Meta-Analysis Identifies 13 New Loci Associated with Waist-Hip Ratio And
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genome-Wide Association Studies Identify Genetic Loci Associated With
SUPPLEMENTARY DATA Genome-wide Association Studies Identify Genetic Loci Associated with Albuminuria in Diabetes SUPPLEMENTAL MATERIALS This work is dedicated to the memory of our colleague Dr. Wen Hong Linda Kao, a wonderful person, brilliant scientist and central member of the CKDGen Consortium. 1 ©2016 American Diabetes Association. Published online at http://diabetes.diabetesjournals.org/lookup/suppl/doi:10.2337/db15-1313/-/DC1 SUPPLEMENTARY DATA Table of Contents SUPPLEMENTARY FIGURE 1: QQ PLOTS FOR ALL GWAS META-ANALYSES ............................................. 3 SUPPLEMENTARY FIGURE 2: MANHATTAN PLOTS FOR ALL GWAS META-ANALYSES ............................. 4 SUPPLEMENTARY FIGURE 3: REGIONAL ASSOCIATION PLOTS............................................................... 6 SUPPLEMENTARY FIGURE 4: EVALUATION OF GLOMERULOSCLEROSIS IN RAB38 KO, CONGENIC AND TRANSGENIC RATS. ........................................................................................................................... 17 SUPPLEMENTARY TABLE 1: CHARACTERISTICS OF THE STUDY POPULATIONS ..................................... 18 SUPPLEMENTARY TABLE 2: INFORMATION ABOUT STUDY DESIGN AND UACR MEASUREMENT .......... 20 SUPPLEMENTARY TABLE 3: STUDY-SPECIFIC INFORMATION ABOUT GENOTYPING, IMPUTATION AND DATA MANAGEMENT AND ANALYSIS ................................................................................................ 31 SUPPLEMENTARY TABLE 4: SNPS ASSOCIATED WITH UACR AMONG ALL INDIVIDUALS WITH A P-VALUE OF <1E-05. ....................................................................................................................................... -

Diseasespecific and Inflammationindependent Stromal
Full Length Arthritis & Rheumatism DOI 10.1002/art.37704 Disease-specific and inflammation-independent stromal alterations in spondyloarthritis synovitis Nataliya Yeremenko1,2, Troy Noordenbos1,2, Tineke Cantaert1,3, Melissa van Tok1,2, Marleen van de Sande1, Juan D. Cañete4, Paul P. Tak1,5*, Dominique Baeten1,2 1Department of Clinical Immunology and Rheumatology and 2Department of Experimental Immunology, Academic Medical Center/University of Amsterdam, the Netherlands. 3Department of Immunobiology, Yale University School of Medicine, New Haven, CT, USA. 4Department of Rheumatology, Hospital Clinic de Barcelona and IDIBAPS, Spain. 5Arthrogen B.V., Amsterdam, the Netherlands. *Currently also: GlaxoSmithKline, Stevenage, U.K. Corresponding author: Dominique Baeten, MD, PhD, Department of Clinical Immunology and Rheumatology, F4-105, Academic Medical Center/University of Amsterdam, Meibergdreef 9, 1105 AZ Amsterdam, The Netherlands. E-mail: [email protected] This article has been accepted for publication and undergone full peer review but has not been through the copyediting, typesetting, pagination and proofreading process which may lead to differences between this version and the Version of Record. Please cite this article as an ‘Accepted Article’, doi: 10.1002/art.37704 © 2012 American College of Rheumatology Received: Apr 11, 2012; Revised: Jul 25, 2012; Accepted: Sep 06, 2012 Arthritis & Rheumatism Page 2 of 36 Abstract Objective: The molecular processes driving the distinct patterns of synovial inflammation and tissue remodelling in spondyloarthritis (SpA) versus rheumatoid arthritis (RA) remain largely unknown. Therefore, we aimed to identify novel and unsuspected disease- specific pathways in SpA by a systematic and unbiased synovial gene expression analysis. Methods: Differentially expressed genes were identified by pan-genomic microarray and confirmed by quantitative PCR and immunohistochemistry using synovial tissue biopsies of SpA (n=63), RA (n=28) and gout (n=9) patients. -

A Context-Dependent Role for the RNF146 Ubiquitin Ligase in Wingless/Wnt Signaling in Drosophila
| INVESTIGATION A Context-Dependent Role for the RNF146 Ubiquitin Ligase in Wingless/Wnt Signaling in Drosophila Zhenghan Wang,* Ofelia Tacchelly-Benites,* Geoffrey P. Noble,* Megan K. Johnson,* Jean-Philippe Gagné,† Guy G. Poirier,† and Yashi Ahmed*,1 *Department of Molecular and Systems Biology, Norris Cotton Cancer Center, Geisel School of Medicine at Dartmouth College HB7400, Hanover, New Hampshire 03755 and †Centre de Recherche du Centre Hospitalier Universitaire de Québec-Université Laval, CHUL Pavilion, Axe Oncologie, Québec G1V 4G2, Canada ABSTRACT Aberrant activation of the Wnt signal transduction pathway triggers the development of colorectal cancer. The ADP-ribose polymerase Tankyrase (TNKS) mediates proteolysis of Axin—a negative regulator of Wnt signaling—and provides a promising ther- apeutic target for Wnt-driven diseases. Proteolysis of TNKS substrates is mediated through their ubiquitination by the poly-ADP-ribose (pADPr)-dependent RING-domain E3 ubiquitin ligase RNF146/Iduna. Like TNKS, RNF146 promotes Axin proteolysis and Wnt pathway activation in some cultured cell lines, but in contrast with TNKS, RNF146 is dispensable for Axin degradation in colorectal carcinoma cells. Thus, the contexts in which RNF146 is essential for TNKS-mediated Axin destabilization and Wnt signaling remain uncertain. Herein, we tested the requirement for RNF146 in TNKS-mediated Axin proteolysis and Wnt pathway activation in a range of in vivo settings. Using null mutants in Drosophila, we provide genetic and biochemical evidence that Rnf146 and Tnks function in the same proteolysis pathway in vivo. Furthermore, like Tnks, Drosophila Rnf146 promotes Wingless signaling in multiple developmental contexts by buffering Axin levels to ensure they remain below the threshold at which Wingless signaling is inhibited. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -
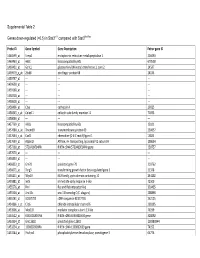
0.5) in Stat3∆/∆ Compared with Stat3flox/Flox
Supplemental Table 2 Genes down-regulated (<0.5) in Stat3∆/∆ compared with Stat3flox/flox Probe ID Gene Symbol Gene Description Entrez gene ID 1460599_at Ermp1 endoplasmic reticulum metallopeptidase 1 226090 1460463_at H60c histocompatibility 60c 670558 1460431_at Gcnt1 glucosaminyl (N-acetyl) transferase 1, core 2 14537 1459979_x_at Zfp68 zinc finger protein 68 24135 1459747_at --- --- --- 1459608_at --- --- --- 1459168_at --- --- --- 1458718_at --- --- --- 1458618_at --- --- --- 1458466_at Ctsa cathepsin A 19025 1458345_s_at Colec11 collectin sub-family member 11 71693 1458046_at --- --- --- 1457769_at H60a histocompatibility 60a 15101 1457680_a_at Tmem69 transmembrane protein 69 230657 1457644_s_at Cxcl1 chemokine (C-X-C motif) ligand 1 14825 1457639_at Atp6v1h ATPase, H+ transporting, lysosomal V1 subunit H 108664 1457260_at 5730409E04Rik RIKEN cDNA 5730409E04Rik gene 230757 1457070_at --- --- --- 1456893_at --- --- --- 1456823_at Gm70 predicted gene 70 210762 1456671_at Tbrg3 transforming growth factor beta regulated gene 3 21378 1456211_at Nlrp10 NLR family, pyrin domain containing 10 244202 1455881_at Ier5l immediate early response 5-like 72500 1455576_at Rinl Ras and Rab interactor-like 320435 1455304_at Unc13c unc-13 homolog C (C. elegans) 208898 1455241_at BC037703 cDNA sequence BC037703 242125 1454866_s_at Clic6 chloride intracellular channel 6 209195 1453906_at Med13l mediator complex subunit 13-like 76199 1453522_at 6530401N04Rik RIKEN cDNA 6530401N04 gene 328092 1453354_at Gm11602 predicted gene 11602 100380944 1453234_at -

Bioinformatic Analysis Suggests That UGT2B15 Activates the Hippo‑YAP Signaling Pathway Leading to the Pathogenesis of Gastric Cancer
ONCOLOGY REPORTS 40: 1855-1862, 2018 Bioinformatic analysis suggests that UGT2B15 activates the Hippo‑YAP signaling pathway leading to the pathogenesis of gastric cancer XUANMIN CHEN1*, DEFENG LI4,5*, NANNAN WANG1*, MEIFENG YANG2*, AIJUN LIAO1*, SHULING WANG3*, GUANGSHENG HU1, BING ZENG1, YUHONG YAO1, DIQUN LIU1, HAN LIU1, WEIWEI ZHOU1, WEISHENG XIAO1, PEIYUAN LI1, CHEN MING1, SONG PING2, PINGFANG CHEN1, LI JING1, YU BAI3 and JUN YAO4 Departments of 1Gastroenterology and 2Hematology, The First Affiliated Hospital of University of South China, University of South China, Hengyang, Hunan 421001; 3Department of Gastroenterology, Changhai Hospital, Second Military Medical University, Shanghai 200433; 4Department of Gastroenterology, The 2nd Clinical Μedicine College (Shenzhen People's Hospital) of Jinan University, Shenzhen, Guangdong 518020; 5Integrated Chinese and Western Medicine Postdoctoral Research Station, Jinan University, Guangzhou, Guangdong 510632, P.R. China Received November 30, 2017; Accepted May 25, 2018 DOI: 10.3892/or.2018.6604 Abstract. Gastric cancer (GC) is one of the most common development of GC. Taken together, our findings demonstrate malignancies that threatens human health. As the molecular that UGT2B15 may be an oncogene in GC and is a promising mechanisms unerlying GC are not completely understood, therapeutic target for cancer treatment. identification of genes related to GC could provide new insights into gene function as well as potential treatment targets. We Introduction discovered that UGT2B15 may contribute to the pathogenesis and progression of GC using GEO data and bioinformatic Gastric cancer (GC) is a type of malignant digestive tract analysis. Using TCGA data, UGT2B15 mRNA was found to be tumor with a poor prognosis. GLOBOCAN 2015 reported GC significantly overexpressed in GC tissues; patients with higher as the third leading cause of cancer-related mortality in the UGT2B15 had a poorer prognosis. -

Long, Noncoding RNA Dysregulation in Glioblastoma
cancers Review Long, Noncoding RNA Dysregulation in Glioblastoma Patrick A. DeSouza 1,2 , Xuan Qu 1, Hao Chen 1,3, Bhuvic Patel 1 , Christopher A. Maher 2,4,5,6 and Albert H. Kim 1,6,* 1 Department of Neurological Surgery, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA; [email protected] (P.A.D.); [email protected] (X.Q.); [email protected] (H.C.); [email protected] (B.P.) 2 Department of Internal Medicine, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA; [email protected] 3 Department of Neuroscience, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA 4 Department of Biomedical Engineering, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA 5 McDonnell Genome Institute, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA 6 Siteman Cancer Center, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA * Correspondence: [email protected] Simple Summary: Developing effective therapies for glioblastoma (GBM), the most common primary brain cancer, remains challenging due to the heterogeneity within tumors and therapeutic resistance that drives recurrence. Noncoding RNAs are transcribed from a large proportion of the genome and remain largely unexplored in their contribution to the evolution of GBM tumors. Here, we will review the general mechanisms of long, noncoding RNAs and the current knowledge of how these impact heterogeneity and therapeutic resistance in GBM. A better understanding of the molecular drivers required for these aggressive tumors is necessary to improve the management and outcomes Citation: DeSouza, P.A.; Qu, X.; of this challenging disease. -

Cellular and Molecular Signatures in the Disease Tissue of Early
Cellular and Molecular Signatures in the Disease Tissue of Early Rheumatoid Arthritis Stratify Clinical Response to csDMARD-Therapy and Predict Radiographic Progression Frances Humby1,* Myles Lewis1,* Nandhini Ramamoorthi2, Jason Hackney3, Michael Barnes1, Michele Bombardieri1, Francesca Setiadi2, Stephen Kelly1, Fabiola Bene1, Maria di Cicco1, Sudeh Riahi1, Vidalba Rocher-Ros1, Nora Ng1, Ilias Lazorou1, Rebecca E. Hands1, Desiree van der Heijde4, Robert Landewé5, Annette van der Helm-van Mil4, Alberto Cauli6, Iain B. McInnes7, Christopher D. Buckley8, Ernest Choy9, Peter Taylor10, Michael J. Townsend2 & Costantino Pitzalis1 1Centre for Experimental Medicine and Rheumatology, William Harvey Research Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, Charterhouse Square, London EC1M 6BQ, UK. Departments of 2Biomarker Discovery OMNI, 3Bioinformatics and Computational Biology, Genentech Research and Early Development, South San Francisco, California 94080 USA 4Department of Rheumatology, Leiden University Medical Center, The Netherlands 5Department of Clinical Immunology & Rheumatology, Amsterdam Rheumatology & Immunology Center, Amsterdam, The Netherlands 6Rheumatology Unit, Department of Medical Sciences, Policlinico of the University of Cagliari, Cagliari, Italy 7Institute of Infection, Immunity and Inflammation, University of Glasgow, Glasgow G12 8TA, UK 8Rheumatology Research Group, Institute of Inflammation and Ageing (IIA), University of Birmingham, Birmingham B15 2WB, UK 9Institute of -

RING-Type E3 Ligases: Master Manipulators of E2 Ubiquitin-Conjugating Enzymes and Ubiquitination☆
Biochimica et Biophysica Acta 1843 (2014) 47–60 Contents lists available at ScienceDirect Biochimica et Biophysica Acta journal homepage: www.elsevier.com/locate/bbamcr Review RING-type E3 ligases: Master manipulators of E2 ubiquitin-conjugating enzymes and ubiquitination☆ Meredith B. Metzger a,1, Jonathan N. Pruneda b,1, Rachel E. Klevit b,⁎, Allan M. Weissman a,⁎⁎ a Laboratory of Protein Dynamics and Signaling, Center for Cancer Research, National Cancer Institute, 1050 Boyles Street, Frederick, MD 21702, USA b Department of Biochemistry, Box 357350, University of Washington, Seattle, WA 98195, USA article info abstract Article history: RING finger domain and RING finger-like ubiquitin ligases (E3s), such as U-box proteins, constitute the vast Received 5 March 2013 majority of known E3s. RING-type E3s function together with ubiquitin-conjugating enzymes (E2s) to medi- Received in revised form 23 May 2013 ate ubiquitination and are implicated in numerous cellular processes. In part because of their importance in Accepted 29 May 2013 human physiology and disease, these proteins and their cellular functions represent an intense area of study. Available online 6 June 2013 Here we review recent advances in RING-type E3 recognition of substrates, their cellular regulation, and their varied architecture. Additionally, recent structural insights into RING-type E3 function, with a focus on im- Keywords: RING finger portant interactions with E2s and ubiquitin, are reviewed. This article is part of a Special Issue entitled: U-box Ubiquitin–Proteasome System. Guest Editors: Thomas Sommer and Dieter H. Wolf. Ubiquitin ligase (E3) Published by Elsevier B.V. Ubiquitin-conjugating enzyme (E2) Protein degradation Catalysis 1. -

NICU Gene List Generator.Xlsx
Neonatal Crisis Sequencing Panel Gene List Genes: A2ML1 - B3GLCT A2ML1 ADAMTS9 ALG1 ARHGEF15 AAAS ADAMTSL2 ALG11 ARHGEF9 AARS1 ADAR ALG12 ARID1A AARS2 ADARB1 ALG13 ARID1B ABAT ADCY6 ALG14 ARID2 ABCA12 ADD3 ALG2 ARL13B ABCA3 ADGRG1 ALG3 ARL6 ABCA4 ADGRV1 ALG6 ARMC9 ABCB11 ADK ALG8 ARPC1B ABCB4 ADNP ALG9 ARSA ABCC6 ADPRS ALK ARSL ABCC8 ADSL ALMS1 ARX ABCC9 AEBP1 ALOX12B ASAH1 ABCD1 AFF3 ALOXE3 ASCC1 ABCD3 AFF4 ALPK3 ASH1L ABCD4 AFG3L2 ALPL ASL ABHD5 AGA ALS2 ASNS ACAD8 AGK ALX3 ASPA ACAD9 AGL ALX4 ASPM ACADM AGPS AMELX ASS1 ACADS AGRN AMER1 ASXL1 ACADSB AGT AMH ASXL3 ACADVL AGTPBP1 AMHR2 ATAD1 ACAN AGTR1 AMN ATL1 ACAT1 AGXT AMPD2 ATM ACE AHCY AMT ATP1A1 ACO2 AHDC1 ANK1 ATP1A2 ACOX1 AHI1 ANK2 ATP1A3 ACP5 AIFM1 ANKH ATP2A1 ACSF3 AIMP1 ANKLE2 ATP5F1A ACTA1 AIMP2 ANKRD11 ATP5F1D ACTA2 AIRE ANKRD26 ATP5F1E ACTB AKAP9 ANTXR2 ATP6V0A2 ACTC1 AKR1D1 AP1S2 ATP6V1B1 ACTG1 AKT2 AP2S1 ATP7A ACTG2 AKT3 AP3B1 ATP8A2 ACTL6B ALAS2 AP3B2 ATP8B1 ACTN1 ALB AP4B1 ATPAF2 ACTN2 ALDH18A1 AP4M1 ATR ACTN4 ALDH1A3 AP4S1 ATRX ACVR1 ALDH3A2 APC AUH ACVRL1 ALDH4A1 APTX AVPR2 ACY1 ALDH5A1 AR B3GALNT2 ADA ALDH6A1 ARFGEF2 B3GALT6 ADAMTS13 ALDH7A1 ARG1 B3GAT3 ADAMTS2 ALDOB ARHGAP31 B3GLCT Updated: 03/15/2021; v.3.6 1 Neonatal Crisis Sequencing Panel Gene List Genes: B4GALT1 - COL11A2 B4GALT1 C1QBP CD3G CHKB B4GALT7 C3 CD40LG CHMP1A B4GAT1 CA2 CD59 CHRNA1 B9D1 CA5A CD70 CHRNB1 B9D2 CACNA1A CD96 CHRND BAAT CACNA1C CDAN1 CHRNE BBIP1 CACNA1D CDC42 CHRNG BBS1 CACNA1E CDH1 CHST14 BBS10 CACNA1F CDH2 CHST3 BBS12 CACNA1G CDK10 CHUK BBS2 CACNA2D2 CDK13 CILK1 BBS4 CACNB2 CDK5RAP2 -

Product Description SALSA MLPA Probemix P476-A1 ZNRF3
MRC-Holland ® Product Description version A1-02; Issued 25 September 2018 MLPA Product Description SALSA ® MLPA ® Probemix P476-A1 ZNRF3 To be used with the MLPA General Protocol. Version A1. New Product. Catalogue numbers: • P476-025R: SALSA MLPA Probemix P476 ZNRF3, 25 reactions. • P476-050R: SALSA MLPA Probemix P476 ZNRF3, 50 reactions. • P476-100R: SALSA MLPA Probemix P476 ZNRF3, 100 reactions. To be used in combination with a SALSA MLPA reagent kit, available for various number of reactions. MLPA reagent kits are either provided with FAM or Cy5.0 dye-labelled PCR primer, suitable for Applied Biosystems and Beckman capillary sequencers, respectively (see www.mlpa.com ). Certificate of Analysis: Information regarding storage conditions, quality tests, and a sample electropherogram from the current sales lot is available at www.mlpa.com . Precautions and warnings: For professional use only. Always consult the most recent product description AND the MLPA General Protocol before use: www.mlpa.com . It is the responsibility of the user to be aware of the latest scientific knowledge of the application before drawing any conclusions from findings generated with this product. General information: The SALSA MLPA Probemix P476 ZNRF3 is a research use only (RUO) assay for the detection of deletions and duplications in the ZNRF3 gene. ZNRF3 is an E3 ubiquitin-protein ligase that acts as a negative feedback regulator of Wnt signalling (Hao et al. 2012). Three independent studies show homozygous deletions of the ZNRF3 gene in 10 to 16% of adrenocortical carcinoma cases (Assié et al. 2014; Juhlin et al. 2015; Zheng et al. 2016). Moreover, in 51% of microsatellite stable colorectal cancers deletion events at the ZNRF3 locus are detected (Bond et al. -

Noelia Díaz Blanco
Effects of environmental factors on the gonadal transcriptome of European sea bass (Dicentrarchus labrax), juvenile growth and sex ratios Noelia Díaz Blanco Ph.D. thesis 2014 Submitted in partial fulfillment of the requirements for the Ph.D. degree from the Universitat Pompeu Fabra (UPF). This work has been carried out at the Group of Biology of Reproduction (GBR), at the Department of Renewable Marine Resources of the Institute of Marine Sciences (ICM-CSIC). Thesis supervisor: Dr. Francesc Piferrer Professor d’Investigació Institut de Ciències del Mar (ICM-CSIC) i ii A mis padres A Xavi iii iv Acknowledgements This thesis has been made possible by the support of many people who in one way or another, many times unknowingly, gave me the strength to overcome this "long and winding road". First of all, I would like to thank my supervisor, Dr. Francesc Piferrer, for his patience, guidance and wise advice throughout all this Ph.D. experience. But above all, for the trust he placed on me almost seven years ago when he offered me the opportunity to be part of his team. Thanks also for teaching me how to question always everything, for sharing with me your enthusiasm for science and for giving me the opportunity of learning from you by participating in many projects, collaborations and scientific meetings. I am also thankful to my colleagues (former and present Group of Biology of Reproduction members) for your support and encouragement throughout this journey. To the “exGBRs”, thanks for helping me with my first steps into this world. Working as an undergrad with you Dr.