Diseasespecific and Inflammationindependent Stromal
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

RNA Sequencing Reveals a Slow to Fast Muscle Fiber Type Transition After Olanzapine Infusion in Rats
RESEARCH ARTICLE RNA Sequencing Reveals a Slow to Fast Muscle Fiber Type Transition after Olanzapine Infusion in Rats Christopher J. Lynch1*, Yuping Xu1, Andras Hajnal2, Anna C. Salzberg3, Yuka Imamura Kawasawa4,5,6 1 Department of Cellular and Molecular Physiology, College of Medicine, Penn State University, Hershey, Pennsylvania, 17033, United States of America, 2 Department of Neural and Behavioral Sciences, College of Medicine, Penn State University, Hershey, Pennsylvania, 17033, United States of America, 3 Department a11111 of Public Health Sciences, College of Medicine, Penn State University, Hershey, Pennsylvania, 17033, United States of America, 4 Department of Pharmacology, College of Medicine, Penn State University, Hershey, Pennsylvania, 17033, United States of America, 5 Department of Biochemistry and Molecular Biology, College of Medicine, Penn State University, Hershey, Pennsylvania, 17033, United States of America, 6 The Institute for Personalized Medicine, College of Medicine, Penn State University, Hershey, Pennsylvania, 17033, United States of America * [email protected] OPEN ACCESS Citation: Lynch CJ, Xu Y, Hajnal A, Salzberg AC, Kawasawa YI (2015) RNA Sequencing Reveals a Abstract Slow to Fast Muscle Fiber Type Transition after Olanzapine Infusion in Rats. PLoS ONE 10(4): Second generation antipsychotics (SGAs), like olanzapine, exhibit acute metabolic side ef- e0123966. doi:10.1371/journal.pone.0123966 fects leading to metabolic inflexibility, hyperglycemia, adiposity and diabetes. Understand- Academic Editor: Guillermo López Lluch, ing how SGAs affect the skeletal muscle transcriptome could elucidate approaches for Universidad Pablo de Olavide, Centro Andaluz de mitigating these side effects. Male Sprague-Dawley rats were infused intravenously with ve- Biología del Desarrollo-CSIC, SPAIN hicle or olanzapine for 24h using a dose leading to a mild hyperglycemia. -

Genetic Mutations and Mechanisms in Dilated Cardiomyopathy
Genetic mutations and mechanisms in dilated cardiomyopathy Elizabeth M. McNally, … , Jessica R. Golbus, Megan J. Puckelwartz J Clin Invest. 2013;123(1):19-26. https://doi.org/10.1172/JCI62862. Review Series Genetic mutations account for a significant percentage of cardiomyopathies, which are a leading cause of congestive heart failure. In hypertrophic cardiomyopathy (HCM), cardiac output is limited by the thickened myocardium through impaired filling and outflow. Mutations in the genes encoding the thick filament components myosin heavy chain and myosin binding protein C (MYH7 and MYBPC3) together explain 75% of inherited HCMs, leading to the observation that HCM is a disease of the sarcomere. Many mutations are “private” or rare variants, often unique to families. In contrast, dilated cardiomyopathy (DCM) is far more genetically heterogeneous, with mutations in genes encoding cytoskeletal, nucleoskeletal, mitochondrial, and calcium-handling proteins. DCM is characterized by enlarged ventricular dimensions and impaired systolic and diastolic function. Private mutations account for most DCMs, with few hotspots or recurring mutations. More than 50 single genes are linked to inherited DCM, including many genes that also link to HCM. Relatively few clinical clues guide the diagnosis of inherited DCM, but emerging evidence supports the use of genetic testing to identify those patients at risk for faster disease progression, congestive heart failure, and arrhythmia. Find the latest version: https://jci.me/62862/pdf Review series Genetic mutations and mechanisms in dilated cardiomyopathy Elizabeth M. McNally, Jessica R. Golbus, and Megan J. Puckelwartz Department of Human Genetics, University of Chicago, Chicago, Illinois, USA. Genetic mutations account for a significant percentage of cardiomyopathies, which are a leading cause of conges- tive heart failure. -

Watsonjn2018.Pdf (1.780Mb)
UNIVERSITY OF CENTRAL OKLAHOMA Edmond, Oklahoma Department of Biology Investigating Differential Gene Expression in vivo of Cardiac Birth Defects in an Avian Model of Maternal Phenylketonuria A THESIS SUBMITTED TO THE GRADUATE FACULTY In partial fulfillment of the requirements For the degree of MASTER OF SCIENCE IN BIOLOGY By Jamie N. Watson Edmond, OK June 5, 2018 J. Watson/Dr. Nikki Seagraves ii J. Watson/Dr. Nikki Seagraves Acknowledgements It is difficult to articulate the amount of gratitude I have for the support and encouragement I have received throughout my master’s thesis. Many people have added value and support to my life during this time. I am thankful for the education, experience, and friendships I have gained at the University of Central Oklahoma. First, I would like to thank Dr. Nikki Seagraves for her mentorship and friendship. I lucked out when I met her. I have enjoyed working on this project and I am very thankful for her support. I would like thank Thomas Crane for his support and patience throughout my master’s degree. I would like to thank Dr. Shannon Conley for her continued mentorship and support. I would like to thank Liz Bullen and Dr. Eric Howard for their training and help on this project. I would like to thank Kristy Meyer for her friendship and help throughout graduate school. I would like to thank my committee members Dr. Robert Brennan and Dr. Lilian Chooback for their advisement on this project. Also, I would like to thank the biology faculty and staff. I would like to thank the Seagraves lab members: Jailene Canales, Kayley Pate, Mckayla Muse, Grace Thetford, Kody Harvey, Jordan Guffey, and Kayle Patatanian for their hard work and support. -
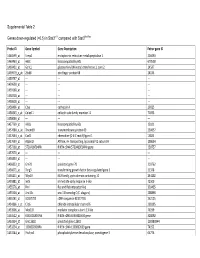
0.5) in Stat3∆/∆ Compared with Stat3flox/Flox
Supplemental Table 2 Genes down-regulated (<0.5) in Stat3∆/∆ compared with Stat3flox/flox Probe ID Gene Symbol Gene Description Entrez gene ID 1460599_at Ermp1 endoplasmic reticulum metallopeptidase 1 226090 1460463_at H60c histocompatibility 60c 670558 1460431_at Gcnt1 glucosaminyl (N-acetyl) transferase 1, core 2 14537 1459979_x_at Zfp68 zinc finger protein 68 24135 1459747_at --- --- --- 1459608_at --- --- --- 1459168_at --- --- --- 1458718_at --- --- --- 1458618_at --- --- --- 1458466_at Ctsa cathepsin A 19025 1458345_s_at Colec11 collectin sub-family member 11 71693 1458046_at --- --- --- 1457769_at H60a histocompatibility 60a 15101 1457680_a_at Tmem69 transmembrane protein 69 230657 1457644_s_at Cxcl1 chemokine (C-X-C motif) ligand 1 14825 1457639_at Atp6v1h ATPase, H+ transporting, lysosomal V1 subunit H 108664 1457260_at 5730409E04Rik RIKEN cDNA 5730409E04Rik gene 230757 1457070_at --- --- --- 1456893_at --- --- --- 1456823_at Gm70 predicted gene 70 210762 1456671_at Tbrg3 transforming growth factor beta regulated gene 3 21378 1456211_at Nlrp10 NLR family, pyrin domain containing 10 244202 1455881_at Ier5l immediate early response 5-like 72500 1455576_at Rinl Ras and Rab interactor-like 320435 1455304_at Unc13c unc-13 homolog C (C. elegans) 208898 1455241_at BC037703 cDNA sequence BC037703 242125 1454866_s_at Clic6 chloride intracellular channel 6 209195 1453906_at Med13l mediator complex subunit 13-like 76199 1453522_at 6530401N04Rik RIKEN cDNA 6530401N04 gene 328092 1453354_at Gm11602 predicted gene 11602 100380944 1453234_at -

Long, Noncoding RNA Dysregulation in Glioblastoma
cancers Review Long, Noncoding RNA Dysregulation in Glioblastoma Patrick A. DeSouza 1,2 , Xuan Qu 1, Hao Chen 1,3, Bhuvic Patel 1 , Christopher A. Maher 2,4,5,6 and Albert H. Kim 1,6,* 1 Department of Neurological Surgery, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA; [email protected] (P.A.D.); [email protected] (X.Q.); [email protected] (H.C.); [email protected] (B.P.) 2 Department of Internal Medicine, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA; [email protected] 3 Department of Neuroscience, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA 4 Department of Biomedical Engineering, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA 5 McDonnell Genome Institute, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA 6 Siteman Cancer Center, Washington University School of Medicine in St. Louis, St. Louis, MO 63110, USA * Correspondence: [email protected] Simple Summary: Developing effective therapies for glioblastoma (GBM), the most common primary brain cancer, remains challenging due to the heterogeneity within tumors and therapeutic resistance that drives recurrence. Noncoding RNAs are transcribed from a large proportion of the genome and remain largely unexplored in their contribution to the evolution of GBM tumors. Here, we will review the general mechanisms of long, noncoding RNAs and the current knowledge of how these impact heterogeneity and therapeutic resistance in GBM. A better understanding of the molecular drivers required for these aggressive tumors is necessary to improve the management and outcomes Citation: DeSouza, P.A.; Qu, X.; of this challenging disease. -

Product Description SALSA MLPA Probemix P476-A1 ZNRF3
MRC-Holland ® Product Description version A1-02; Issued 25 September 2018 MLPA Product Description SALSA ® MLPA ® Probemix P476-A1 ZNRF3 To be used with the MLPA General Protocol. Version A1. New Product. Catalogue numbers: • P476-025R: SALSA MLPA Probemix P476 ZNRF3, 25 reactions. • P476-050R: SALSA MLPA Probemix P476 ZNRF3, 50 reactions. • P476-100R: SALSA MLPA Probemix P476 ZNRF3, 100 reactions. To be used in combination with a SALSA MLPA reagent kit, available for various number of reactions. MLPA reagent kits are either provided with FAM or Cy5.0 dye-labelled PCR primer, suitable for Applied Biosystems and Beckman capillary sequencers, respectively (see www.mlpa.com ). Certificate of Analysis: Information regarding storage conditions, quality tests, and a sample electropherogram from the current sales lot is available at www.mlpa.com . Precautions and warnings: For professional use only. Always consult the most recent product description AND the MLPA General Protocol before use: www.mlpa.com . It is the responsibility of the user to be aware of the latest scientific knowledge of the application before drawing any conclusions from findings generated with this product. General information: The SALSA MLPA Probemix P476 ZNRF3 is a research use only (RUO) assay for the detection of deletions and duplications in the ZNRF3 gene. ZNRF3 is an E3 ubiquitin-protein ligase that acts as a negative feedback regulator of Wnt signalling (Hao et al. 2012). Three independent studies show homozygous deletions of the ZNRF3 gene in 10 to 16% of adrenocortical carcinoma cases (Assié et al. 2014; Juhlin et al. 2015; Zheng et al. 2016). Moreover, in 51% of microsatellite stable colorectal cancers deletion events at the ZNRF3 locus are detected (Bond et al. -

Identification of Active Signaling Pathways by Integrating Gene Expression and Protein Interaction Data Md Humayun Kabir1,2,3, Ralph Patrick2,4,5, Joshua W
Kabir et al. BMC Systems Biology 2018, 12(Suppl 9):120 https://doi.org/10.1186/s12918-018-0655-x RESEARCH Open Access Identification of active signaling pathways by integrating gene expression and protein interaction data Md Humayun Kabir1,2,3, Ralph Patrick2,4,5, Joshua W. K. Ho2,4,6* and Michael D. O’Connor1,7* From 29th International Conference on Genome Informatics Yunnan, China. 3-5 December 2018 Abstract Background: Signaling pathways are the key biological mechanisms that transduce extracellular signals to affect transcription factor mediated gene regulation within cells. A number of computational methods have been developed to identify the topological structure of a specific signaling pathway using protein-protein interaction data, but they are not designed for identifying active signaling pathways in an unbiased manner. On the other hand, there are statistical methods based on gene sets or pathway data that can prioritize likely active signaling pathways, but they do not make full use of active pathway structure that link receptor, kinases and downstream transcription factors. Results: Here, we present a method to simultaneously predict the set of active signaling pathways, together with their pathway structure, by integrating protein-protein interaction network and gene expression data. We evaluated the capacity for our method to predict active signaling pathways for dental epithelial cells, ocular lens epithelial cells, human pluripotent stem cell-derived lens epithelial cells, and lens fiber cells. This analysis showed our approach could identify all the known active pathways that are associated with tooth formation and lens development. Conclusions: The results suggest that SPAGI can be a useful approach to identify the potential active signaling pathways given a gene expression profile. -

140503 IPF Signatures Supplement Withfigs Thorax
Supplementary material for Heterogeneous gene expression signatures correspond to distinct lung pathologies and biomarkers of disease severity in idiopathic pulmonary fibrosis Daryle J. DePianto1*, Sanjay Chandriani1⌘*, Alexander R. Abbas1, Guiquan Jia1, Elsa N. N’Diaye1, Patrick Caplazi1, Steven E. Kauder1, Sabyasachi Biswas1, Satyajit K. Karnik1#, Connie Ha1, Zora Modrusan1, Michael A. Matthay2, Jasleen Kukreja3, Harold R. Collard2, Jackson G. Egen1, Paul J. Wolters2§, and Joseph R. Arron1§ 1Genentech Research and Early Development, South San Francisco, CA 2Department of Medicine, University of California, San Francisco, CA 3Department of Surgery, University of California, San Francisco, CA ⌘Current address: Novartis Institutes for Biomedical Research, Emeryville, CA. #Current address: Gilead Sciences, Foster City, CA. *DJD and SC contributed equally to this manuscript §PJW and JRA co-directed this project Address correspondence to Paul J. Wolters, MD University of California, San Francisco Department of Medicine Box 0111 San Francisco, CA 94143-0111 [email protected] or Joseph R. Arron, MD, PhD Genentech, Inc. MS 231C 1 DNA Way South San Francisco, CA 94080 [email protected] 1 METHODS Human lung tissue samples Tissues were obtained at UCSF from clinical samples from IPF patients at the time of biopsy or lung transplantation. All patients were seen at UCSF and the diagnosis of IPF was established through multidisciplinary review of clinical, radiological, and pathological data according to criteria established by the consensus classification of the American Thoracic Society (ATS) and European Respiratory Society (ERS), Japanese Respiratory Society (JRS), and the Latin American Thoracic Association (ALAT) (ref. 5 in main text). Non-diseased normal lung tissues were procured from lungs not used by the Northern California Transplant Donor Network. -

Nuclear Envelope Laminopathies: Evidence for Developmentally Inappropriate Nuclear Envelope-Chromatin Associations
Nuclear Envelope Laminopathies: Evidence for Developmentally Inappropriate Nuclear Envelope-Chromatin Associations by Jelena Perovanovic M.S. in Molecular Biology and Physiology, September 2009, University of Belgrade M.Phil. in Molecular Medicine, August 2013, The George Washington University A Dissertation submitted to The Faculty of The Columbian College of Arts and Sciences of The George Washington University in partial fulfillment of the requirements for the degree of Doctor of Philosophy August 31, 2015 Dissertation directed by Eric P. Hoffman Professor of Integrative Systems Biology The Columbian College of Arts and Sciences of The George Washington University certifies that Jelena Perovanovic has passed the Final Examination for the degree of Doctor of Philosophy as of May 5, 2015. This is the final and approved form of the dissertation. Nuclear Envelope Laminopathies: Evidence for Developmentally Inappropriate Nuclear Envelope-Chromatin Associations Jelena Perovanovic Dissertation Research Committee: Eric P. Hoffman, Professor of Integrative Systems Biology, Dissertation Director Anamaris Colberg-Poley, Professor of Integrative Systems Biology, Committee Member Robert J. Freishtat, Associate Professor of Pediatrics, Committee Member Vittorio Sartorelli, Senior Investigator, National Institutes of Health, Committee Member ii © Copyright 2015 by Jelena Perovanovic All rights reserved iii Acknowledgments I am deeply indebted to countless individuals for their support and encouragement during the past five years of graduate studies. First and foremost, I would like to express my gratitude to my mentor, Dr. Eric P. Hoffman, for his unwavering support and guidance, and keen attention to my professional development. This Dissertation would not have been possible without the critical input he provided and the engaging environment he created. -

Novel Mutations in TPM2 and PIEZO2 Are Responsible for Distal
Li et al. BMC Medical Genetics (2018) 19:179 https://doi.org/10.1186/s12881-018-0692-8 RESEARCH ARTICLE Open Access Novel mutations in TPM2 and PIEZO2 are responsible for distal arthrogryposis (DA) 2B and mild DA in two Chinese families Shan Li1†, Yi You1†, Jinsong Gao2, Bin Mao1, Yixuan Cao1, Xiuli Zhao1* and Xue Zhang1* Abstract Background: Distal arthrogryposis (DA) is a group of clinically and genetically heterogeneous disorders that involve multiple congenital limb contractures and comprise at least 10 clinical subtypes. Here, we describe our findings in two Chinese families: Family 1 with DA2B (MIM 601680) and Family 2 with mild DA. Methods: To map the disease locus, two-point linkage analysis was performed with microsatellite markers closed to TPM2, TNNI2/TNNT3 and TNNC2. In Family 1, a positive LOD (logarithm of odds) score was only obtained at the microsatellite marker close to TPM2 and mutation screening was performed using direct sequencing of TPM2 in the proband. In Family 2, for the LOD score that did not favor linkage to any markers, whole-exome sequencing (WES) was performed on the proband. PCR–restriction fragment length polymorphism (RFLP) and bioinformatics analysis were then applied to identify the pathogenic mutations in two families. In order to correlate genotype with phenotype in DA, retrospective analyses of phenotypic features according to the TPM2 and PIEZO2 mutation spectrums were carried out. Results: A heterozygous missense mutation c.308A > G (p.Q103R) in TPM2 in Family 1, and a novel variation c.8153G > A (p.R2718Q) in PIEZO2 in Family 2 were identified. -

Spatial Distribution of Leading Pacemaker Sites in the Normal, Intact Rat Sinoa
Supplementary Material Supplementary Figure 1: Spatial distribution of leading pacemaker sites in the normal, intact rat sinoatrial 5 nodes (SAN) plotted along a normalized y-axis between the superior vena cava (SVC) and inferior vena 6 cava (IVC) and a scaled x-axis in millimeters (n = 8). Colors correspond to treatment condition (black: 7 baseline, blue: 100 µM Acetylcholine (ACh), red: 500 nM Isoproterenol (ISO)). 1 Supplementary Figure 2: Spatial distribution of leading pacemaker sites before and after surgical 3 separation of the rat SAN (n = 5). Top: Intact SAN preparations with leading pacemaker sites plotted during 4 baseline conditions. Bottom: Surgically cut SAN preparations with leading pacemaker sites plotted during 5 baseline conditions (black) and exposure to pharmacological stimulation (blue: 100 µM ACh, red: 500 nM 6 ISO). 2 a &DUGLDFIoQChDQQHOV .FQM FOXVWHU &DFQDG &DFQDK *MD &DFQJ .FQLS .FQG .FQK .FQM &DFQDF &DFQE .FQM í $WSD .FQD .FQM í .FQN &DVT 5\U .FQM &DFQJ &DFQDG ,WSU 6FQD &DFQDG .FQQ &DFQDJ &DFQDG .FQD .FQT 6FQD 3OQ 6FQD +FQ *MD ,WSU 6FQE +FQ *MG .FQN .FQQ .FQN .FQD .FQE .FQQ +FQ &DFQDD &DFQE &DOP .FQM .FQD .FQN .FQG .FQN &DOP 6FQD .FQD 6FQE 6FQD 6FQD ,WSU +FQ 6FQD 5\U 6FQD 6FQE 6FQD .FQQ .FQH 6FQD &DFQE 6FQE .FQM FOXVWHU V6$1 L6$1 5$ /$ 3 b &DUGLDFReFHSWRUV $GUDF FOXVWHU $GUDD &DY &KUQE &KUP &KJD 0\O 3GHG &KUQD $GUE $GUDG &KUQE 5JV í 9LS $GUDE 7SP í 5JV 7QQF 3GHE 0\K $GUE *QDL $QN $GUDD $QN $QN &KUP $GUDE $NDS $WSE 5DPS &KUP 0\O &KUQD 6UF &KUQH $GUE &KUQD FOXVWHU V6$1 L6$1 5$ /$ 4 c 1HXURQDOPURWHLQV -

Targeting the Mild-Hypoxia Driving Force for Metabolic and Muscle Transcriptional Reprogramming of Gilthead Sea Bream (Sparus Aurata) Juveniles
biology Article Targeting the Mild-Hypoxia Driving Force for Metabolic and Muscle Transcriptional Reprogramming of Gilthead Sea Bream (Sparus aurata) Juveniles Fernando Naya-Català 1, Juan A. Martos-Sitcha 1,2 , Verónica de las Heras 1, Paula Simó-Mirabet 1 , Josep À. Calduch-Giner 1 and Jaume Pérez-Sánchez 1,* 1 Nutrigenomics and Fish Growth Endocrinology Group, Institute of Aquaculture Torre de la Sal, CSIC, 12595 Ribera de Cabanes, Spain; [email protected] (F.N.-C.); [email protected] (J.A.M.-S.); [email protected] (V.d.l.H.); [email protected] (P.S.-M.); [email protected] (J.À.C.-G.) 2 Department of Biology, Faculty of Marine and Environmental Sciences, Instituto Universitario de Investigación Marina (INMAR), Campus de Excelencia Internacional del Mar (CEI-MAR), University of Cádiz, 11519 Cádiz, Spain * Correspondence: [email protected] Simple Summary: Reduced oxygen availability generates a number of adaptive features across all the animal kingdom, and the goal of this study was targeting the mild-hypoxia driving force for metabolic and muscle transcriptional reprogramming of gilthead sea bream juveniles. Attention was focused on blood metabolic and muscle transcriptomic landmarks before and after exhaustive exercise. Our results after mild-hypoxia conditioning highlighted an increased contribution of lipid Citation: Naya-Català, F.; metabolism to whole energy supply to preserve the aerobic energy production, a better swimming Martos-Sitcha, J.A.; de las Heras, V.; performance regardless of changes in feed intake, as well as reduced protein turnover and improved Simó-Mirabet, P.; Calduch-Giner, J.À.; anaerobic fitness with the restoration of normoxia.