Program Bound to Inspire
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Folic Acid Antagonists: Antimicrobial and Immunomodulating Mechanisms and Applications
International Journal of Molecular Sciences Review Folic Acid Antagonists: Antimicrobial and Immunomodulating Mechanisms and Applications Daniel Fernández-Villa 1, Maria Rosa Aguilar 1,2 and Luis Rojo 1,2,* 1 Instituto de Ciencia y Tecnología de Polímeros, Consejo Superior de Investigaciones Científicas, CSIC, 28006 Madrid, Spain; [email protected] (D.F.-V.); [email protected] (M.R.A.) 2 Consorcio Centro de Investigación Biomédica en Red de Bioingeniería, Biomateriales y Nanomedicina, 28029 Madrid, Spain * Correspondence: [email protected]; Tel.: +34-915-622-900 Received: 18 September 2019; Accepted: 7 October 2019; Published: 9 October 2019 Abstract: Bacterial, protozoan and other microbial infections share an accelerated metabolic rate. In order to ensure a proper functioning of cell replication and proteins and nucleic acids synthesis processes, folate metabolism rate is also increased in these cases. For this reason, folic acid antagonists have been used since their discovery to treat different kinds of microbial infections, taking advantage of this metabolic difference when compared with human cells. However, resistances to these compounds have emerged since then and only combined therapies are currently used in clinic. In addition, some of these compounds have been found to have an immunomodulatory behavior that allows clinicians using them as anti-inflammatory or immunosuppressive drugs. Therefore, the aim of this review is to provide an updated state-of-the-art on the use of antifolates as antibacterial and immunomodulating agents in the clinical setting, as well as to present their action mechanisms and currently investigated biomedical applications. Keywords: folic acid antagonists; antifolates; antibiotics; antibacterials; immunomodulation; sulfonamides; antimalarial 1. -

Recent Advances in Biocatalysts Engineering for Polyethylene Terephthalate Plastic Waste Green Recycling
Environment International 145 (2020) 106144 Contents lists available at ScienceDirect Environment International journal homepage: www.elsevier.com/locate/envint Review article Recent advances in biocatalysts engineering for polyethylene terephthalate plastic waste green recycling Nadia A. Samak a,b,c,1, Yunpu Jia a,b,1, Moustafa M. Sharshar a,b, Tingzhen Mu a, Maohua Yang a, Sumit Peh a,b, Jianmin Xing a,b,* a CAS Key Laboratory of Green Process and Engineering & State Key Laboratory of Biochemical Engineering, Institute of Process Engineering, Chinese Academy of Sciences, Beijing 100190, PR China b College of Chemical Engineering, University of Chinese Academy of Sciences, 19 A Yuquan Road, Beijing 100049, PR China c Processes Design and Development Department, Egyptian Petroleum Research Institute, Nasr City, 11727 Cairo, Egypt ARTICLE INFO ABSTRACT Handling Editor: Guo-ping Sheng The massive waste of poly(ethylene terephthalate) (PET) that ends up in the landfills and oceans and needs hundreds of years for degradation has attracted global concern. The poor stability and productivity of the Keywords: available PET biocatalysts hinder their industrial applications. Active PET biocatalysts can provide a promising Plastic waste avenue for PET bioconversion and recycling. Therefore, there is an urgent need to develop new strategies that Poly(ethylene terephthalate) could enhance the stability, catalytic activity, solubility, productivity, and re-usability of these PET biocatalysts Recycling under harsh conditions such as high temperatures, pH, and salinity. This has raised great attention in using Biocatalysts ’ Bioengineering bioengineering strategies to improve PET biocatalysts robustness and catalytic behavior. Herein, historical and forecasting data of plastic production and disposal were critically reviewed. -

1 SUPPLEMENTARY DATA Table S1
Electronic Supplementary Material (ESI) for Food & Function. This journal is © The Royal Society of Chemistry 2020 SUPPLEMENTARY DATA Table S1: Effects of catalase, protease, papain and pH treatments on the antimicrobial activity (%) of the CFS of S. thermophilus strain against G. vaginalis ATCC 14018. The antimicrobial activity of the strain was highest under acidic pH conditions. Remaining antimicrobial activity (%) Strains 5 mg mL-1 1 mg mL-1 1 mg mL-1 pH treatments catalase proteinase K papain 3.5 4.0 5.0 6.0 6.5 7.0 S. thermophilus KLDS 3.1003 90.49±0.62 100±0 100±0 100±0 100±0 67.02±0.08 6.35±0.03 0±0 0±0 All values were determined in triplicate Values were expressed as mean ± SD 1 Table S2: Weekly weights of study animals before and after S. aureus ATCC25923 infection. Significant variations (at P > 0.05) were observed in the weight of study animals during the study. Period C C C C Week 0 22.00b 23.57b 19.34e 23.24a 20.90d 22.84c 20.19d 22.60b 23.92a 23.70b 20.56c 23.78a 20.89d 23.10c 20.00d 23.51a Average 22.43 23.30 20.02 23.28 SD 1.29 0.40 0.51 0.51 C TST TSA TSTSA 24.92a 24.15a 21.19c 20.98c Week 1 23.34a 23.13c 20.26d 22.68b 21.98b 22.96d 20.85c 22.86b 22.60b 22.36d 24.28a 22.33b Average 23.21 23.15 21.65 22.21 SD 1.27 0.74 1.80 0.86 23.25a 24.28a 19.94d 20.39d Week 2 21.32c 22.09e 18.46f 20.35d 20.29d 21.82e 19.40e 16.68e 21.27c 22.89d 22.74b 20.09d Average 21.53 22.77 20.14 19.38 SD 1.24 1.10 1.84 1.80 Values with the same alphabet along the same column are not significantly different (P > 0.05) Composition of control and trial diets are given in Table 1. -

Roles of Vitamin Metabolizing Genes in Multidrug-Resistant Plasmids of Superbugs
bioRxiv preprint doi: https://doi.org/10.1101/285403; this version posted March 20, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Roles of Vitamin Metabolizing Genes in Multidrug-Resistant Plasmids of Superbugs Asit Kumar Chakraborty, Genetic Engineering Laboratory, Department of Biotechnology & Biochemistry, Oriental Institute of Science & Technology, Vidyasagar University, Midnapore 721102, West Bengal, India. Abstract Superbug crisis has rocked this world with million deaths due to failure of potent antibiotics. Thousands mdr genes with hundreds of mutant isomers are generated. Small integrons and R-plasmids have combined with F'-plasmids creating a space for >10-20 of mdr genes that inactivate antibiotics in different mechanisms. Mdr genes are created to save bacteria from antibiotics because gut microbiota synthesize >20 vitamins and complex bio-molecules needed for >30000 biochemical reactions of human metabolosome. In other words, mdr gene creation is protected from both sides, intestinal luminal cells and gut bacteria in a tight symbiotic signalling system. We have proposed, to avert the crisis all vitamin metabolizing genes will be acquired in MDR- plasmids if we continue oral antibiotics therapy. Therefore, we have checked the plasmid databases and have detected thiamine, riboflavin, folate, cobalamine and biotin metabolizing enzymes in MDR plasmids. Thus vit genes may mobilise recently into MDR-plasmids and are likely essential for gut microbiota protection. Analysis found that cob and thi genes are abundant and likely very essential than other vit genes. -

Letters to Nature
letters to nature Received 7 July; accepted 21 September 1998. 26. Tronrud, D. E. Conjugate-direction minimization: an improved method for the re®nement of macromolecules. Acta Crystallogr. A 48, 912±916 (1992). 1. Dalbey, R. E., Lively, M. O., Bron, S. & van Dijl, J. M. The chemistry and enzymology of the type 1 27. Wolfe, P. B., Wickner, W. & Goodman, J. M. Sequence of the leader peptidase gene of Escherichia coli signal peptidases. Protein Sci. 6, 1129±1138 (1997). and the orientation of leader peptidase in the bacterial envelope. J. Biol. Chem. 258, 12073±12080 2. Kuo, D. W. et al. Escherichia coli leader peptidase: production of an active form lacking a requirement (1983). for detergent and development of peptide substrates. Arch. Biochem. Biophys. 303, 274±280 (1993). 28. Kraulis, P.G. Molscript: a program to produce both detailed and schematic plots of protein structures. 3. Tschantz, W. R. et al. Characterization of a soluble, catalytically active form of Escherichia coli leader J. Appl. Crystallogr. 24, 946±950 (1991). peptidase: requirement of detergent or phospholipid for optimal activity. Biochemistry 34, 3935±3941 29. Nicholls, A., Sharp, K. A. & Honig, B. Protein folding and association: insights from the interfacial and (1995). the thermodynamic properties of hydrocarbons. Proteins Struct. Funct. Genet. 11, 281±296 (1991). 4. Allsop, A. E. et al.inAnti-Infectives, Recent Advances in Chemistry and Structure-Activity Relationships 30. Meritt, E. A. & Bacon, D. J. Raster3D: photorealistic molecular graphics. Methods Enzymol. 277, 505± (eds Bently, P. H. & O'Hanlon, P. J.) 61±72 (R. Soc. Chem., Cambridge, 1997). -

2019 Abstracts
We are delighted EDITORIAL to welcome the worldwide yeast community in Gothenburg for ICYGMB2019! The “International Yeast Conferences” started in the 1960s with a handful of delegates and since then have become THE most important event in yeast research. Now the yeast meeting to returns to Gothenburg. Many yeast researchers still remember the meeting in 2003 with over 1,100 delegates, a truly memorable event. The Life Sciences are changing, and yeast research remains at their forefront. Advancements in genome sequencing and genome editing just make yeast more exciting as model organism in basic cell biological research, genome evolution and as a tool for synthetic biology and biotechnology. One of the most important reasons for the enormous success of yeast research lies in the unique character of the international yeast research community. No other community employs such a free exchange and access to information and research tools. Nor has any other community had the ability to build – even intercontinental – consortia of critical mass to tackle large‐scale projects, such as in sequencing the first eukaryotic genome or the first comprehensive yeast knockout library. Yeast2019 is the meeting of the international yeast research community where the latest, and even unpublished results are exchanged, and new projects, alliances, and collaborations are founded. A do‐not‐miss‐event. We attempt to incorporate the present excitement in yeast research in the programme of yeast2019. We are confident that this conference will contain important news and information for all yeast researchers. Taken together, yeast2019 will provide an up‐ to‐date overview in yeast research and it will set the scene for years to come. -

Teaching About the Unintended Consequences of Plastics Tamara
From Disposable Culture to Disposable People: Teaching About the Unintended Consequences of Plastics Tamara "Sasha" Adkins, MPH Dissertation Committee Dr. Alesia Maltz, Committee Chair Core Faculty, Environmental Studies, Antioch University New England Dr. Abigail Abrash Walton, Teaching Faculty, Environmental Studies, Antioch University New England Dr. Janaki Natarajan Tschannerl, Director, SPARK Teacher Education / Educational Praxis & Director, Bapagrama Educational Center, Bangalore, India From Disposable Culture to Disposable People: Teaching About the Unintended Consequences of Plastics By Tamara "Sasha" Adkins, DISSERTATION Submitted in partial fulfillment of the requirements for the degree of Ph.D. in the Department of Environmental Studies Antioch University New England Keene, New Hampshire November 2017 Copyright 2017 Tamara “Sasha” Adkins All Rights Reserved Dedication Zendaya, To you I dedicate not only this dissertation but all my work to shape a world where no one is disposable. i Acknowledgments Alesia, my adviser and mentor for the past decade, sets a high bar for wisdom, compassion, and kindness. I will always remember you saying, "Oh, sweetie, too many sighs! If you are not having fun (with the dissertation research) then something is wrong." Thank you for not only patiently drawing out my best work, but for making the process a delightful one. Janaki, thank you for all I am learning from you in Spark and through this process. Abi joined my committee officially in my final semester, but had already been giving me encouragement and support for many years. It is much appreciated. I'd like to thank Charles Curtin, who served on my committee for a time, but due to extenuating circumstances, was not able to continue in that role. -

Product Sheet Info
Master Clone List for NR-19279 ® Vibrio cholerae Gateway Clone Set, Recombinant in Escherichia coli, Plates 1-46 Catalog No. NR-19279 Table 1: Vibrio cholerae Gateway® Clones, Plate 1 (NR-19679) Clone ID Well ORF Locus ID Symbol Product Accession Position Length Number 174071 A02 367 VC2271 ribD riboflavin-specific deaminase NP_231902.1 174346 A03 336 VC1877 lpxK tetraacyldisaccharide 4`-kinase NP_231511.1 174354 A04 342 VC0953 holA DNA polymerase III, delta subunit NP_230600.1 174115 A05 388 VC2085 sucC succinyl-CoA synthase, beta subunit NP_231717.1 174310 A06 506 VC2400 murC UDP-N-acetylmuramate--alanine ligase NP_232030.1 174523 A07 132 VC0644 rbfA ribosome-binding factor A NP_230293.2 174632 A08 322 VC0681 ribF riboflavin kinase-FMN adenylyltransferase NP_230330.1 174930 A09 433 VC0720 phoR histidine protein kinase PhoR NP_230369.1 174953 A10 206 VC1178 conserved hypothetical protein NP_230823.1 174976 A11 213 VC2358 hypothetical protein NP_231988.1 174898 A12 369 VC0154 trmA tRNA (uracil-5-)-methyltransferase NP_229811.1 174059 B01 73 VC2098 hypothetical protein NP_231730.1 174075 B02 82 VC0561 rpsP ribosomal protein S16 NP_230212.1 174087 B03 378 VC1843 cydB-1 cytochrome d ubiquinol oxidase, subunit II NP_231477.1 174099 B04 383 VC1798 eha eha protein NP_231433.1 174294 B05 494 VC0763 GTP-binding protein NP_230412.1 174311 B06 314 VC2183 prsA ribose-phosphate pyrophosphokinase NP_231814.1 174603 B07 108 VC0675 thyA thymidylate synthase NP_230324.1 174474 B08 466 VC1297 asnS asparaginyl-tRNA synthetase NP_230942.2 174933 B09 198 -

Supplementary Information
Supplementary information (a) (b) Figure S1. Resistant (a) and sensitive (b) gene scores plotted against subsystems involved in cell regulation. The small circles represent the individual hits and the large circles represent the mean of each subsystem. Each individual score signifies the mean of 12 trials – three biological and four technical. The p-value was calculated as a two-tailed t-test and significance was determined using the Benjamini-Hochberg procedure; false discovery rate was selected to be 0.1. Plots constructed using Pathway Tools, Omics Dashboard. Figure S2. Connectivity map displaying the predicted functional associations between the silver-resistant gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S3. Connectivity map displaying the predicted functional associations between the silver-sensitive gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S4. Metabolic overview of the pathways in Escherichia coli. The pathways involved in silver-resistance are coloured according to respective normalized score. Each individual score represents the mean of 12 trials – three biological and four technical. Amino acid – upward pointing triangle, carbohydrate – square, proteins – diamond, purines – vertical ellipse, cofactor – downward pointing triangle, tRNA – tee, and other – circle. -

Studies with PET-Hydrolyzing Enzymes
MASTERARBEIT / MASTER’S THESIS Titel der Masterarbeit / Title of the Master‘s Thesis „Investigating the properties of PET-hydrolase and PBS- depolymerase“ verfasst von / submitted by Christoph Clemens Hinterberger, BSc angestrebter akademischer Grad / in partial fulfilment of the requirements for the degree of Master of Science (MSc) Wien, 2018/ Vienna 2018 Studienkennzahl lt. Studienblatt / A 066 830 degree programme code as it appears on the student record sheet: Studienrichtung lt. Studienblatt / Masterstudium Molekulare degree programme as it appears on Mikrobiologie, Mikrobielle Ökologie und the student record sheet: Immunbiologie Betreut von / Supervisor: Univ.-Prof. Dr. Udo Bläsi Declaration I hereby declare that this thesis was composed by myself, that the work contained herein is my own except where explicitly stated otherwise in the text, and that this work has not been submitted for any other degree or processional qualification except as specified. Acknowledgements First and foremost, I have to thank Prof. Uwe T. Bornscheuer, who gave me the opportunity of working on this topic, and Dr. Dominique Böttcher, for supervising my work. My thanks also go out to everyone in Dr. Bornscheuer’s group, who helped me along the way, especially to the ever smiling Lukas. Furthermore I have to thank people in my life which made all of this possible: My parents who support me on every step on the road, my siblings, Paul, Max, Tao Su and everyone else in the hard core, Nils and everyone else from the Pack, Linda and Michelle and finally Julia. You’ve all done more than you think. Table of Contents Declaration .............................................................................................................................................. 2 Acknowledgements ................................................................................................................................ -

Enzymatic PET Degradation
GREEN AND SUSTAINABLE CHEMISTRY CHIMIA 2019, 73, No. 9 743 doi:10.2533/chimia.2019.743 Chimia 73 (2019) 743–749 © Swiss Chemical Society Enzymatic PET Degradation Athena Papadopoulou§, Katrin Hecht§, and Rebecca Buller* Abstract: Plastic, in the form of packaging material, disposables, clothing and other articles with a short lifespan, has become an indispensable part of our everyday life. The increased production and use of plastic, however, accelerates the accumulation of plastic waste and poses an increasing burden on the environment with negative effects on biodiversity and human health. PET, a common thermoplastic, is recycled in many countries via ther- mal, mechanical and chemical means. Recently, several enzymes have been identified capable of degrading this recalcitrant plastic, opening possibilities for the biological recycling of the omnipresent material. In this review, we analyze the current knowledge of enzymatic PET degradation and discuss advances in improving the involved enzymes via protein engineering. Looking forward, the use of plastic degrading enzymes may facilitate sustain- able plastic waste management and become an important tool for the realization of a circular plastic economy. Keywords: Biocatalysis · Biodegradation · Enzyme engineering · Plastic recycling · PET Dr. Athena Papadopoulou studied Biology 1. Introduction and received her BSc from the University Plastic has become an omnipresent material in our daily life of Ioannina. She completed her MSc in and, as a consequence, the plastic industry has become the seventh Chemistry with a focus on Chemical and most important industry in Europe, employing more than 1.5 mil- Biochemical Technologies. In 2013 she lion people with a turnover of 355 billion Euros in 2017.[1] Plastic moved to Biotechnology group at the production is cheap and the generated plastic items are durable University of Ioannina to pursue her PhD and versatile. -
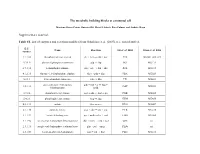
The Metabolic Building Blocks of a Minimal Cell Supplementary
The metabolic building blocks of a minimal cell Mariana Reyes-Prieto, Rosario Gil, Mercè Llabrés, Pere Palmer and Andrés Moya Supplementary material. Table S1. List of enzymes and reactions modified from Gabaldon et. al. (2007). n.i.: non identified. E.C. Name Reaction Gil et. al. 2004 Glass et. al. 2006 number 2.7.1.69 phosphotransferase system glc + pep → g6p + pyr PTS MG041, 069, 429 5.3.1.9 glucose-6-phosphate isomerase g6p ↔ f6p PGI MG111 2.7.1.11 6-phosphofructokinase f6p + atp → fbp + adp PFK MG215 4.1.2.13 fructose-1,6-bisphosphate aldolase fbp ↔ gdp + dhp FBA MG023 5.3.1.1 triose-phosphate isomerase gdp ↔ dhp TPI MG431 glyceraldehyde-3-phosphate gdp + nad + p ↔ bpg + 1.2.1.12 GAP MG301 dehydrogenase nadh 2.7.2.3 phosphoglycerate kinase bpg + adp ↔ 3pg + atp PGK MG300 5.4.2.1 phosphoglycerate mutase 3pg ↔ 2pg GPM MG430 4.2.1.11 enolase 2pg ↔ pep ENO MG407 2.7.1.40 pyruvate kinase pep + adp → pyr + atp PYK MG216 1.1.1.27 lactate dehydrogenase pyr + nadh ↔ lac + nad LDH MG460 1.1.1.94 sn-glycerol-3-phosphate dehydrogenase dhp + nadh → g3p + nad GPS n.i. 2.3.1.15 sn-glycerol-3-phosphate acyltransferase g3p + pal → mag PLSb n.i. 2.3.1.51 1-acyl-sn-glycerol-3-phosphate mag + pal → dag PLSc MG212 acyltransferase 2.7.7.41 phosphatidate cytidyltransferase dag + ctp → cdp-dag + pp CDS MG437 cdp-dag + ser → pser + 2.7.8.8 phosphatidylserine synthase PSS n.i. cmp 4.1.1.65 phosphatidylserine decarboxylase pser → peta PSD n.i.