Product Description SALSA MLPA Probemix P476-A1 ZNRF3
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Diseasespecific and Inflammationindependent Stromal
Full Length Arthritis & Rheumatism DOI 10.1002/art.37704 Disease-specific and inflammation-independent stromal alterations in spondyloarthritis synovitis Nataliya Yeremenko1,2, Troy Noordenbos1,2, Tineke Cantaert1,3, Melissa van Tok1,2, Marleen van de Sande1, Juan D. Cañete4, Paul P. Tak1,5*, Dominique Baeten1,2 1Department of Clinical Immunology and Rheumatology and 2Department of Experimental Immunology, Academic Medical Center/University of Amsterdam, the Netherlands. 3Department of Immunobiology, Yale University School of Medicine, New Haven, CT, USA. 4Department of Rheumatology, Hospital Clinic de Barcelona and IDIBAPS, Spain. 5Arthrogen B.V., Amsterdam, the Netherlands. *Currently also: GlaxoSmithKline, Stevenage, U.K. Corresponding author: Dominique Baeten, MD, PhD, Department of Clinical Immunology and Rheumatology, F4-105, Academic Medical Center/University of Amsterdam, Meibergdreef 9, 1105 AZ Amsterdam, The Netherlands. E-mail: [email protected] This article has been accepted for publication and undergone full peer review but has not been through the copyediting, typesetting, pagination and proofreading process which may lead to differences between this version and the Version of Record. Please cite this article as an ‘Accepted Article’, doi: 10.1002/art.37704 © 2012 American College of Rheumatology Received: Apr 11, 2012; Revised: Jul 25, 2012; Accepted: Sep 06, 2012 Arthritis & Rheumatism Page 2 of 36 Abstract Objective: The molecular processes driving the distinct patterns of synovial inflammation and tissue remodelling in spondyloarthritis (SpA) versus rheumatoid arthritis (RA) remain largely unknown. Therefore, we aimed to identify novel and unsuspected disease- specific pathways in SpA by a systematic and unbiased synovial gene expression analysis. Methods: Differentially expressed genes were identified by pan-genomic microarray and confirmed by quantitative PCR and immunohistochemistry using synovial tissue biopsies of SpA (n=63), RA (n=28) and gout (n=9) patients. -
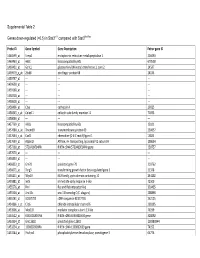
0.5) in Stat3∆/∆ Compared with Stat3flox/Flox
Supplemental Table 2 Genes down-regulated (<0.5) in Stat3∆/∆ compared with Stat3flox/flox Probe ID Gene Symbol Gene Description Entrez gene ID 1460599_at Ermp1 endoplasmic reticulum metallopeptidase 1 226090 1460463_at H60c histocompatibility 60c 670558 1460431_at Gcnt1 glucosaminyl (N-acetyl) transferase 1, core 2 14537 1459979_x_at Zfp68 zinc finger protein 68 24135 1459747_at --- --- --- 1459608_at --- --- --- 1459168_at --- --- --- 1458718_at --- --- --- 1458618_at --- --- --- 1458466_at Ctsa cathepsin A 19025 1458345_s_at Colec11 collectin sub-family member 11 71693 1458046_at --- --- --- 1457769_at H60a histocompatibility 60a 15101 1457680_a_at Tmem69 transmembrane protein 69 230657 1457644_s_at Cxcl1 chemokine (C-X-C motif) ligand 1 14825 1457639_at Atp6v1h ATPase, H+ transporting, lysosomal V1 subunit H 108664 1457260_at 5730409E04Rik RIKEN cDNA 5730409E04Rik gene 230757 1457070_at --- --- --- 1456893_at --- --- --- 1456823_at Gm70 predicted gene 70 210762 1456671_at Tbrg3 transforming growth factor beta regulated gene 3 21378 1456211_at Nlrp10 NLR family, pyrin domain containing 10 244202 1455881_at Ier5l immediate early response 5-like 72500 1455576_at Rinl Ras and Rab interactor-like 320435 1455304_at Unc13c unc-13 homolog C (C. elegans) 208898 1455241_at BC037703 cDNA sequence BC037703 242125 1454866_s_at Clic6 chloride intracellular channel 6 209195 1453906_at Med13l mediator complex subunit 13-like 76199 1453522_at 6530401N04Rik RIKEN cDNA 6530401N04 gene 328092 1453354_at Gm11602 predicted gene 11602 100380944 1453234_at -

Long, Noncoding RNA Dysregulation in Glioblastoma
cancers Review Long, Noncoding RNA Dysregulation in Glioblastoma Patrick A. DeSouza 1,2 , Xuan Qu 1, Hao Chen 1,3, Bhuvic Patel 1 , Christopher A. Maher 2,4,5,6 and Albert H. Kim 1,6,* 1 Department of Neurological Surgery, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA; [email protected] (P.A.D.); [email protected] (X.Q.); [email protected] (H.C.); [email protected] (B.P.) 2 Department of Internal Medicine, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA; [email protected] 3 Department of Neuroscience, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA 4 Department of Biomedical Engineering, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA 5 McDonnell Genome Institute, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA 6 Siteman Cancer Center, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA * Correspondence: [email protected] Simple Summary: Developing effective therapies for glioblastoma (GBM), the most common primary brain cancer, remains challenging due to the heterogeneity within tumors and therapeutic resistance that drives recurrence. Noncoding RNAs are transcribed from a large proportion of the genome and remain largely unexplored in their contribution to the evolution of GBM tumors. Here, we will review the general mechanisms of long, noncoding RNAs and the current knowledge of how these impact heterogeneity and therapeutic resistance in GBM. A better understanding of the molecular drivers required for these aggressive tumors is necessary to improve the management and outcomes Citation: DeSouza, P.A.; Qu, X.; of this challenging disease. -

Identification of Active Signaling Pathways by Integrating Gene Expression and Protein Interaction Data Md Humayun Kabir1,2,3, Ralph Patrick2,4,5, Joshua W
Kabir et al. BMC Systems Biology 2018, 12(Suppl 9):120 https://doi.org/10.1186/s12918-018-0655-x RESEARCH Open Access Identification of active signaling pathways by integrating gene expression and protein interaction data Md Humayun Kabir1,2,3, Ralph Patrick2,4,5, Joshua W. K. Ho2,4,6* and Michael D. O’Connor1,7* From 29th International Conference on Genome Informatics Yunnan, China. 3-5 December 2018 Abstract Background: Signaling pathways are the key biological mechanisms that transduce extracellular signals to affect transcription factor mediated gene regulation within cells. A number of computational methods have been developed to identify the topological structure of a specific signaling pathway using protein-protein interaction data, but they are not designed for identifying active signaling pathways in an unbiased manner. On the other hand, there are statistical methods based on gene sets or pathway data that can prioritize likely active signaling pathways, but they do not make full use of active pathway structure that link receptor, kinases and downstream transcription factors. Results: Here, we present a method to simultaneously predict the set of active signaling pathways, together with their pathway structure, by integrating protein-protein interaction network and gene expression data. We evaluated the capacity for our method to predict active signaling pathways for dental epithelial cells, ocular lens epithelial cells, human pluripotent stem cell-derived lens epithelial cells, and lens fiber cells. This analysis showed our approach could identify all the known active pathways that are associated with tooth formation and lens development. Conclusions: The results suggest that SPAGI can be a useful approach to identify the potential active signaling pathways given a gene expression profile. -

Kremen Is Beyond a Subsidiary Co-Receptor of Wnt Signaling: an in Silico Validation
Turkish Journal of Biology Turk J Biol (2015) 39: 501-510 http://journals.tubitak.gov.tr/biology/ © TÜBİTAK Research Article doi:10.3906/biy-1409-1 Kremen is beyond a subsidiary co-receptor of Wnt signaling: an in silico validation 1 2, 3 Hemn MOHAMMADPOUR , Saeed KHALILI *, Zahra Sadat HASHEMI 1 Department of Medical Immunology, Faculty of Medical Science, Tarbiat Modares University, Tehran, Iran 2 Department of Medical Biotechnology, Faculty of Medical Science, Tarbiat Modares University, Tehran, Iran 3 Department of Medical Biotechnology, School of Advanced Medical Technologies, Tehran University of Medical Science, Tehran, Iran Received: 01.09.2014 Accepted/Published Online: 12.02.2015 Printed: 15.06.2015 Abstract: Kremen-1 is a co-receptor of Wnt signaling, interacting with the Dkk-3 protein, which is an antagonist of the Wnt/β catenin pathway. In the present study we attempted to shed some light on the possible orientations of Kremen/Dkk-3 interactions, explaining the mechanisms of Kremen and Dkk-3 functions. Employing state-of-the-art software, a Kremen model was built and subsequently refined. The quality of the final model was evaluated using RMSD calculations. Ultimately, we used the Kremen model for docking analysis between Kremen and Dkk-3 molecules. A model built by the Robetta server showed the best quality scores. Near native coordination of the final model was verified getting < 2 Å RMSD values between our model and an experimentally resolved structure. Docking analysis indicates that one low energy orientation for Kremen/Dkk-3 involves all extracellular domains of Kremen, while another only involves the CUB and WSC domains. -

Meta-Analysis Identifies 13 New Loci Associated with Waist-Hip Ratio And
ARTICLES Meta-analysis identifies 13 new loci associated with waist-hip ratio and reveals sexual dimorphism in the genetic basis of fat distribution Waist-hip ratio (WHR) is a measure of body fat distribution and a predictor of metabolic consequences independent of overall adiposity. WHR is heritable, but few genetic variants influencing this trait have been identified. We conducted a meta-analysis of 32 genome-wide association studies for WHR adjusted for body mass index (comprising up to 77,167 participants), following up 16 loci in an additional 29 studies (comprising up to 113,636 subjects). We identified 13 new loci in or near RSPO3, VEGFA, TBX15-WARS2, NFE2L3, GRB14, DNM3-PIGC, ITPR2-SSPN, LY86, HOXC13, ADAMTS9, ZNRF3-KREMEN1, NISCH-STAB1 and CPEB4 (P = 1.9 × 10−9 to P = 1.8 × 10−40) and the known signal at LYPLAL1. Seven of these loci exhibited marked sexual dimorphism, all with a stronger effect on WHR in women than men (P for sex difference = 1.9 × 10−3 to P = 1.2 × 10−13). These findings provide evidence for multiple loci that modulate body fat distribution independent of overall adiposity and reveal strong gene-by-sex interactions. Central obesity and body fat distribution, as measured by waist discovery stage, up to 2,850,269 imputed and genotyped SNPs circumference and WHR, are associated with individual risk of type were examined in 32 GWAS comprising up to 77,167 participants 2 diabetes (T2D)1,2 and coronary heart disease3 and with mortality informative for anthropometric measures of body fat distribution. from all causes4. -

Anti-KREMEN1 Antibody (ARG40158)
Product datasheet [email protected] ARG40158 Package: 50 μg anti-KREMEN1 antibody Store at: -20°C Summary Product Description Rabbit Polyclonal antibody recognizes KREMEN1 Tested Reactivity Hu, Rat Tested Application IHC-P, WB Specificity Three isoforms of KREMEN1 exists as a result of alternative splicing event. Host Rabbit Clonality Polyclonal Isotype IgG Target Name KREMEN1 Antigen Species Human Immunogen A 18 amino acid peptide within the last 50 amino acids of Human KREMEN1. Conjugation Un-conjugated Alternate Names KREMEN; KRM1; Dickkopf receptor; Kringle domain-containing transmembrane protein 1; Kremen protein 1; Kringle-containing protein marking the eye and the nose Application Instructions Application table Application Dilution IHC-P 5 - 20 µg/ml WB 0.125 - 0.25 µg/ml Application Note * The dilutions indicate recommended starting dilutions and the optimal dilutions or concentrations should be determined by the scientist. Calculated Mw 52 kDa Properties Form Liquid Purification Affinity purification with immunogen. Buffer PBS and 0.02% Sodium azide. Preservative 0.02% Sodium azide Concentration 1 mg/ml Storage instruction For continuous use, store undiluted antibody at 2-8°C for up to a week. For long-term storage, aliquot and store at -20°C or below. Storage in frost free freezers is not recommended. Avoid repeated freeze/thaw cycles. Suggest spin the vial prior to opening. The antibody solution should be gently mixed before use. www.arigobio.com 1/3 Note For laboratory research only, not for drug, diagnostic or other use. Bioinformation Gene Symbol KREMEN1 Gene Full Name kringle containing transmembrane protein 1 Background This gene encodes a high-affinity dickkopf homolog 1 (DKK1) transmembrane receptor that functionally cooperates with DKK1 to block wingless (WNT)/beta-catenin signaling. -

The Regulation of Self-Renewal in Normal Human Urothelial Cells
The Regulation of Self-Renewal in Normal Human Urothelial Cells Lisa A. Kirkwood PhD University of York Department of Biology April 2012 Abstract The urinary tract is lined by a mitotically-quiescent, but highly regenerative epithelium, the urothelium. The mechanisms regulating urothelial regeneration are incompletely understood although autocrine stimulation of the Epidermal Growth Factor Receptor (EGFR) signalling pathway has been implicated. The hypothesis developed in this thesis is that urothelial homeostasis is regulated through resolution of interactive signal transduction networks downstream of local environmental cues, such as cell:cell contact. Here, canonical Wnt signalling was examined as a candidate key pathway due to the pivotal role of β-catenin in both nuclear transcription and intercellular adherens junctions. Normal human urothelial (NHU) cells isolated from surgical biopsies were grown as finite cell lines in monolayer culture. mRNA analysis from proliferating cultures inferred all components for a functional autocrine-activated canonical Wnt cascade were present. In proliferating cells, β-catenin was nuclear and Axin2 expression provided an objective hallmark of β-catenin/TCF transcription factor activity. This endogenous activity was not mediated by Wnt receptor activation, as Wnt ligand was produced in inactive (non-palmitylated) form in serum-free culture, but instead -catenin activation was driven via EGFR- mediated phosphorylation of GSK3 and inhibition of the β-catenin destruction complex. In quiescent, contact–inhibited cultures, β-catenin was seen to re- localise to the adherens junctions and GSK3β activity was re-established. Knock-down of β-catenin using RNA interference led to significant changes in p-ERK and p-AKT activity as well as an increase in E-cadherin protein expression. -

WISP1 Regulation by Micrornas in Pulmonary Fibrosis“
Aus dem Institut für Prophylaxe und Epidemiologie der Kreislaufkrankheiten der Ludwig-Maximilians-Universität München Direktor: Prof. Dr. med. Christian Weber WISP1 Regulation by MicroRNAs in Pulmonary Fibrosis Dissertation zum Erwerb des Doktorgrades der Naturwissenschaften an der Medizinischen Fakultät der Ludwig-Maximilians-Universität München vorgelegt von Barbara Berschneider aus Regensburg 2014 Meiner Familie Gedruckt mit Genehmigung der Medizinischen Fakultät der Ludwig-Maximilians-Universität München Betreuer: PD Dr. rer. nat. Peter Neth Zweitgutachter: Prof. Dr. rer.nat. Wolfgang Zimmermann Dekan: Prof. Dr.med. Dr.h.c. Maximilian Reiser, FACR, FRCR Tag der mündlichen Prüfung: 6. November 2014 TABLE OF CONTENTS Table of Contents ........................................................................................................... I Abbreviations .............................................................................................................. VI 1 Zusammenfassung .................................................................................................. 1 2 Summary .................................................................................................................... 2 3 Introduction ............................................................................................................. 3 3.1 Idiopathic pulmonary fibrosis ............................................................................................................... 3 3.1.1 Clinical features of IPF ................................................................................................................... -

Altered Gene Expression in Circulating Immune Cells Following a 24-Hour Passive Dehydration
University of Connecticut OpenCommons@UConn Master's Theses University of Connecticut Graduate School 5-10-2020 Altered Gene Expression in Circulating Immune Cells Following a 24-Hour Passive Dehydration Aidan Fiol [email protected] Follow this and additional works at: https://opencommons.uconn.edu/gs_theses Recommended Citation Fiol, Aidan, "Altered Gene Expression in Circulating Immune Cells Following a 24-Hour Passive Dehydration" (2020). Master's Theses. 1480. https://opencommons.uconn.edu/gs_theses/1480 This work is brought to you for free and open access by the University of Connecticut Graduate School at OpenCommons@UConn. It has been accepted for inclusion in Master's Theses by an authorized administrator of OpenCommons@UConn. For more information, please contact [email protected]. Altered Gene Expression in Circulating Immune Cells Following a 24- Hour Passive Dehydration Aidan Fiol B.S., University of Connecticut, 2018 A Thesis Submitted in Partial Fulfillment of the Requirements for the Degree of Master of Science At the University of Connecticut 2020 i copyright by Aidan Fiol 2020 ii APPROVAL PAGE Master of Science Thesis Altered Gene Expression in Circulating Immune Cells Following a 24- Hour Passive Dehydration Presented by Aidan Fiol, B.S. Major Advisor__________________________________________________________________ Elaine Choung-Hee Lee, Ph.D. Associate Advisor_______________________________________________________________ Douglas J. Casa, Ph.D. Associate Advisor_______________________________________________________________ Robert A. Huggins, Ph.D. University of Connecticut 2020 iii ACKNOWLEDGEMENTS I’d like to thank my committee members. Dr. Lee, you have been an amazing advisor, mentor and friend to me these past few years. Your advice, whether it was how to be a better writer or scientist, or just general life advice given on our way to get coffee, has helped me grow as a researcher and as a person. -

DKK1 Antibody (N-Terminus) Rabbit Polyclonal Antibody Catalog # ALS10601
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 DKK1 Antibody (N-Terminus) Rabbit Polyclonal Antibody Catalog # ALS10601 Specification DKK1 Antibody (N-Terminus) - Product Information Application IHC Primary Accession O94907 Reactivity Human, Monkey, Pig, Horse Host Rabbit Clonality Polyclonal Calculated MW 29kDa KDa DKK1 Antibody (N-Terminus) - Additional Information Gene ID 22943 Anti-DKK1 antibody ALS10601 IHC of human uterus, endometrium. Other Names Dickkopf-related protein 1, Dickkopf-1, Dkk-1, hDkk-1, SK, DKK1 DKK1 Antibody (N-Terminus) - Background Target/Specificity Antagonizes canonical Wnt signaling by Human DKK1. BLAST analysis of the peptide inhibiting LRP5/6 interaction with Wnt and by immunogen showed no homology with forming a ternary complex with the other human proteins. transmembrane protein KREMEN that promotes internalization of LRP5/6. DKKs play Reconstitution & Storage an important role in vertebrate development, Long term: -70°C; Short term: +4°C where they locally inhibit Wnt regulated processes such as antero- posterior axial Precautions patterning, limb development, somitogenesis DKK1 Antibody (N-Terminus) is for research and eye formation. In the adult, Dkks are use only and not for use in diagnostic or implicated in bone formation and bone therapeutic procedures. disease, cancer and Alzheimer disease. DKK1 Antibody (N-Terminus) - References DKK1 Antibody (N-Terminus) - Protein Information Fedi P.,et al.J. Biol. Chem. 274:19465-19472(1999). Name DKK1 Krupnik V.E.,et al.Gene 238:301-313(1999). Tate G.,et al.Submitted (NOV-1998) to the Function EMBL/GenBank/DDBJ databases. Antagonizes canonical Wnt signaling by Roessler E.,et al.Cytogenet. -

UC San Diego Electronic Theses and Dissertations
UC San Diego UC San Diego Electronic Theses and Dissertations Title Cardiac Stretch-Induced Transcriptomic Changes are Axis-Dependent Permalink https://escholarship.org/uc/item/7m04f0b0 Author Buchholz, Kyle Stephen Publication Date 2016 Peer reviewed|Thesis/dissertation eScholarship.org Powered by the California Digital Library University of California UNIVERSITY OF CALIFORNIA, SAN DIEGO Cardiac Stretch-Induced Transcriptomic Changes are Axis-Dependent A dissertation submitted in partial satisfaction of the requirements for the degree Doctor of Philosophy in Bioengineering by Kyle Stephen Buchholz Committee in Charge: Professor Jeffrey Omens, Chair Professor Andrew McCulloch, Co-Chair Professor Ju Chen Professor Karen Christman Professor Robert Ross Professor Alexander Zambon 2016 Copyright Kyle Stephen Buchholz, 2016 All rights reserved Signature Page The Dissertation of Kyle Stephen Buchholz is approved and it is acceptable in quality and form for publication on microfilm and electronically: Co-Chair Chair University of California, San Diego 2016 iii Dedication To my beautiful wife, Rhia. iv Table of Contents Signature Page ................................................................................................................... iii Dedication .......................................................................................................................... iv Table of Contents ................................................................................................................ v List of Figures ...................................................................................................................