Human Ribonuclease P: Subunits, Function, and Intranuclear Localization
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Yeast Genome Gazetteer P35-65
gazetteer Metabolism 35 tRNA modification mitochondrial transport amino-acid metabolism other tRNA-transcription activities vesicular transport (Golgi network, etc.) nitrogen and sulphur metabolism mRNA synthesis peroxisomal transport nucleotide metabolism mRNA processing (splicing) vacuolar transport phosphate metabolism mRNA processing (5’-end, 3’-end processing extracellular transport carbohydrate metabolism and mRNA degradation) cellular import lipid, fatty-acid and sterol metabolism other mRNA-transcription activities other intracellular-transport activities biosynthesis of vitamins, cofactors and RNA transport prosthetic groups other transcription activities Cellular organization and biogenesis 54 ionic homeostasis organization and biogenesis of cell wall and Protein synthesis 48 plasma membrane Energy 40 ribosomal proteins organization and biogenesis of glycolysis translation (initiation,elongation and cytoskeleton gluconeogenesis termination) organization and biogenesis of endoplasmic pentose-phosphate pathway translational control reticulum and Golgi tricarboxylic-acid pathway tRNA synthetases organization and biogenesis of chromosome respiration other protein-synthesis activities structure fermentation mitochondrial organization and biogenesis metabolism of energy reserves (glycogen Protein destination 49 peroxisomal organization and biogenesis and trehalose) protein folding and stabilization endosomal organization and biogenesis other energy-generation activities protein targeting, sorting and translocation vacuolar and lysosomal -
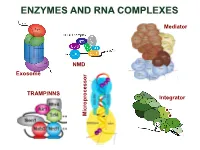
Enzymes and Rna Complexes
ENZYMES AND RNA COMPLEXES Mediator NMD Exosome NMD TRAMP/NNS Integrator Microprocessor RNA PROCESSING and DECAY machinery: RNases Protein Function Characteristics Exonucleases 5’ 3’ Xrn1 cytoplasmic, mRNA degradation processsive Rat1 nuclear, pre-rRNA, sn/snoRNA, pre-mRNA processing and degradation Rrp17/hNol12 nuclear, pre-rRNA processing Exosome 3’ 5’ multisubunit exo/endo complex subunits organized as in bacterial PNPase Rrp44/Dis3 catalytic subunit Exo/PIN domains, processsive Rrp4, Rrp40 pre-rRNA, sn/snoRNA processing, mRNA degradation Rrp41-43, 45-46 participates in NMD, ARE-dependent, non-stop decay Mtr3, Ski4 Mtr4 nuclear helicase cofactor DEAD box Rrp6 (Rrp47) nuclear exonuclease ( Rrp6 BP, cofactor) RNAse D homolog, processsive Ski2,3,7,8 cytoplasmic exosome cofactors. SKI complex helicase, GTPase Other 3’ 5’ Rex1-4 3’-5’ exonucleases, rRNA, snoRNA, tRNA processing RNase D homolog DXO 3’-5’ exonuclease in addition to decapping mtEXO 3’ 5’ mitochondrial degradosome RNA degradation in yeast Suv3/ Dss1 helicase/ 3’-5’ exonuclease DExH box/ RNase II homolog Deadenylation Ccr4/NOT/Pop2 major deadenylase complex (Ccr, Caf, Pop, Not proteins) Ccr4- Mg2+ dependent endonuclease Pan2p/Pan3 additional deadenylases (poliA tail length) RNase D homolog, poly(A) specific nuclease PARN mammalian deadenylase RNase D homolog, poly(A) specific nuclease Endonucleases RNase III -Rnt1 pre-rRNA, sn/snoRNA processing, mRNA degradation dsRNA specific -Dicer, Drosha siRNA/miRNA biogenesis, functions in RNAi PAZ, RNA BD, RNase III domains Ago2 Slicer -

A Disease-Linked Lncrna Mutation in Rnase MRP Inhibits Ribosome Synthesis
bioRxiv preprint doi: https://doi.org/10.1101/2021.03.29.437572; this version posted March 29, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. A disease-linked lncRNA mutation in RNase MRP inhibits ribosome synthesis Nic Roberston1, Vadim Shchepachev1, David Wright2, Tomasz W. Turowski1, Christos Spanos1, Aleksandra Helwak1, Rose Zamoyska2, David Tollervey1 1 Wellcome Centre for Cell Biology, University of Edinburgh, Edinburgh, UK 2 Ashworth Laboratories, Institute of Immunology and Infection Research, University of Edinburgh, Edinburgh, UK Keywords: protein-RNA interaction; RNA-binding sites; UV crosslinking; mass spectrometry; genetic disease; Cartilage Hair Hypoplasia; ribosome synthesis; T cell activation Running title: RNase MRP defects cause ribosomopathy Highlights: • Mutations in RMRP lncRNA impair pre-rRNA processing and T cell activation • Patient derived fibroblasts show impaired pre-rRNA processing • Cells with the most common disease-linked mutation have specific processing defects • Cytoplasmic ribosomes and intact RNase MRP complexes are also reduced in these cells 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.03.29.437572; this version posted March 29, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Abstract Mutations in the human RMRP gene cause Cartilage Hair Hypoplasia (CHH), an autosomal recessive disorder characterized by skeletal abnormalities and impaired T cell activation. -

A Specialized Processing Body That Is Temporally and Asymmetrically Regulated During the Cell Cycle in Saccharomyces Cerevisiae
JCB: ARTICLE A specialized processing body that is temporally and asymmetrically regulated during the cell cycle in Saccharomyces cerevisiae Tina Gill, Jason Aulds, and Mark E. Schmitt Department of Biochemistry and Molecular Biology, State University of New York Upstate Medical University, Syracuse, NY 13210 Nase mitochondrial RNA processing (MRP) is an this spot was asymmetrically found in daughter cells, essential ribonucleoprotein endoribonuclease that where the RNase MRP substrate, CLB2 mRNA, localizes. R functions in the degradation of specifi c mRNAs in- Both the mitotic exit network and fourteen early ana- volved in cell cycle regulation. We have investigated phase release pathways are nonessential but important where this processing event occurs and how it is regu- for the temporal changes in localization. Asymmetric lo- lated. As expected, results demonstrate that RNase MRP calization was found to be dependent on the locasome. is predominantly localized in the nucleolus, where it The evidence suggests that these spots are specialized processes ribosomal RNAs. However, after the initiation processing bodies for the degradation of transcripts that of mitosis, RNase MRP localizes throughout the entire are cell cycle regulated and daughter cell localized. We nucleus and in a single discrete cytoplasmic spot that have called these TAM bodies for temporal asymmetric persists until the completion of telophase. Furthermore, MRP bodies. Introduction RNase mitochondrial RNA processing (MRP) is an essential ri- 27SA preribosomal RNA at the A3 site, forming the 5.8S(s) bonucleoprotein endoribonuclease that cleaves RNA substrates ribosomal RNA (rRNA; Schmitt and Clayton, 1993; Lygerou in a site-specifi c manner and is highly conserved in eukaryotes et al., 1996). -

The Rnase MRP and Rnase P Complexes, Will Be Summarised
PDF hosted at the Radboud Repository of the Radboud University Nijmegen The following full text is a publisher's version. For additional information about this publication click this link. http://hdl.handle.net/2066/19147 Please be advised that this information was generated on 2021-09-28 and may be subject to change. The human The human RNase MRP complex RNase MRP complex Composition, assembly and role in human disease Hans van Eenennaam Hans van Eenennaam The human RNase MRP complex Composition, assembly and role in human disease Hans van Eenennaam, 2002 The human RNase MRP complex Composition, assembly and role in human disease een wetenschappelijke proeve op het gebied van de Natuurwetenschappen, Wiskunde en Informatica PROEFSCHRIFT ter verkrijging van de graad van doctor aan de Katholieke Universiteit Nijmegen, volgens besluit van het College van Decanen in het openbaar te verdedigen op vrijdag 14 juni 2002 des namiddags om 1:30 uur precies door Hans van Eenennaam Cover illustration Statue of the Dwarf Seneb and his family, painted limestone, height 34 cm, width 22.5 cm, geboren op 3 november 1973 Giza, Tomb of Seneb, Late Fifth - Early Sixth te Middelburg Dynasty. From: The Cairo museum Masterpieces of Egyptian Art, Francesco Tiradritti, Thames & Hudson Ltd, London, 1998 Promotor Prof. Dr. W.J. van Venrooij Co-promotor Dr. G.J.M. Pruijn voor mijn ouders Manuscriptcommissie Prof. Dr. L.B.A. van de Putte Prof. Dr. F.P.J.T. Rutjes Prof. Dr. H.F. Tabak (Universiteit van Amsterdam) ISBN 90-9015696-8 © 2002 by Hans van Eenennaam The research described in this thesis was performed at the Department of Biochemistry, Faculty of Science, University of Nijmegen, the Netherlands. -

A Novel Model for the Rnase MRP-Induced Switch Between the Different Forms of 5.8S Rrna
Preprints (www.preprints.org) | NOT PEER-REVIEWED | Posted: 29 April 2021 doi:10.20944/preprints202104.0765.v1 Research article A novel model for the RNase MRP-induced switch between the different forms of 5.8S rRNA Xiao Li1,2,3, Janice M Zengel1 and Lasse Lindahl1 1 Department of Biological Sciences, University of Maryland Baltimore County (UMBC) 1000 Hilltop Circle, Baltimore, Maryland 21250, USA 2 Department of Biology, University of Rochester, Rochester, New York 14627, USA 3 Current address: Abbott Laboratories, San Diego, CA ribosome biogenesis; rRNA processing; RNase MRP; long/short 5.8S rRNA © 2021 by the author(s). Distributed under a Creative Commons CC BY license. Preprints (www.preprints.org) | NOT PEER-REVIEWED | Posted: 29 April 2021 doi:10.20944/preprints202104.0765.v1 Abstract: Processing of the RNA polymerase I pre-rRNA transcript into the mature 18S, 5.8S, and 25S rRNAs requires removing the “spacer” sequences. The canonical pathway for the removal of the ITS1 spacer, located between 18S and 5.8S rRNAs in the primary transcript, involves cleavages at the 3’ end of 18S rRNA and at two sites inside ITS1. The process generates a long and a short 5.8S rRNA that differ in the number of ITS1 nucleotides retained at the 5.8S 5’ end. Here we document a novel pathway that generates the long 5.8S for ITS1 while bypassing cleavage within ITS1. It entails a single endonuclease cut at the 3’-end of 18S rRNA followed by exonuclease Xrn1 degradation of ITS1. Mutations in RNase MRP increase the accumulation of long relative to short 5.8S rRNA; traditionally this is attributed to a decreased rate of RNase MRP cleavage at its target in ITS1, called A3. -

Subject Index Proc
13088 Subject Index Proc. Natl. Acad. Sci. USA 91 (1994) Apoptosis in substantia nigra following developmental striatal excitotoxic Brine shrimp injury, 8117 See Artemia Visualizing hippocampal synaptic function by optical detection of Ca2l Broccol entry through the N-methyl-D-aspartate channel, 8170 See Brassica Amygdala modulation of hippocampal-dependent and caudate Bromophenacyl bromide nucleus-dependent memory processes, 8477 Bromophenacyl bromide binding to the actin-bundling protein I-plastin Distribution of corticotropin-releasing factor receptor mRNA expression inhibits inositol trisphosphate-independent increase in Ca2l in human in the rat brain and pituitary, 8777 neutrophils, 3534 Brownian dynamics Preproenkephalin promoter yields region-specific and long-term Adhesion of hard spheres under the influence of double-layer, van der expression in adult brain after direct in vivo gene transfer via a Waals, and gravitational potentials at a solid/liquid interface, 3004 defective herpes simplex viral vector, 8979 Browsers Intravenous administration of a transferrin receptor antibody-nerve Thorn-like prickles and heterophylly in Cyanea: Adaptations to extinct growth factor conjugate prevents the degeneration of cholinergic avian browsers on Hawaii?, 2810 striatal neurons in a model of Huntington disease, 9077 Bruton agammaglobulinemia Axotomy induces the expression of vasopressin receptors in cranial and Genomic organization and structure of Bruton agammaglobulinemia spinal motor nuclei in the adult rat, 9636 tyrosine kinase: Localization -

Genome-Wide Investigation of Cellular Functions for Trna Nucleus
Genome-wide Investigation of Cellular Functions for tRNA Nucleus- Cytoplasm Trafficking in the Yeast Saccharomyces cerevisiae DISSERTATION Presented in Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy in the Graduate School of The Ohio State University By Hui-Yi Chu Graduate Program in Molecular, Cellular and Developmental Biology The Ohio State University 2012 Dissertation Committee: Anita K. Hopper, Advisor Stephen Osmani Kurt Fredrick Jane Jackman Copyright by Hui-Yi Chu 2012 Abstract In eukaryotic cells tRNAs are transcribed in the nucleus and exported to the cytoplasm for their essential role in protein synthesis. This export event was thought to be unidirectional. Surprisingly, several lines of evidence showed that mature cytoplasmic tRNAs shuttle between nucleus and cytoplasm and their distribution is nutrient-dependent. This newly discovered tRNA retrograde process is conserved from yeast to vertebrates. Although how exactly the tRNA nuclear-cytoplasmic trafficking is regulated is still under investigation, previous studies identified several transporters involved in tRNA subcellular dynamics. At least three members of the β-importin family function in tRNA nuclear-cytoplasmic intracellular movement: (1) Los1 functions in both the tRNA primary export and re-export processes; (2) Mtr10, directly or indirectly, is responsible for the constitutive retrograde import of cytoplasmic tRNA to the nucleus; (3) Msn5 functions solely in the re-export process. In this thesis I focus on the physiological role(s) of the tRNA nuclear retrograde pathway. One possibility is that nuclear accumulation of cytoplasmic tRNA serves to modulate translation of particular transcripts. To test this hypothesis, I compared expression profiles from non-translating mRNAs and polyribosome-bound translating mRNAs collected from msn5Δ and mtr10Δ mutants and wild-type cells, in fed or acute amino acid starvation conditions. -

Nuclear Domains of the RNA Subunit of Rnase P
Journal of Cell Science 110, 829-837 (1997) 829 Printed in Great Britain © The Company of Biologists Limited 1997 JCS8126 Nuclear domains of the RNA subunit of RNase P Marty R. Jacobson1, Long-Guang Cao1,*, Krishan Taneja2, Robert H. Singer2,†, Yu-li Wang1 and Thoru Pederson1,‡ 1Cell Biology Group, Worcester Foundation for Biomedical Research, Shrewsbury, MA 01545, USA 2Department of Cell Biology, University of Massachusetts Medical Center, Worcester, MA 01655, USA *Present address: Department of Molecular Genetics and Microbiology, University of Florida, PO Box 100266, Gainesville, FL 32610, USA †Present address: Department of Anatomy and Structural Biology, Albert Einstein College of Medicine, Bronx, NY 10461-1975, USA ‡Author for correspondence (e-mail: [email protected]) SUMMARY The ribonucleoprotein enzyme RNase P catalyzes the 5′ microinjection. In contrast, a truncated RNase P RNA con- processing of pre-transfer RNA, and has also recently been taining the To binding domain but lacking nucleotides 89- implicated in pre-ribosomal RNA processing. In the 341 became rapidly localized in nucleoli following nuclear present investigation, in situ hybridization revealed that microinjection. However, unlike the full-length RNase P RNase P RNA is present throughout the nucleus of RNA, this 3′ truncated RNA remained stably associated mammalian cells. However, rhodamine-labeled human with the nucleoli and did not translocate to the nucleo- RNase P RNA microinjected into the nucleus of rat kidney plasm. These results suggest a nucleolar phase in the mat- (NRK) epithelial cells or human (HeLa) cells initially uration, ribonucleoprotein assembly or function of RNase localized in nucleoli, and subsequently became more evenly P RNA, mediated at least in part by the nucleolar To distributed throughout the nucleus, similar to the steady- antigen. -

Title 1 Transcriptomic Analysis of Ribosome Biogenesis and Pre-Rrna
bioRxiv preprint doi: https://doi.org/10.1101/2021.08.01.454488; this version posted August 1, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Title 2 Transcriptomic analysis of ribosome biogenesis and pre-rRNA processing during growth 3 stress in Entamoeba histolytica ∗1 4 Sarah Naiyer ,2, Shashi Shekhar Singh2,5, Devinder Kaur2,6, Yatendra Pratap Singh2, Amartya 5 Mukherjee2,3, Alok Bhattacharya4 and Sudha Bhattacharya2,4. 6 2- School of Environmental Sciences, Jawaharlal Nehru University, New Delhi, India-110067 7 3- Present Address: Department of Molecular Reproduction, Development and Genetics, Indian 8 Institute of Science Bangalore-560012 9 4- Present Address: Ashoka University, Rajiv Gandhi Education City, Sonipat, Haryana, India - 10 131029 11 5- Present Address: Department of inflammation and Immunity, Cleveland Clinic, Cleveland, 12 OH, USA- 44195 13 6- Present Address: Central University of Punjab, Bathinda- 151401 14 15 16 17 *-Corresponding author 18 1- Present Address 19 Sarah Naiyer, PhD 20 Department of Immunology and Microbiology 21 University of Illinois at Chicago, USA, 60612 22 Email: [email protected], [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.08.01.454488; this version posted August 1, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

Cryo-EM Structure of Catalytic Ribonucleoprotein Complex Rnase MRP
bioRxiv preprint doi: https://doi.org/10.1101/2020.03.17.996132; this version posted March 18, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Cryo-EM structure of catalytic ribonucleoprotein complex RNase MRP Authors: Anna Perederina1, Di Li1, Hyunwook Lee1, Carol Bator1, Igor Berezin1, Susan L. Hafenstein1,2, Andrey S. Krasilnikov1,3* Affiliations: 1Department of Biochemistry and Molecular Biology, Pennsylvania State University, University Park, PA 16802 2Department of Medicine, Pennsylvania State University, Hershey, PA 17033 3Center for RNA Biology, Pennsylvania State University, University Park, PA 16802 *Correspondence to: [email protected] Abstract: RNase MRP is an essential eukaryotic ribonucleoprotein complex involved in the maturation of rRNA and the regulation of the cell cycle. RNase MRP is related to the ribozyme-based RNase P, but it has evolved to have distinct cellular roles. We report a cryo-EM structure of the S. cerevisiae RNase MRP holoenzyme solved to 3.0 Å. We describe the structure of this 450 kDa complex, interactions between its components, and the organization of its catalytic RNA. We show that while the catalytic center of RNase MRP is inherited from the ancestral enzyme RNase P, the substrate binding pocket of RNase MRP is significantly altered by the addition of unique RNA and protein elements, as well as by RNA-driven protein remodeling. One Sentence Summary: Changes in peripheral RNA elements and RNA-driven protein remodeling result in diversification of related catalytic RNPs 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.03.17.996132; this version posted March 18, 2020. -

Targeted CRISPR Disruption Reveals a Role for Rnase MRP RNA in Human Preribosomal RNA Processing
Downloaded from genesdev.cshlp.org on September 27, 2021 - Published by Cold Spring Harbor Laboratory Press Targeted CRISPR disruption reveals a role for RNase MRP RNA in human preribosomal RNA processing Katherine C. Goldfarb1,2 and Thomas R. Cech1,2 1Department of Chemistry and Biochemistry, BioFrontiers Institute, University of Colorado at Boulder, Boulder, Colorado 80302, USA; 2Howard Hughes Medical Institute, University of Colorado at Boulder, Boulder, Colorado 80302, USA MRP RNA is an abundant, essential noncoding RNA whose functions have been proposed in yeast but are incom- pletely understood in humans. Mutations in the genomic locus for MRP RNA cause pleiotropic human diseases, including cartilage hair hypoplasia (CHH). Here we applied CRISPR–Cas9 genome editing to disrupt the endogenous human MRP RNA locus, thereby attaining what has eluded RNAi and RNase H experiments: elimination of MRP RNA in the majority of cells. The resulting accumulation of ribosomal RNA (rRNA) precursor—analyzed by RNA fluorescent in situ hybridization (FISH), Northern blots, and RNA sequencing—implicates MRP RNA in pre-rRNA processing. Amelioration of pre-rRNA imbalance is achieved through rescue of MRP RNA levels by ectopic ex- pression. Furthermore, affinity-purified MRP ribonucleoprotein (RNP) from HeLa cells cleaves the human pre-rRNA in vitro at at least one site used in cells, while RNP isolated from cells with CRISPR-edited MRP loci loses this activity, and ectopic MRP RNA expression restores cleavage activity. Thus, a role for RNase MRP in human pre-rRNA processing is established. As demonstrated here, targeted CRISPR disruption is a valuable tool for functional studies of essential noncoding RNAs that are resistant to RNAi and RNase H-based degradation.