Roles of Vitamin Metabolizing Genes in Multidrug-Resistant Plasmids of Superbugs
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Folic Acid Antagonists: Antimicrobial and Immunomodulating Mechanisms and Applications
International Journal of Molecular Sciences Review Folic Acid Antagonists: Antimicrobial and Immunomodulating Mechanisms and Applications Daniel Fernández-Villa 1, Maria Rosa Aguilar 1,2 and Luis Rojo 1,2,* 1 Instituto de Ciencia y Tecnología de Polímeros, Consejo Superior de Investigaciones Científicas, CSIC, 28006 Madrid, Spain; [email protected] (D.F.-V.); [email protected] (M.R.A.) 2 Consorcio Centro de Investigación Biomédica en Red de Bioingeniería, Biomateriales y Nanomedicina, 28029 Madrid, Spain * Correspondence: [email protected]; Tel.: +34-915-622-900 Received: 18 September 2019; Accepted: 7 October 2019; Published: 9 October 2019 Abstract: Bacterial, protozoan and other microbial infections share an accelerated metabolic rate. In order to ensure a proper functioning of cell replication and proteins and nucleic acids synthesis processes, folate metabolism rate is also increased in these cases. For this reason, folic acid antagonists have been used since their discovery to treat different kinds of microbial infections, taking advantage of this metabolic difference when compared with human cells. However, resistances to these compounds have emerged since then and only combined therapies are currently used in clinic. In addition, some of these compounds have been found to have an immunomodulatory behavior that allows clinicians using them as anti-inflammatory or immunosuppressive drugs. Therefore, the aim of this review is to provide an updated state-of-the-art on the use of antifolates as antibacterial and immunomodulating agents in the clinical setting, as well as to present their action mechanisms and currently investigated biomedical applications. Keywords: folic acid antagonists; antifolates; antibiotics; antibacterials; immunomodulation; sulfonamides; antimalarial 1. -

The Influence of Vitamin B12on the Content, Distribution and in Vivo
The Influence of Vitamin B12on the Content, Distribution and in Vivo Synthesis of Thiamine Pyrophosphate, Flavin Adenine Dinucleotide and Pyridine Nucleotides in Rat Liver1 URMILA MARFATIA, D. V. REGE, H. P. TIPNIS ANDA. SREENIVASAN Department of Chemical Technology, University of Bombay, Matunga, Bombay Apart from the well-known interrelation (FAD) and pyridine nucleotides (PN), and ships among the B vitamins, there is rea to their in vivo synthesis from the corre son to believe that folie acid and vitamin sponding administered vitamins, in the rat Downloaded from Bu may influence the functioning of other liver. Data on the distribution of these vitamins as cofactors. Thus, dietary folie cofactors in liver cells of the normal rat acid has been known to determine rat are available in the works of Goethart ('52) liver stores of coenzyme A (CoA) and and Dianzani and Dianzani Mor ('57) on adenotriphosphate (ATP) (Popp and Tot TPP, of Carruthers and Suntzeff ('54) and ter, '52; Totter, '53); a decrease in liver Dianzani ('55) on pyridine nucleotides DPN is also caused by aminopterin2 (PN) and of Schneider and Hogeboom jn.nutrition.org (Strength et al., '54). The in vivo incor (Schneider, '56) on FAD. poration of nitcotinamide into pyridino- nucleotides in rat liver is affected in a EXPERIMENTAL deficiency of vitamin B«(Nadkarni et al., Young, male Wistar rats weighing ap '57). Low blood level of citrovorum factor proximately 100 gm each were used. The by guest on April 19, 2011 in the hyperthyroid, vitamin Bi2-deficient animals, housed individually in raised rat is corrected by administration of vita mesh-bottom cages, were initially depleted min B«(Pfander et al., '52). -

1 SUPPLEMENTARY DATA Table S1
Electronic Supplementary Material (ESI) for Food & Function. This journal is © The Royal Society of Chemistry 2020 SUPPLEMENTARY DATA Table S1: Effects of catalase, protease, papain and pH treatments on the antimicrobial activity (%) of the CFS of S. thermophilus strain against G. vaginalis ATCC 14018. The antimicrobial activity of the strain was highest under acidic pH conditions. Remaining antimicrobial activity (%) Strains 5 mg mL-1 1 mg mL-1 1 mg mL-1 pH treatments catalase proteinase K papain 3.5 4.0 5.0 6.0 6.5 7.0 S. thermophilus KLDS 3.1003 90.49±0.62 100±0 100±0 100±0 100±0 67.02±0.08 6.35±0.03 0±0 0±0 All values were determined in triplicate Values were expressed as mean ± SD 1 Table S2: Weekly weights of study animals before and after S. aureus ATCC25923 infection. Significant variations (at P > 0.05) were observed in the weight of study animals during the study. Period C C C C Week 0 22.00b 23.57b 19.34e 23.24a 20.90d 22.84c 20.19d 22.60b 23.92a 23.70b 20.56c 23.78a 20.89d 23.10c 20.00d 23.51a Average 22.43 23.30 20.02 23.28 SD 1.29 0.40 0.51 0.51 C TST TSA TSTSA 24.92a 24.15a 21.19c 20.98c Week 1 23.34a 23.13c 20.26d 22.68b 21.98b 22.96d 20.85c 22.86b 22.60b 22.36d 24.28a 22.33b Average 23.21 23.15 21.65 22.21 SD 1.27 0.74 1.80 0.86 23.25a 24.28a 19.94d 20.39d Week 2 21.32c 22.09e 18.46f 20.35d 20.29d 21.82e 19.40e 16.68e 21.27c 22.89d 22.74b 20.09d Average 21.53 22.77 20.14 19.38 SD 1.24 1.10 1.84 1.80 Values with the same alphabet along the same column are not significantly different (P > 0.05) Composition of control and trial diets are given in Table 1. -

Letters to Nature
letters to nature Received 7 July; accepted 21 September 1998. 26. Tronrud, D. E. Conjugate-direction minimization: an improved method for the re®nement of macromolecules. Acta Crystallogr. A 48, 912±916 (1992). 1. Dalbey, R. E., Lively, M. O., Bron, S. & van Dijl, J. M. The chemistry and enzymology of the type 1 27. Wolfe, P. B., Wickner, W. & Goodman, J. M. Sequence of the leader peptidase gene of Escherichia coli signal peptidases. Protein Sci. 6, 1129±1138 (1997). and the orientation of leader peptidase in the bacterial envelope. J. Biol. Chem. 258, 12073±12080 2. Kuo, D. W. et al. Escherichia coli leader peptidase: production of an active form lacking a requirement (1983). for detergent and development of peptide substrates. Arch. Biochem. Biophys. 303, 274±280 (1993). 28. Kraulis, P.G. Molscript: a program to produce both detailed and schematic plots of protein structures. 3. Tschantz, W. R. et al. Characterization of a soluble, catalytically active form of Escherichia coli leader J. Appl. Crystallogr. 24, 946±950 (1991). peptidase: requirement of detergent or phospholipid for optimal activity. Biochemistry 34, 3935±3941 29. Nicholls, A., Sharp, K. A. & Honig, B. Protein folding and association: insights from the interfacial and (1995). the thermodynamic properties of hydrocarbons. Proteins Struct. Funct. Genet. 11, 281±296 (1991). 4. Allsop, A. E. et al.inAnti-Infectives, Recent Advances in Chemistry and Structure-Activity Relationships 30. Meritt, E. A. & Bacon, D. J. Raster3D: photorealistic molecular graphics. Methods Enzymol. 277, 505± (eds Bently, P. H. & O'Hanlon, P. J.) 61±72 (R. Soc. Chem., Cambridge, 1997). -

Coenzymes and Prosthetic Groups Nomenclature
Coenzymes and prosthetic groups Nomenclature • Cofactor: nonprotein component of enzymes • Cofactor - a co-catalyst required for enzyme activity • Coenzyme - a dissociable cofactor, usually organic • Prosthetic group - non-dissociable cofactor • Vitamin - a required micro-nutrient (organism cannot synthesize adequate quantities for normal health - may vary during life-cycle). – water soluble - not stored, generally no problem with overdose – lipid soluble - stored, often toxic with overdose. • Apoenzyme - enzyme lacking cofactor (inactive) • Holoenzyme - enzyme with cofactors (active) Vitamins are precursors of cofactors Why cofactors? Adenine Nucleotide Coenzymes All use the adenine nucleotide group solely for binding to the enzyme! • pyridine dinucleotides (NADH, NADPH) • flavin mono- and dinucleotides (FMN, FADH) • coenzyme A Nucleotide triphosphates • ATP hydrolysis – resonance stabilizes products – reactants cannot be resonance stabilized because of competition with adjacent bridging anhydrides – charge density greater on reactants than products Coenzyme A • Activation of acyl groups for transfer by nucleophilic attack • activation of the alpha- hydrogen of the acyl group for abstraction as a proton • Both these functions are mediated by the reactive -SH group on CoA, which forms thioesters Coenzyme A Nicotinic Acid/Nicotinamide Coenzymes • These coenzymes are two-electron carriers • They transfer hydride anion (H-) to and from substrates • Two important coenzymes in this class: • Nicotinamide adenine dinucleotide (NAD+) • Nicotinamide -

Product Sheet Info
Master Clone List for NR-19279 ® Vibrio cholerae Gateway Clone Set, Recombinant in Escherichia coli, Plates 1-46 Catalog No. NR-19279 Table 1: Vibrio cholerae Gateway® Clones, Plate 1 (NR-19679) Clone ID Well ORF Locus ID Symbol Product Accession Position Length Number 174071 A02 367 VC2271 ribD riboflavin-specific deaminase NP_231902.1 174346 A03 336 VC1877 lpxK tetraacyldisaccharide 4`-kinase NP_231511.1 174354 A04 342 VC0953 holA DNA polymerase III, delta subunit NP_230600.1 174115 A05 388 VC2085 sucC succinyl-CoA synthase, beta subunit NP_231717.1 174310 A06 506 VC2400 murC UDP-N-acetylmuramate--alanine ligase NP_232030.1 174523 A07 132 VC0644 rbfA ribosome-binding factor A NP_230293.2 174632 A08 322 VC0681 ribF riboflavin kinase-FMN adenylyltransferase NP_230330.1 174930 A09 433 VC0720 phoR histidine protein kinase PhoR NP_230369.1 174953 A10 206 VC1178 conserved hypothetical protein NP_230823.1 174976 A11 213 VC2358 hypothetical protein NP_231988.1 174898 A12 369 VC0154 trmA tRNA (uracil-5-)-methyltransferase NP_229811.1 174059 B01 73 VC2098 hypothetical protein NP_231730.1 174075 B02 82 VC0561 rpsP ribosomal protein S16 NP_230212.1 174087 B03 378 VC1843 cydB-1 cytochrome d ubiquinol oxidase, subunit II NP_231477.1 174099 B04 383 VC1798 eha eha protein NP_231433.1 174294 B05 494 VC0763 GTP-binding protein NP_230412.1 174311 B06 314 VC2183 prsA ribose-phosphate pyrophosphokinase NP_231814.1 174603 B07 108 VC0675 thyA thymidylate synthase NP_230324.1 174474 B08 466 VC1297 asnS asparaginyl-tRNA synthetase NP_230942.2 174933 B09 198 -

Supplementary Information
Supplementary information (a) (b) Figure S1. Resistant (a) and sensitive (b) gene scores plotted against subsystems involved in cell regulation. The small circles represent the individual hits and the large circles represent the mean of each subsystem. Each individual score signifies the mean of 12 trials – three biological and four technical. The p-value was calculated as a two-tailed t-test and significance was determined using the Benjamini-Hochberg procedure; false discovery rate was selected to be 0.1. Plots constructed using Pathway Tools, Omics Dashboard. Figure S2. Connectivity map displaying the predicted functional associations between the silver-resistant gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S3. Connectivity map displaying the predicted functional associations between the silver-sensitive gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S4. Metabolic overview of the pathways in Escherichia coli. The pathways involved in silver-resistance are coloured according to respective normalized score. Each individual score represents the mean of 12 trials – three biological and four technical. Amino acid – upward pointing triangle, carbohydrate – square, proteins – diamond, purines – vertical ellipse, cofactor – downward pointing triangle, tRNA – tee, and other – circle. -
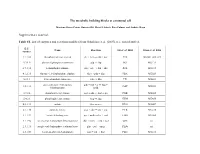
The Metabolic Building Blocks of a Minimal Cell Supplementary
The metabolic building blocks of a minimal cell Mariana Reyes-Prieto, Rosario Gil, Mercè Llabrés, Pere Palmer and Andrés Moya Supplementary material. Table S1. List of enzymes and reactions modified from Gabaldon et. al. (2007). n.i.: non identified. E.C. Name Reaction Gil et. al. 2004 Glass et. al. 2006 number 2.7.1.69 phosphotransferase system glc + pep → g6p + pyr PTS MG041, 069, 429 5.3.1.9 glucose-6-phosphate isomerase g6p ↔ f6p PGI MG111 2.7.1.11 6-phosphofructokinase f6p + atp → fbp + adp PFK MG215 4.1.2.13 fructose-1,6-bisphosphate aldolase fbp ↔ gdp + dhp FBA MG023 5.3.1.1 triose-phosphate isomerase gdp ↔ dhp TPI MG431 glyceraldehyde-3-phosphate gdp + nad + p ↔ bpg + 1.2.1.12 GAP MG301 dehydrogenase nadh 2.7.2.3 phosphoglycerate kinase bpg + adp ↔ 3pg + atp PGK MG300 5.4.2.1 phosphoglycerate mutase 3pg ↔ 2pg GPM MG430 4.2.1.11 enolase 2pg ↔ pep ENO MG407 2.7.1.40 pyruvate kinase pep + adp → pyr + atp PYK MG216 1.1.1.27 lactate dehydrogenase pyr + nadh ↔ lac + nad LDH MG460 1.1.1.94 sn-glycerol-3-phosphate dehydrogenase dhp + nadh → g3p + nad GPS n.i. 2.3.1.15 sn-glycerol-3-phosphate acyltransferase g3p + pal → mag PLSb n.i. 2.3.1.51 1-acyl-sn-glycerol-3-phosphate mag + pal → dag PLSc MG212 acyltransferase 2.7.7.41 phosphatidate cytidyltransferase dag + ctp → cdp-dag + pp CDS MG437 cdp-dag + ser → pser + 2.7.8.8 phosphatidylserine synthase PSS n.i. cmp 4.1.1.65 phosphatidylserine decarboxylase pser → peta PSD n.i. -

The Biosynthesis of the Molybdenum Cofactor in Escherichia Coli and Its Connection to Fes Cluster Assembly and the Thiolation of Trna
Hindawi Publishing Corporation Advances in Biology Volume 2014, Article ID 808569, 21 pages http://dx.doi.org/10.1155/2014/808569 Review Article The Biosynthesis of the Molybdenum Cofactor in Escherichia coli and Its Connection to FeS Cluster Assembly and the Thiolation of tRNA Silke Leimkühler Department of Molecular Enzymology, Institute of Biochemistry and Biology, University of Potsdam, Karl-Liebknecht-Straße 24-25, 14476 Potsdam, Germany Correspondence should be addressed to Silke Leimkuhler;¨ [email protected] Received 16 January 2014; Accepted 28 March 2014; Published 29 April 2014 Academic Editor: Paul Rosch¨ Copyright © 2014 Silke Leimkuhler.¨ This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. The thiolation of biomolecules is a complex process that involves the activation of sulfur. The L-cysteine desulfurase IscS is the main sulfur mobilizing protein in Escherichia coli that provides the sulfur from L-cysteine to several important biomolecules in the cell such as iron sulfur (FeS) clusters, molybdopterin (MPT), thiamine, and thionucleosides of tRNA. Various proteins mediate the transfer of sulfur from IscS to various biomolecules using different interaction partners. A direct connection between the sulfur- containing molecules FeS clusters, thiolated tRNA, and the molybdenum cofactor (Moco) has been identified. The first step of Moco biosynthesis involves the conversion of 5 GTP to cyclic pyranopterin monophosphate (cPMP), a reaction catalyzed by a FeS cluster containing protein. Formed cPMP is further converted to MPT by insertion of two sulfur atoms. The sulfur for this reaction is provided by the L-cysteine desulfurase IscS in addition to the involvement of the TusA protein. -

Program Bound to Inspire
A PROGRAM BOUND TO INSPIRE Held in conjunction with Conference Program Feeling lucky? Stop by the ASBMB booth #1421 for your chance to win one of three $250 Amazon gift cards. To participate in the contest, you must update your ASBMB member profile or join the society for the first time. WWW.ASBMB.ORG Table of Contents 2 At-a-glance 10 Sunday oral program 20 Monday oral program 30 Tuesday oral program 39 Poster sessions Program at-a-glance At-a-glance Saturday April 6 Time Location Event Convention Center 8:30 AM – 4:30 PM ASBMB Graduate Student and Postdoctoral Researcher Travel W307ABC Awardee Career Development Event Convention Center 11:30 AM – 12:00 PM ASBMB Annual Meeting Orientation for Undergraduate W306AB Students Convention Center 11:30 AM – 6:00 PM ASBMB Undergraduate Poster Competition Judges' W303ABC Orientation Convention Center 1:00 PM – 4:30 PM W304 ASBMB Undergraduate Student Research Poster Competition Convention Center 1:15 PM – 2:45 PM Career Development Workshop for Grads and Postdocs: W305A Networking Skills Convention Center 1:15 PM – 2:45 PM Career Development Workshop for Grads and Postdocs: W307D Constructing Your Elevator Pitch Convention Center 1:15 PM – 2:45 PM Career Development Workshop for Grads and Postdocs: W305B Practical Tools for Navigating Your Career Path Evolution Convention Center 4:45 PM – 5:45 PM ASBMB Undergraduate Workshop: Exploring Careers Speed W306AB Networking Convention Center 7:00 PM – 8:30 PM Valencia Ballroom EB Welcome Reception with Science Outreach Poster Session ABCD GRANT WRITING WORKSHOP June 13–15 • Washington, D.C. -

Supplementary Informations SI2. Supplementary Table 1
Supplementary Informations SI2. Supplementary Table 1. M9, soil, and rhizosphere media composition. LB in Compound Name Exchange Reaction LB in soil LBin M9 rhizosphere H2O EX_cpd00001_e0 -15 -15 -10 O2 EX_cpd00007_e0 -15 -15 -10 Phosphate EX_cpd00009_e0 -15 -15 -10 CO2 EX_cpd00011_e0 -15 -15 0 Ammonia EX_cpd00013_e0 -7.5 -7.5 -10 L-glutamate EX_cpd00023_e0 0 -0.0283302 0 D-glucose EX_cpd00027_e0 -0.61972444 -0.04098397 0 Mn2 EX_cpd00030_e0 -15 -15 -10 Glycine EX_cpd00033_e0 -0.0068175 -0.00693094 0 Zn2 EX_cpd00034_e0 -15 -15 -10 L-alanine EX_cpd00035_e0 -0.02780553 -0.00823049 0 Succinate EX_cpd00036_e0 -0.0056245 -0.12240603 0 L-lysine EX_cpd00039_e0 0 -10 0 L-aspartate EX_cpd00041_e0 0 -0.03205557 0 Sulfate EX_cpd00048_e0 -15 -15 -10 L-arginine EX_cpd00051_e0 -0.0068175 -0.00948672 0 L-serine EX_cpd00054_e0 0 -0.01004986 0 Cu2+ EX_cpd00058_e0 -15 -15 -10 Ca2+ EX_cpd00063_e0 -15 -100 -10 L-ornithine EX_cpd00064_e0 -0.0068175 -0.00831712 0 H+ EX_cpd00067_e0 -15 -15 -10 L-tyrosine EX_cpd00069_e0 -0.0068175 -0.00233919 0 Sucrose EX_cpd00076_e0 0 -0.02049199 0 L-cysteine EX_cpd00084_e0 -0.0068175 0 0 Cl- EX_cpd00099_e0 -15 -15 -10 Glycerol EX_cpd00100_e0 0 0 -10 Biotin EX_cpd00104_e0 -15 -15 0 D-ribose EX_cpd00105_e0 -0.01862144 0 0 L-leucine EX_cpd00107_e0 -0.03596182 -0.00303228 0 D-galactose EX_cpd00108_e0 -0.25290619 -0.18317325 0 L-histidine EX_cpd00119_e0 -0.0068175 -0.00506825 0 L-proline EX_cpd00129_e0 -0.01102953 0 0 L-malate EX_cpd00130_e0 -0.03649016 -0.79413596 0 D-mannose EX_cpd00138_e0 -0.2540567 -0.05436649 0 Co2 EX_cpd00149_e0 -

Vitamin Biosynthesis As an Antifungal Target
Journal of Fungi Review Vitamin Biosynthesis as an Antifungal Target Zohar Meir and Nir Osherov * Department of Clinical Microbiology and Immunology, Sackler School of Medicine, Tel-Aviv University, Ramat-Aviv, Tel-Aviv 69978, Israel; [email protected] * Correspondence: [email protected]; Tel.: +972-3-640-9599; Fax: +972-3-640-9160 Received: 29 May 2018; Accepted: 13 June 2018; Published: 17 June 2018 Abstract: The large increase in the population of immunosuppressed patients, coupled with the limited efficacy of existing antifungals and rising resistance toward them, have dramatically highlighted the need to develop novel drugs for the treatment of invasive fungal infections. An attractive possibility is the identification of possible drug targets within essential fungal metabolic pathways not shared with humans. Here, we review the vitamin biosynthetic pathways (vitamins A–E, K) as candidates for the development of antifungals. We present a set of ranking criteria that identify the vitamin B2 (riboflavin), B5 (pantothenic acid), and B9 (folate) biosynthesis pathways as being particularly rich in new antifungal targets. We propose that recent scientific advances in the fields of drug design and fungal genomics have developed sufficiently to merit a renewed look at these pathways as promising sources for the development of novel classes of antifungals. Keywords: antifungals; fungal vitamin metabolism; drug target; essential genes 1. Introduction The number of life-threatening fungal infections has risen dramatically over the last twenty years. Recent estimates have identified a global burden of almost two million patients with systemic and invasive fungal infections, including ~700,000 cases of invasive candidiasis, ~500,000 cases of Pneumocystis jirovecii pneumonia, ~250,000 cases of invasive aspergillosis, ~220,000 cases of cryptococcal meningitis, and ~100,000 cases of disseminated histoplasmosis [1,2].