Supplementary Information
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Hypopituitarism May Be an Additional Feature of SIM1 and POU3F2 Containing 6Q16 Deletions in Children with Early Onset Obesity
Open Access Journal of Pediatric Endocrinology Case Report Hypopituitarism may be an Additional Feature of SIM1 and POU3F2 Containing 6q16 Deletions in Children with Early Onset Obesity Rutteman B1, De Rademaeker M2, Gies I1, Van den Bogaert A2, Zeevaert R1, Vanbesien J1 and De Abstract Schepper J1* Over the past decades, 6q16 deletions have become recognized as a 1Department of Pediatric Endocrinology, UZ Brussel, frequent cause of the Prader-Willi-like syndrome. Involvement of SIM1 in the Belgium deletion has been linked to the development of obesity. Although SIM1 together 2Department of Genetics, UZ Brussel, Belgium with POU3F2, which is located close to the SIM1 locus, are involved in pituitary *Corresponding author: De Schepper J, Department development and function, pituitary dysfunction has not been reported frequently of Pediatric Endocrinology, UZ Brussel, Belgium in cases of 6q16.1q16.3 deletion involving both genes. Here we report on a case of a girl with typical Prader-Willi-like symptoms including early-onset Received: December 23, 2016; Accepted: February 21, hyperphagic obesity, hypotonia, short hands and feet and neuro-psychomotor 2017; Published: February 22, 2017 development delay. Furthermore, she suffered from central diabetes insipidus, central hypothyroidism and hypocortisolism. Her genetic defect is a 6q16.1q16.3 deletion, including SIM1 and POU3F2. We recommend searching for a 6q16 deletion in children with early onset hyperphagic obesity associated with biological signs of hypopituitarism and/or polyuria-polydipsia syndrome. The finding of a combined SIM1 and POU3F2 deletion should prompt monitoring for the development of hypopituitarism, if not already present at diagnosis. Keywords: 6q16 deletion; SIM1; POU3F2; Prader-Willi-like syndrome; Hypopituitarism Introduction insipidus and of some specific pituitary hormone deficiencies in children with a combined SIM1 and POU3F2 deficiency. -

Download Download
Supplementary Figure S1. Results of flow cytometry analysis, performed to estimate CD34 positivity, after immunomagnetic separation in two different experiments. As monoclonal antibody for labeling the sample, the fluorescein isothiocyanate (FITC)- conjugated mouse anti-human CD34 MoAb (Mylteni) was used. Briefly, cell samples were incubated in the presence of the indicated MoAbs, at the proper dilution, in PBS containing 5% FCS and 1% Fc receptor (FcR) blocking reagent (Miltenyi) for 30 min at 4 C. Cells were then washed twice, resuspended with PBS and analyzed by a Coulter Epics XL (Coulter Electronics Inc., Hialeah, FL, USA) flow cytometer. only use Non-commercial 1 Supplementary Table S1. Complete list of the datasets used in this study and their sources. GEO Total samples Geo selected GEO accession of used Platform Reference series in series samples samples GSM142565 GSM142566 GSM142567 GSM142568 GSE6146 HG-U133A 14 8 - GSM142569 GSM142571 GSM142572 GSM142574 GSM51391 GSM51392 GSE2666 HG-U133A 36 4 1 GSM51393 GSM51394 only GSM321583 GSE12803 HG-U133A 20 3 GSM321584 2 GSM321585 use Promyelocytes_1 Promyelocytes_2 Promyelocytes_3 Promyelocytes_4 HG-U133A 8 8 3 GSE64282 Promyelocytes_5 Promyelocytes_6 Promyelocytes_7 Promyelocytes_8 Non-commercial 2 Supplementary Table S2. Chromosomal regions up-regulated in CD34+ samples as identified by the LAP procedure with the two-class statistics coded in the PREDA R package and an FDR threshold of 0.5. Functional enrichment analysis has been performed using DAVID (http://david.abcc.ncifcrf.gov/) -

S41467-020-18249-3.Pdf
ARTICLE https://doi.org/10.1038/s41467-020-18249-3 OPEN Pharmacologically reversible zonation-dependent endothelial cell transcriptomic changes with neurodegenerative disease associations in the aged brain Lei Zhao1,2,17, Zhongqi Li 1,2,17, Joaquim S. L. Vong2,3,17, Xinyi Chen1,2, Hei-Ming Lai1,2,4,5,6, Leo Y. C. Yan1,2, Junzhe Huang1,2, Samuel K. H. Sy1,2,7, Xiaoyu Tian 8, Yu Huang 8, Ho Yin Edwin Chan5,9, Hon-Cheong So6,8, ✉ ✉ Wai-Lung Ng 10, Yamei Tang11, Wei-Jye Lin12,13, Vincent C. T. Mok1,5,6,14,15 &HoKo 1,2,4,5,6,8,14,16 1234567890():,; The molecular signatures of cells in the brain have been revealed in unprecedented detail, yet the ageing-associated genome-wide expression changes that may contribute to neurovas- cular dysfunction in neurodegenerative diseases remain elusive. Here, we report zonation- dependent transcriptomic changes in aged mouse brain endothelial cells (ECs), which pro- minently implicate altered immune/cytokine signaling in ECs of all vascular segments, and functional changes impacting the blood–brain barrier (BBB) and glucose/energy metabolism especially in capillary ECs (capECs). An overrepresentation of Alzheimer disease (AD) GWAS genes is evident among the human orthologs of the differentially expressed genes of aged capECs, while comparative analysis revealed a subset of concordantly downregulated, functionally important genes in human AD brains. Treatment with exenatide, a glucagon-like peptide-1 receptor agonist, strongly reverses aged mouse brain EC transcriptomic changes and BBB leakage, with associated attenuation of microglial priming. We thus revealed tran- scriptomic alterations underlying brain EC ageing that are complex yet pharmacologically reversible. -

Methods in and Applications of the Sequencing of Short Non-Coding Rnas" (2013)
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations 2013 Methods in and Applications of the Sequencing of Short Non- Coding RNAs Paul Ryvkin University of Pennsylvania, [email protected] Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Bioinformatics Commons, Genetics Commons, and the Molecular Biology Commons Recommended Citation Ryvkin, Paul, "Methods in and Applications of the Sequencing of Short Non-Coding RNAs" (2013). Publicly Accessible Penn Dissertations. 922. https://repository.upenn.edu/edissertations/922 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/922 For more information, please contact [email protected]. Methods in and Applications of the Sequencing of Short Non-Coding RNAs Abstract Short non-coding RNAs are important for all domains of life. With the advent of modern molecular biology their applicability to medicine has become apparent in settings ranging from diagonistic biomarkers to therapeutics and fields angingr from oncology to neurology. In addition, a critical, recent technological development is high-throughput sequencing of nucleic acids. The convergence of modern biotechnology with developments in RNA biology presents opportunities in both basic research and medical settings. Here I present two novel methods for leveraging high-throughput sequencing in the study of short non- coding RNAs, as well as a study in which they are applied to Alzheimer's Disease (AD). The computational methods presented here include High-throughput Annotation of Modified Ribonucleotides (HAMR), which enables researchers to detect post-transcriptional covalent modifications ot RNAs in a high-throughput manner. In addition, I describe Classification of RNAs by Analysis of Length (CoRAL), a computational method that allows researchers to characterize the pathways responsible for short non-coding RNA biogenesis. -

Human Cumulus Cells in Long-Term in Vitro Culture Reflect Differential
cells Article Human Cumulus Cells in Long-Term In Vitro Culture Reflect Differential Expression Profile of Genes Responsible for Planned Cell Death and Aging—A Study of New Molecular Markers Bła˙zejChermuła 1, Wiesława Kranc 2 , Karol Jopek 3, Joanna Budna-Tukan 3 , Greg Hutchings 2,4, Claudia Dompe 3,4, Lisa Moncrieff 3,4 , Krzysztof Janowicz 2,4, Małgorzata Józkowiak 5, Michal Jeseta 6 , Jim Petitte 7 , Paul Mozdziak 8 , Leszek Pawelczyk 1, Robert Z. Spaczy ´nski 1 and Bartosz Kempisty 2,3,6,9,* 1 Division of Infertility and Reproductive Endocrinology, Department of Gynecology, Obstetrics and Gynecological Oncology, Poznan University of Medical Sciences, 33 Polna St., 60-535 Poznan, Poland; [email protected] (B.C.); [email protected] (L.P.); [email protected] (R.Z.S.) 2 Department of Anatomy, Poznan University of Medical Sciences, 6 Swiecickiego St., 60-781 Poznan, Poland; [email protected] (W.K.); [email protected] (G.H.); [email protected] (K.J.) 3 Department of Histology and Embryology, Poznan University of Medical Sciences, 6 Swiecickiego St., 60-781 Poznan, Poland; [email protected] (K.J.); [email protected] (J.B.-T.); [email protected] (C.D.); l.moncrieff[email protected] (L.M.) 4 The School of Medicine, Medical Sciences and Nutrition, University of Aberdeen, Aberdeen AB25 2ZD, UK 5 Department of Toxicology, Poznan University of Medical Sciences, 30 Dojazd St., 60-631 Poznan, Poland; [email protected] 6 Department of Obstetrics and Gynecology, University Hospital and -

The Interaction of DNA Repair Factors ASCC2 and ASCC3 Is Affected by Somatic Cancer Mutations
ARTICLE https://doi.org/10.1038/s41467-020-19221-x OPEN The interaction of DNA repair factors ASCC2 and ASCC3 is affected by somatic cancer mutations Junqiao Jia 1, Eva Absmeier1,5, Nicole Holton1, Agnieszka J. Pietrzyk-Brzezinska1,6, Philipp Hackert2, ✉ Katherine E. Bohnsack 2, Markus T. Bohnsack2,3 & Markus C. Wahl 1,4 The ASCC3 subunit of the activating signal co-integrator complex is a dual-cassette Ski2-like nucleic acid helicase that provides single-stranded DNA for alkylation damage repair by the 1234567890():,; α-ketoglutarate-dependent dioxygenase AlkBH3. Other ASCC components integrate ASCC3/ AlkBH3 into a complex DNA repair pathway. We mapped and structurally analyzed inter- acting ASCC2 and ASCC3 regions. The ASCC3 fragment comprises a central helical domain and terminal, extended arms that clasp the compact ASCC2 unit. ASCC2–ASCC3 interfaces are evolutionarily highly conserved and comprise a large number of residues affected by somatic cancer mutations. We quantified contributions of protein regions to the ASCC2–ASCC3 interaction, observing that changes found in cancers lead to reduced ASCC2–ASCC3 affinity. Functional dissection of ASCC3 revealed similar organization and regulation as in the spliceosomal RNA helicase Brr2. Our results delineate functional regions in an important DNA repair complex and suggest possible molecular disease principles. 1 Laboratory of Structural Biochemistry, Freie Universität Berlin, D-14195 Berlin, Germany. 2 Department of Molecular Biology, University Medical Centre Göttingen, Göttingen, Germany. 3 Göttingen Center for Molecular Biosciences, Georg-August-Universität, Göttingen, Germany. 4 Helmholtz-Zentrum Berlin für Materialien und Energie, Macromolecular Crystallography, D-12489 Berlin, Germany. 5Present address: MRC Laboratory of Molecular Biology, Cambridge Biomedical Campus, Cambridge CB2 0QH, UK. -

Cerebral Cavernous Malformations: from Genes to Proteins to Disease
See the corresponding editorial in this issue, pp 119–121. J Neurosurg 116:122–132, 2012 Cerebral cavernous malformations: from genes to proteins to disease Clinical article DANIEL D. CAVALCANTI, M.D., M. YASHAR S. KALANI, M.D., PH.D., NIKOLAY L. MARTIROSYAN, M.D., JUSTIN EALES, B.S., ROBERT F. SPETZLER, M.D., AND MARK C. PREUL, M.D. Division of Neurological Surgery, Barrow Neurological Institute, St. Joseph’s Hospital and Medical Center, Phoenix, Arizona Over the past half century molecular biology has led to great advances in our understanding of angio- and vas- culogenesis and in the treatment of malformations resulting from these processes gone awry. Given their sporadic and familial distribution, their developmental and pathological link to capillary telangiectasias, and their observed chromosomal abnormalities, cerebral cavernous malformations (CCMs) are regarded as akin to cancerous growths. Although the exact pathological mechanisms involved in the formation of CCMs are still not well understood, the identification of 3 genetic loci has begun to shed light on key developmental pathways involved in CCM pathogen- esis. Cavernous malformations can occur sporadically or in an autosomal dominant fashion. Familial forms of CCMs have been attributed to mutations at 3 different loci implicated in regulating important processes such as proliferation and differentiation of angiogenic precursors and members of the apoptotic machinery. These processes are important for the generation, maintenance, and pruning of every vessel in the body. In this review the authors highlight the lat- est discoveries pertaining to the molecular genetics of CCMs, highlighting potential new therapeutic targets for the treatment of these lesions. -
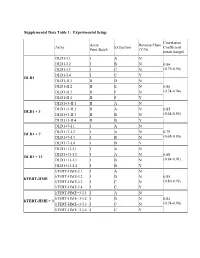
Table 3: Average Gene Expression Profiles by Chromosome
Supplemental Data Table 1: Experimental Setup Correlation Array Reverse Fluor Array Extraction Coefficient Print Batch (Y/N) mean (range) DLD1-I.1 I A N DLD1-I.2 I B N 0.86 DLD1-I.3 I C N (0.79-0.90) DLD1-I.4 I C Y DLD1 DLD1-II.1 II D N DLD1-II.2 II E N 0.86 DLD1-II.3 II F N (0.74-0.94) DLD1-II.4 II F Y DLD1+3-II.1 II A N DLD1+3-II.2 II A N 0.85 DLD1 + 3 DLD1+3-II.3 II B N (0.64-0.95) DLD1+3-II.4 II B Y DLD1+7-I.1 I A N DLD1+7-I.2 I A N 0.79 DLD1 + 7 DLD1+7-I.3 I B N (0.68-0.90) DLD1+7-I.4 I B Y DLD1+13-I.1 I A N DLD1+13-I.2 I A N 0.88 DLD1 + 13 DLD1+13-I.3 I B N (0.84-0.91) DLD1+13-I.4 I B Y hTERT-HME-I.1 I A N hTERT-HME-I.2 I B N 0.85 hTERT-HME hTERT-HME-I.3 I C N (0.80-0.92) hTERT-HME-I.4 I C Y hTERT-HME+3-I.1 I A N hTERT-HME+3-I.2 I B N 0.84 hTERT-HME + 3 hTERT-HME+3-I.3 I C N (0.74-0.90) hTERT-HME+3-I.4 I C Y Supplemental Data Table 2: Average gene expression profiles by chromosome arm DLD1 hTERT-HME Ratio.7 Ratio.1 Ratio.3 Ratio.3 Chrom. -

Multiomic Analysis of the UV-Induced DNA Damage Response
Resource Multiomic Analysis of the UV-Induced DNA Damage Response Graphical Abstract Authors Stefan Boeing, Laura Williamson, Vesela Encheva, ..., Michael Howell, Ambrosius P. Snijders, Jesper Q. Svejstrup Correspondence [email protected] In Brief Boeing et al. investigate the UV-induced DNA damage response by combining a range of proteomic and genomic screens. A function in this response for the melanoma driver STK19 as well as a number of other factors are uncovered. Highlights d A multiomic screening approach examines the UV-induced DNA damage response d Multiple factors are connected to the transcription-related DNA damage response d Melanoma gene STK19 is required for a normal DNA damage response Boeing et al., 2016, Cell Reports 15, 1597–1610 May 17, 2016 ª 2016 The Author(s) http://dx.doi.org/10.1016/j.celrep.2016.04.047 Cell Reports Resource Multiomic Analysis of the UV-Induced DNA Damage Response Stefan Boeing,1,5 Laura Williamson,1 Vesela Encheva,2 Ilaria Gori,3 Rebecca E. Saunders,3 Rachael Instrell,3 Ozan Aygun,€ 1,7 Marta Rodriguez-Martinez,1 Juston C. Weems,4 Gavin P. Kelly,5 Joan W. Conaway,4,6 Ronald C. Conaway,4,6 Aengus Stewart,5 Michael Howell,3 Ambrosius P. Snijders,2 and Jesper Q. Svejstrup1,* 1Mechanisms of Transcription Laboratory, the Francis Crick Institute, Clare Hall Laboratories, South Mimms EN6 3LD, UK 2Protein Analysis and Proteomics Laboratory, the Francis Crick Institute, Clare Hall Laboratories, South Mimms EN6 3LD, UK 3High Throughput Screening Laboratory, the Francis Crick Institute, 44 Lincoln’s -

Context-Selective Death of Acute Myeloid Leukemia Cells Triggered by the Novel Hybrid Retinoid-HDAC Inhibitor MC2392
Published OnlineFirst February 24, 2014; DOI: 10.1158/0008-5472.CAN-13-2568 Cancer Therapeutics, Targets, and Chemical Biology Research Context-Selective Death of Acute Myeloid Leukemia Cells Triggered by the Novel Hybrid Retinoid-HDAC Inhibitor MC2392 Floriana De Bellis1,4, Vincenzo Carafa1, Mariarosaria Conte1, Dante Rotili3, Francesca Petraglia1, Filomena Matarese4, Kees-Jan Francoijs¸ 4, Julien Ablain6, Sergio Valente3,Remy Castellano5, Armelle Goubard5, Yves Collette5, Amit Mandoli4, Joost H.A. Martens4, Hugues de The6, Angela Nebbioso1, Antonello Mai3, Hendrik G. Stunnenberg4, and Lucia Altucci1,2 Abstract HDAC inhibitors (HDACi) are widely used in the clinic to sensitize tumorigenic cells for treatment with other anticancer compounds. The major drawback of HDACi is the broad inhibition of the plethora of HDAC-containing complexes. In acute promyelocytic leukemia (APL), repression by the PML-RARa oncofusion protein is mediated by an HDAC-containing complex that can be dissociated by pharmacologic doses of all trans retinoic acid (ATRA) inducing differentiation and cell death at the expense of side effects and recurrence. We hypothesized that the context-specific close physical proximity of a retinoid and HDACi-binding protein in the repressive PML-RARa- HDAC complex may permit selective targeting by a hybrid molecule of ATRA with a 2-aminoanilide tail of the HDAC inhibitor MS-275, yielding MC2392. We show that MC2392 elicits weak ATRA and essentially no HDACi activity in vitro or in vivo. Genome-wide epigenetic analyses revealed that in NB4 cells expressing PML-RARa, MC2392 induces changes in H3 acetylation at a small subset of PML-RARa–binding sites. RNA-seq reveals that MC2392 alters expression of a number of stress-responsive and apoptotic genes. -

Gene Section Review
Atlas of Genetics and Cytogenetics in Oncology and Haematology OPEN ACCESS JOURNAL INIST-CNRS Gene Section Review DAPK2 (death-associated protein kinase 2) Mafalda Pinto, Valdemar Máximo IPATIMUP Institute of Molecular Pathology and Immunology of the University of Porto, (MP, VM); I3S Institute for Innovation and Health Research, University of Porto (MP, VM); Department of Pathology and Oncology, Medical Faculty of the University of Porto, Porto, Portugal (VM) [email protected]; [email protected] Published in Atlas Database: April 2016 Online updated version : http://AtlasGeneticsOncology.org/Genes/DAPK2ID40263ch15q22.html Printable original version : http://documents.irevues.inist.fr/bitstream/handle/2042/66944/04-2016-DAPK2ID40263ch15q22.pdf DOI: 10.4267/2042/66944 This work is licensed under a Creative Commons Attribution-Noncommercial-No Derivative Works 2.0 France Licence. © 2016 Atlas of Genetics and Cytogenetics in Oncology and Haematology Location (base pair): Chromosome 15:63,907,036- Abstract 64,072,033 reverse strand. GRCh38:CM000677.2 (Ensembl.org) Short communication on DAPK2, with data on DNA and on the protein encoded. DNA/RNA Keywords DAPK2; DRP1; DRP-1; calcium/calmodulin; DAPK2 is a gene that codes for a protein that serine/threonine; kinase; apoptosis belongs to the serine/threonine protein kinase family. Identity Transcription Other names: DRP1, DRP-1 DPAK2 has 13 transcripts (3 coding), 75 orthologues and 11 paralogues (MYLK4, MYLK3, STK17A, HGNC (Hugo): DAPK2 DAPK3, DAPK1, OBSCN, SPEG, TTN, MYLK, Location: 15q22.31 STK17B, MYKL2) (ENSG00000035664). Blue highlighting indicates alternating exons; Red highlighting indicates amino acids encoded across a splice junction. Atlas Genet Cytogenet Oncol Haematol. 2016; 20(12) 587 DAPK2 (death-associated protein kinase 2) Pinto M, Máximo V Schematic diagram of DAPK2 protein structure. -

Application of Next-Generation Sequencing for the Genomic
Article Application of Next‐Generation Sequencing for the Genomic Characterization of Patients with Smoldering Myeloma Martina Manzoni, Valentina Marchica, Paola Storti, Bachisio Ziccheddu, Gabriella Sammarelli, Giannalisa Todaro, Francesca Pelizzoni, Simone Salerio, Laura Notarfranchi, Alessandra Pompa, Luca Baldini, Niccolò Bolli, Antonino Neri, Nicola Giuliani and Marta Lionetti Supplementary Materials Supplementary Methods Library Design, CAPP‐Seq Library Preparation, and Ultra‐Deep Next‐Generation Sequencing (NGS) The targeted resequencing gene panel included coding exons and splice sites of 56 genes that emerged as drivers from a recent analysis of whole genome/exome data in more than 800 MM patients [1] (target region: 112 kb: ACTG1, BCL7A, BHLHE41, BRAF, BTG1, CCND1, CDKN1B, CYLD, DIS3, DTX1, DUSP2, EGR1, FAM46C, FGFR3, FUBP1, HIST1H1B, HIST1H1D, HIST1H1E, HIST1H2BK, IGLL5, IRF1, IRF4, KLHL6, KMT2B, KRAS, LCE1D, LTB, MAX, NFKB2, NFKBIA, NRAS, PABPC1, PIM1, POT1, PRDM1, PRKD2, PTPN11, RASA2, RB1, RFTN1, RPL10, RPL5, RPRD1B, RPS3A, SAMHD1, SETD2, SP140, TBC1D29, TCL1A, TGDS, TP53, TRAF2, TRAF3, XBP1, ZNF462, ZNF292). Tumor gDNA (median 400 ng) was sheared through enzymatic fragmentation before library construction to obtain 150–200‐bp fragments. Targeted ultra‐deep‐next generation sequencing was performed by using the CAPP‐seq approach, as described in Newman A.M. et al., Nat. Med. 2014 [2]. The NGS libraries were constructed using the KAPA Library Preparation Kit (Kapa Biosystems, Wilmington, MA, USA) and hybrid selection was performed with the custom SeqCap EZ Choice Library (Roche NimbleGen, Madison, WI, USA), allowing for enrichment by the capture of genomic regions of interest up to 7 Mb for human resequencing studies. Multiplexed libraries were sequenced using 200‐bp paired‐end runs on an Illumina MiSeq sequencer (Illumina, Hayward, CA, USA).