345_s_at Colec11 collectin sub-family member 11 71693 1458046_at --- --- --- 1457769_at H60a histocompatibility 60a 15101 1457680_a_at Tmem69 transmembrane protein 69 230657 1457644_s_at Cxcl1 chemokine (C-X-C motif) ligand 1 14825 1457639_at Atp6v1h ATPase, H+ transporting, lysosomal V1 subunit H 108664 1457260_at 5730409E04Rik RIKEN cDNA 5730409E04Rik gene 230757 1457070_at --- --- --- 1456893_at --- --- --- 1456823_at Gm70 predicted gene 70 210762 1456671_at Tbrg3 transforming growth factor beta regulated gene 3 21378 1456211_at Nlrp10 NLR family, pyrin domain containing 10 244202 1455881_at Ier5l immediate early response 5-like 72500 1455576_at Rinl Ras and Rab interactor-like 320435 1455304_at Unc13c unc-13 homolog C (C. elegans) 208898 1455241_at BC037703 cDNA sequence BC037703 242125 1454866_s_at Clic6 chloride intracellular channel 6 209195 1453906_at Med13l mediator complex subunit 13-like 76199 1453522_at 6530401N04Rik RIKEN cDNA 6530401N04 gene 328092 1453354_at Gm11602 predicted gene 11602 100380944 1453234_at -

Remarkably High Rate of DNA Amplification Promoted by the Mating-Type Switching Mechanism in Schizosaccharomyces Pombe
NOTE Remarkably High Rate of DNA Amplification Promoted by the Mating-Type Switching Mechanism in Schizosaccharomyces pombe Chuanhe Yu, Michael J. Bonaduce, and Amar J. S. Klar1 Developmental Genetics Section, Gene Regulation and Chromosome Biology Laboratory, National Institutes of Health, National Cancer Institute, Frederick, Maryland 21702-1201 ABSTRACT A novel mating-type switching-defective mutant showed a highly unstable rearrangement at the mating-type locus (mat1) in fission yeast. The mutation resulted from local amplification of a 134-bp DNA fragment by the mat1-switching phenomenon. We speculate that the rolling-circle-like replication and homologous recombination might be the general mechanisms for local genome region expansion. he fission yeast (Schizosaccharomyces pombe) cell usually and Klar 2000). The DSB is repaired by the intrachromoso- Texists in a haploid state, with P (for plus) or M (for mi- mal gene conversion event, replacing the mat1 gene with nus) cell/mating type. These cell types efficiently intercon- a copy derived from one of the mat2 or mat3 donor loci, vert by the mat1-switching phenomenon (reviewed in Egel resulting in a cell-type switch. 2005 and Klar 2007). The mating-type loci are composed of three homologous cassettes, spanning a 30-kb region Characterization of a DNA Unstable Mutant located in the middle of chromosome 2 (Figure 1A). The mating-type-determining cassette, mat1, is transcriptionally The homothallic cells, called h90 cells, efficiently switch their active and contains either P or M allele-specific DNA mating type in 40% of cell divisions (Miyata and Miyata sequence. The other two cassettes, mat2P or mat3M, exist 1981; Egel 1984; Egel and Eie 1987; Klar 1990). -

Download Download
Supplementary Figure S1. Results of flow cytometry analysis, performed to estimate CD34 positivity, after immunomagnetic separation in two different experiments. As monoclonal antibody for labeling the sample, the fluorescein isothiocyanate (FITC)- conjugated mouse anti-human CD34 MoAb (Mylteni) was used. Briefly, cell samples were incubated in the presence of the indicated MoAbs, at the proper dilution, in PBS containing 5% FCS and 1% Fc receptor (FcR) blocking reagent (Miltenyi) for 30 min at 4 C. Cells were then washed twice, resuspended with PBS and analyzed by a Coulter Epics XL (Coulter Electronics Inc., Hialeah, FL, USA) flow cytometer. only use Non-commercial 1 Supplementary Table S1. Complete list of the datasets used in this study and their sources. GEO Total samples Geo selected GEO accession of used Platform Reference series in series samples samples GSM142565 GSM142566 GSM142567 GSM142568 GSE6146 HG-U133A 14 8 - GSM142569 GSM142571 GSM142572 GSM142574 GSM51391 GSM51392 GSE2666 HG-U133A 36 4 1 GSM51393 GSM51394 only GSM321583 GSE12803 HG-U133A 20 3 GSM321584 2 GSM321585 use Promyelocytes_1 Promyelocytes_2 Promyelocytes_3 Promyelocytes_4 HG-U133A 8 8 3 GSE64282 Promyelocytes_5 Promyelocytes_6 Promyelocytes_7 Promyelocytes_8 Non-commercial 2 Supplementary Table S2. Chromosomal regions up-regulated in CD34+ samples as identified by the LAP procedure with the two-class statistics coded in the PREDA R package and an FDR threshold of 0.5. Functional enrichment analysis has been performed using DAVID (http://david.abcc.ncifcrf.gov/) -

Transcriptional Control of Tissue-Resident Memory T Cell Generation
Transcriptional control of tissue-resident memory T cell generation Filip Cvetkovski Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Graduate School of Arts and Sciences COLUMBIA UNIVERSITY 2019 © 2019 Filip Cvetkovski All rights reserved ABSTRACT Transcriptional control of tissue-resident memory T cell generation Filip Cvetkovski Tissue-resident memory T cells (TRM) are a non-circulating subset of memory that are maintained at sites of pathogen entry and mediate optimal protection against reinfection. Lung TRM can be generated in response to respiratory infection or vaccination, however, the molecular pathways involved in CD4+TRM establishment have not been defined. Here, we performed transcriptional profiling of influenza-specific lung CD4+TRM following influenza infection to identify pathways implicated in CD4+TRM generation and homeostasis. Lung CD4+TRM displayed a unique transcriptional profile distinct from spleen memory, including up-regulation of a gene network induced by the transcription factor IRF4, a known regulator of effector T cell differentiation. In addition, the gene expression profile of lung CD4+TRM was enriched in gene sets previously described in tissue-resident regulatory T cells. Up-regulation of immunomodulatory molecules such as CTLA-4, PD-1, and ICOS, suggested a potential regulatory role for CD4+TRM in tissues. Using loss-of-function genetic experiments in mice, we demonstrate that IRF4 is required for the generation of lung-localized pathogen-specific effector CD4+T cells during acute influenza infection. Influenza-specific IRF4−/− T cells failed to fully express CD44, and maintained high levels of CD62L compared to wild type, suggesting a defect in complete differentiation into lung-tropic effector T cells. -

Molecular Profiling of Peripheral Blood Is Associated with Circulating Tumor Cells Content and Poor Survival in Metastatic Castration-Resistant Prostate Cancer
www.impactjournals.com/oncotarget/ Oncotarget, Vol. 6, No. 12 Molecular profiling of peripheral blood is associated with circulating tumor cells content and poor survival in metastatic castration-resistant prostate cancer Mercedes Marín-Aguilera1, Òscar Reig1,2, Juan José Lozano3, Natalia Jiménez1, Susana García-Recio1,4, Nadina Erill5, Lydia Gaba2, Andrea Tagliapietra2, Vanesa Ortega2, Gemma Carrera6, Anna Colomer5, Pedro Gascón4 and Begoña Mellado1,2 1 Translational Genomics Group and Targeted Therapeutics in Solid Tumors Group, Institut d’Investigacions Biomèdiques August Pi i Sunyer (IDIBAPS), Barcelona, Spain 2 Medical Oncology Department, Hospital Clínic, Barcelona, Spain 3 Bioinformatics Platform Department, Centro de Investigación Biomédica en Red en el Área temática de Enfermedades Hepáticas y Digestivas (CIBEREHD), Hospital Clínic, Barcelona, Spain 4 Laboratory of Translational Oncology, Fundació Clínic per a la Recerca Biomèdica, Barcelona, Spain 5 Althia, Barcelona, Spain 6 Medical Oncology Department, Hospital Plató, Barcelona, Spain Correspondence to: Begoña Mellado, email: [email protected] Keywords: circulating tumor cells, peripheral blood, microarrays, cell search system Received: January 22, 2015 Accepted: February 14, 2015 Published: March 12, 2015 This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT The enumeration of circulating -

Detection of a 4 Bp Mutation in the 3'UTR Region of Goat Sox9 Gene
animals Article 0 Detection of a 4 bp Mutation in the 3 UTR Region of Goat Sox9 Gene and Its Effect on the Growth Traits Libang He 1,2, Yi Bi 1,2, Ruolan Wang 1,2, Chuanying Pan 1,2, Hong Chen 1,2, Xianyong Lan 1,2,* and Lei Qu 3,4,* 1 College of Animal Science and Technology, Northwest A&F University, Yangling 712100, Shaanxi, China; [email protected] 2 Shaanxi Key Laboratory of Molecular Biology for Agriculture, Yangling 712100, Shaanxi, China 3 Shaanxi Provincial Engineering and Technology Research Center of Cashmere Goats, Yulin University, Yulin 719000, Shaanxi, China 4 Life Science Research Center, Yulin University, Yulin 719000, Shaanxi, China * Correspondence: [email protected] (X.L.); [email protected] (L.Q.); Tel.: +86-137-7207-1502 (X.L.); +86-189-9226-2688 (L.Q.) Received: 4 March 2020; Accepted: 8 April 2020; Published: 13 April 2020 Simple Summary: The sex determining region Y (SRY)-type high mobility group (HMG) box 9 (Sox9) gene is critically important in the formation and development of cartilage and is considered the “main regulator” of chondrogenesis. Additionally, a large number of studies have shown that mutations in a single allele of human Sox9 can lead to campomelic dysplasia syndrome. Therefore, the mutations of Sox9 have been the subject of increasing interest among researchers. However, no studies to date have examined the association between Sox9 gene variants and growth traits in goats. Here, we detected a 4 bp indel in the 30Untranslated Regions (30UTR) region of Sox9 in Shaanbei white cashmere (SBWC) goats (n = 1109) and studied the association between this indel and growth traits. -

Redefining the Specificity of Phosphoinositide-Binding by Human
bioRxiv preprint doi: https://doi.org/10.1101/2020.06.20.163253; this version posted June 21, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. Redefining the specificity of phosphoinositide-binding by human PH domain-containing proteins Nilmani Singh1†, Adriana Reyes-Ordoñez1†, Michael A. Compagnone1, Jesus F. Moreno Castillo1, Benjamin J. Leslie2, Taekjip Ha2,3,4,5, Jie Chen1* 1Department of Cell & Developmental Biology, University of Illinois at Urbana-Champaign, Urbana, IL 61801; 2Department of Biophysics and Biophysical Chemistry, Johns Hopkins University School of Medicine, Baltimore, MD 21205; 3Department of Biophysics, Johns Hopkins University, Baltimore, MD 21218; 4Department of Biomedical Engineering, Johns Hopkins University, Baltimore, MD 21205; 5Howard Hughes Medical Institute, Baltimore, MD 21205, USA †These authors contributed equally to this work. *Correspondence: [email protected]. bioRxiv preprint doi: https://doi.org/10.1101/2020.06.20.163253; this version posted June 21, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. ABSTRACT Pleckstrin homology (PH) domains are presumed to bind phosphoinositides (PIPs), but specific interaction with and regulation by PIPs for most PH domain-containing proteins are unclear. Here we employed a single-molecule pulldown assay to study interactions of lipid vesicles with full-length proteins in mammalian whole cell lysates.