6 Molecular Systematics of Red Algae
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Xenacoelomorpha's Significance for Understanding Bilaterian Evolution
Available online at www.sciencedirect.com ScienceDirect Xenacoelomorpha’s significance for understanding bilaterian evolution Andreas Hejnol and Kevin Pang The Xenacoelomorpha, with its phylogenetic position as sister biology models are the fruitfly Drosophila melanogaster and group of the Nephrozoa (Protostomia + Deuterostomia), plays the nematode Caenorhabditis elegans, in which basic prin- a key-role in understanding the evolution of bilaterian cell types ciples of developmental processes have been studied in and organ systems. Current studies of the morphological and great detail. It might be because the field of evolutionary developmental diversity of this group allow us to trace the developmental biology — EvoDevo — has its origin in evolution of different organ systems within the group and to developmental biology and not evolutionary biology that reconstruct characters of the most recent common ancestor of species under investigation are often called ‘model spe- Xenacoelomorpha. The disparity of the clade shows that there cies’. Criteria for selected representative species are cannot be a single xenacoelomorph ‘model’ species and primarily the ease of access to collected material and strategic sampling is essential for understanding the evolution their ability to be cultivated in the lab [1]. In some cases, of major traits. With this strategy, fundamental insights into the a supposedly larger number of ancestral characters or a evolution of molecular mechanisms and their role in shaping dominant role in ecosystems have played an additional animal organ systems can be expected in the near future. role in selecting model species. These arguments were Address used to attract sufficient funding for genome sequencing Sars International Centre for Marine Molecular Biology, University of and developmental studies that are cost-intensive inves- Bergen, Thormøhlensgate 55, 5008 Bergen, Norway tigations. -

Carbonic Anhydrases: an Ancient Tool in Calcareous Sponge Biomineralization
fgene-12-624533 March 30, 2021 Time: 13:30 # 1 BRIEF RESEARCH REPORT published: 07 April 2021 doi: 10.3389/fgene.2021.624533 Carbonic Anhydrases: An Ancient Tool in Calcareous Sponge Biomineralization Oliver Voigt1*, Benedetta Fradusco1, Carolin Gut1, Charalampos Kevrekidis1, Sergio Vargas1 and Gert Wörheide1,2,3 1 Department of Earth and Environmental Sciences, Palaeontology and Geobiology, Ludwig-Maximilians-Universität München, Munich, Germany, 2 GeoBio-Center, Ludwig-Maximilians-Universität München, Munich, Germany, 3 SNSB-Bayerische Staatssammlung für Paläontologie und Geologie, Munich, Germany Enzymes of the a-carbonic anhydrase gene family (CAs) are essential for the deposition of calcium carbonate biominerals. In calcareous sponges (phylum Porifera, class Calcarea), specific CAs are involved in the formation of calcite spicules, a unique trait and synapomorphy of this class. However, detailed studies on the CA repertoire of calcareous sponges exist for only two species of one of the two Calcarea subclasses, the Calcaronea. The CA repertoire of the second subclass, the Calcinea, has not been investigated so far, leaving a considerable gap in our knowledge about this Edited by: Melanie Debiais-Thibaud, gene family in Calcarea. Here, using transcriptomic analysis, phylogenetics, and in situ Université de Montpellier, France hybridization, we study the CA repertoire of four additional species of calcareous Reviewed by: sponges, including three from the previously unsampled subclass Calcinea. Our data Helena Cetkovi´ c,´ Rudjer Boskovic Institute, Croatia indicate that the last common ancestor of Calcarea had four ancestral CAs with defined Ana Riesgo, subcellular localizations and functions (mitochondrial/cytosolic, membrane-bound, and Natural History Museum, secreted non-catalytic). The evolution of membrane-bound and secreted CAs involved United Kingdom gene duplications and losses, whereas mitochondrial/cytosolic and non-catalytic CAs *Correspondence: Oliver Voigt are evidently orthologous genes. -
S I Section 4
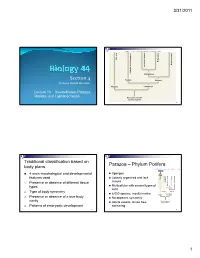
3/31/2011 Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Porifera Ecdysozoa Deuterostomia Lophotrochozoa Cnidaria and Ctenophora Cnidaria and Protostomia SSiection 4 Radiata Bilateria Professor Donald McFarlane Parazoa Eumetazoa Lecture 13 Invertebrates: Parazoa, Radiata, and Lophotrochozoa Ancestral colonial choanoflagellate 2 Traditional classification based on Parazoa – Phylum Porifera body plans Copyright © The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Parazoa 4 main morphological and developmental Sponges features used Loosely organized and lack Porifera Ecdysozoa Cnidaria and Ctenophora tissues euterostomia photrochozoa D 1. o Presence or absence of different tissue L Multicellular with several types of types Protostomia cells 2. Type of body symmetry Radiata Bilateria 8,000 species, mostly marine Parazoa Eumetazoa 3. Presence or absence of a true body No apparent symmetry cavity Ancestral colonial Adults sessile, larvae free- choanoflagellate 4. Patterns of embryonic development swimming 3 4 1 3/31/2011 Water drawn through pores (ostia) into spongocoel Flows out through osculum Reproduce Choanocytes line spongocoel Sexually Most hermapppgggphrodites producing eggs and sperm Trap and eat small particles and plankton Gametes are derived from amoebocytes or Mesohyl between choanocytes and choanocytes epithelial cells Asexually Amoebocytes absorb food from choanocytes, Small fragment or bud may detach and form a new digest it, and carry -

Animal Evolution: Trichoplax, Trees, and Taxonomic Turmoil
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Dispatch R1003 Dispatches Animal Evolution: Trichoplax, Trees, and Taxonomic Turmoil The genome sequence of Trichoplax adhaerens, the founding member of the into the same major classes (C, E/F enigmatic animal phylum Placozoa, has revealed that a surprising level of and B) as do those described from genetic complexity underlies its extremely simple body plan, indicating either Amphimedon [4]. Consistent with that placozoans are secondarily simple or that there is an undiscovered a more derived position, however, morphologically complex life stage. Trichoplax has a number of Antp superclass Hox genes that are absent David J. Miller1 and Eldon E. Ball2 but no other axial differentiation, from the sponge Amphimedon. resembling an amoeba. Grell [3] who These include the ‘ParaHox’ gene With the recent or imminent release formally described these common but Trox-2 [5] and the extended Hox of the whole genome sequences of inconspicuous marine organisms as family gene Not [6] known from a number of key animal species, this belonging to a new phylum, assumed previous work. Particularly intriguing is an exciting time for the ‘evo-devo’ that their simplicity is primary, and is the discovery in Trichoplax of many community. In the last twelve months, that they therefore must represent genes associated with neuroendocrine whole genome analyses of the a key stage in animal evolution. This function across the Bilateria; in cnidarian Nematostella vectensis, view is still held by several prominent common with Amphimedon [7], many the choanoflagellate Monosiga Trichoplax biologists, but has always elements of the post-synaptic scaffold brevicollis and the cephalochordate been contentious; the view that it is are present, but so too are channel Branchiostoma floridae (commonly derived from a more complex ancestor and receptor proteins not known from known as amphioxus) have been has recently been gaining momentum sponges. -

The Evolution of the Mitochondrial Genomes of Calcareous Sponges and Cnidarians Ehsan Kayal Iowa State University
Iowa State University Capstones, Theses and Graduate Theses and Dissertations Dissertations 2012 The evolution of the mitochondrial genomes of calcareous sponges and cnidarians Ehsan Kayal Iowa State University Follow this and additional works at: https://lib.dr.iastate.edu/etd Part of the Evolution Commons, and the Molecular Biology Commons Recommended Citation Kayal, Ehsan, "The ve olution of the mitochondrial genomes of calcareous sponges and cnidarians" (2012). Graduate Theses and Dissertations. 12621. https://lib.dr.iastate.edu/etd/12621 This Dissertation is brought to you for free and open access by the Iowa State University Capstones, Theses and Dissertations at Iowa State University Digital Repository. It has been accepted for inclusion in Graduate Theses and Dissertations by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. The evolution of the mitochondrial genomes of calcareous sponges and cnidarians by Ehsan Kayal A dissertation submitted to the graduate faculty in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY Major: Ecology and Evolutionary Biology Program of Study Committee Dennis V. Lavrov, Major Professor Anne Bronikowski John Downing Eric Henderson Stephan Q. Schneider Jeanne M. Serb Iowa State University Ames, Iowa 2012 Copyright 2012, Ehsan Kayal ii TABLE OF CONTENTS ABSTRACT .......................................................................................................................................... -

Introduction to the Bilateria and the Phylum Xenacoelomorpha Triploblasty and Bilateral Symmetry Provide New Avenues for Animal Radiation
CHAPTER 9 Introduction to the Bilateria and the Phylum Xenacoelomorpha Triploblasty and Bilateral Symmetry Provide New Avenues for Animal Radiation long the evolutionary path from prokaryotes to modern animals, three key innovations led to greatly expanded biological diversification: (1) the evolution of the eukaryote condition, (2) the emergence of the A Metazoa, and (3) the evolution of a third germ layer (triploblasty) and, perhaps simultaneously, bilateral symmetry. We have already discussed the origins of the Eukaryota and the Metazoa, in Chapters 1 and 6, and elsewhere. The invention of a third (middle) germ layer, the true mesoderm, and evolution of a bilateral body plan, opened up vast new avenues for evolutionary expan- sion among animals. We discussed the embryological nature of true mesoderm in Chapter 5, where we learned that the evolution of this inner body layer fa- cilitated greater specialization in tissue formation, including highly specialized organ systems and condensed nervous systems (e.g., central nervous systems). In addition to derivatives of ectoderm (skin and nervous system) and endoderm (gut and its de- Classification of The Animal rivatives), triploblastic animals have mesoder- Kingdom (Metazoa) mal derivatives—which include musculature, the circulatory system, the excretory system, Non-Bilateria* Lophophorata and the somatic portions of the gonads. Bilater- (a.k.a. the diploblasts) PHYLUM PHORONIDA al symmetry gives these animals two axes of po- PHYLUM PORIFERA PHYLUM BRYOZOA larity (anteroposterior and dorsoventral) along PHYLUM PLACOZOA PHYLUM BRACHIOPODA a single body plane that divides the body into PHYLUM CNIDARIA ECDYSOZOA two symmetrically opposed parts—the left and PHYLUM CTENOPHORA Nematoida PHYLUM NEMATODA right sides. -

TRF2 and the Evolution of the Bilateria
Downloaded from genesdev.cshlp.org on September 26, 2021 - Published by Cold Spring Harbor Laboratory Press HYPOTHESIS TRF2 and the evolution of the bilateria Sascha H.C. Duttke,1 Russell F. Doolittle,1,2 Yuan-Liang Wang,1 and James T. Kadonaga1 1Section of Molecular Biology, University of California at San Diego, La Jolla, California 92093, USA; 2Department of Chemistry and Biochemistry, University of California at San Diego, La Jolla, California 92093, USA The development of a complex body plan requires a di- addition, TFIIA has been found to enhance the binding of versity of regulatory networks. Here we consider the TBP to DNA. TBP (as well as the TATA box), TFIIB, TFIIE, concept of TATA-box-binding protein (TBP) family pro- TFIIS, and Pol II are present in Archaea and eukaryotes teins as ‘‘system factors’’ that each supports a distinct set (for review, see Grohmann and Werner 2011). Moreover, of transcriptional programs. For instance, TBP activates many eukaryotes contain TBP as well as TBP-related TATA-box-dependent core promoters, whereas TBP-related factors (TRFs). factor 2 (TRF2) activates TATA-less core promoters that Here we consider the concept that TBP family proteins are dependent on a TCT or downstream core promoter function as ‘‘system factors’’ that support distinct transcrip- element (DPE) motif. These findings led us to investigate tional programs. TBP occurs in Archaea and eukaryotes, the evolution of TRF2. TBP occurs in Archaea and and TBP-related factor 1 (TRF1), TRF2, and TRF3 (for eukaryotes, but TRF2 evolved prior to the emergence reviews, see Goodrich and Tjian 2010; Akhtar and Veenstra of the bilateria and subsequent to the evolutionary split 2011) evolved independently via duplications of the between bilaterians and nonbilaterian animals. -

Lack of Support for Deuterostomia Prompts Reinterpretation of the First Bilateria. Authors and Affiliations Paschalia Kapli1, Pa
bioRxiv preprint doi: https://doi.org/10.1101/2020.07.01.182915; this version posted July 2, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Lack of support for Deuterostomia prompts reinterpretation of the first Bilateria. 2 3 Authors and Affiliations 4 Paschalia Kapli1, Paschalis Natsidis1, Daniel J. Leite1, Maximilian Fursman1,&, Nadia Jeffrie1, 5 Imran A. Rahman2, Hervé Philippe3, Richard R. Copley4, Maximilian J. Telford1* 6 7 1. Centre for Life’s Origins and Evolution, Department of Genetics, Evolution and Environment, 8 University College London, Gower Street, London WC1E 6BT, UK 9 10 2. Oxford University Museum of Natural History, Parks Road, Oxford OX1 3PW, UK 11 3. Station d’Ecologie Théorique et Expérimentale, UMR CNRS 5321, Université Paul Sabatier, 12 09200 Moulis, France 13 4. Laboratoire de Biologie du Développement de Villefranche-sur-mer (LBDV), Sorbonne 14 Université, CNRS, 06230 Villefranche-sur-mer, France 15 & Current Address: Geology and Palaeoenvironmental Research Department, Institute of 16 Geosciences, Goethe University Frankfurt - Riedberg Campus, Altenhöferallee 1, 60438 17 Frankfurt am Main 18 19 20 21 22 23 *Correspondence: [email protected]. 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.07.01.182915; this version posted July 2, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

Cambrian Explosion: the Evolution the Evolution of Animal Body
Cambrian Explosion: The Evolution of Animal Body Plans I. What is a body plan? II. Cambrian Explosion III. Why have no new plans evolved? What is a body plan? (bauplan) At the upper levels of the taxonomic hierarchy, ppyhyla- or class-level clades are characterized by their possession of particular assemblages of homologous architectural and structural features… --J.J. W. Valentine (1986) Characteristics of Body Plans • Symmetry ••SkeletonSkeleton • Body Cavity • Segmentation • Appendages 1 Traditional Phylogeny Four Major Splits: 1) Parazoa-Eumetazoa 2) Radiata-Bilateria 3) Acoelomates-Coelomates 4) Protostomes-Deuterostomes Split 1: Several characters define Eumetazoa. • Tissues separated by basement membranes • Special Cell Junctions (other than septate junctions) • Neurons and Synapses • Muscle Split 2: Body symmetry defines Bilateria. 2 Two Phyla compose the Radiata. Ctenophora Cnidaria Bilateria are worms. Split 3: The presence of a body cavity defines Coelomates. 3 2 cells 4 cells morula blastula Split 4: Developmental characters define Deuterostomes. Traditional Phylogeny Four Major Splits: 1) Parazoa-Eumetazoa 2) Radiata-Bilateria 3) Acoelomates-Coelomates 4) Protostomes-Deuterostomes 4 Burgess Shale 540 mya Cambrian Sponges Cambrian Annelids 5 Cambrian Arthropods Cambrian Brachiopod Cambrian Chordate 6 Traditional Phylogeny Four Major Splits: 1) Parazoa-Eumetazoa 2) Radiata-Bilateria 3) Acoelomates-Coelomates 4) Protostomes-Deuterostomes Simon Conway Morris Opabinia Wiwaxia Hallucigenia 1. What selective factors caused the Cambrian Explosion? 2. Why have no new body plans evolved since? 7 What was so special about the Cambrian? • abiotic environment Atmospheric oxygen was fairly stable during the Cambrian. Cambrian Morris (1995) At the beginning of the Cambrian period, sea level was very similar to that of present day. -

Insights Into Bilaterian Evolution from Three Spiralian Genomes
LETTER OPEN doi:10.1038/nature11696 Insights into bilaterian evolution from three spiralian genomes Oleg Simakov1,2, Ferdinand Marletaz1{, Sung-Jin Cho2, Eric Edsinger-Gonzales2, Paul Havlak3, Uffe Hellsten4, Dian-Han Kuo2{, Tomas Larsson1, Jie Lv3, Detlev Arendt1, Robert Savage5, Kazutoyo Osoegawa6, Pieter de Jong6, Jane Grimwood4,7, Jarrod A. Chapman4, Harris Shapiro4, Andrea Aerts4, Robert P. Otillar4, Astrid Y. Terry4, Jeffrey L. Boore4{, Igor V. Grigoriev4, David R. Lindberg8, Elaine C. Seaver9{, David A. Weisblat2, Nicholas H. Putnam3,10 & Daniel S. Rokhsar2,4,11 Current genomic perspectives on animal diversity neglect two Supplementary Table 2.2.2 and Supplementary Note 2.2. The repetitive prominent phyla, the molluscs and annelids, that together account landscape of these genomes is discussed in Supplementary Note 3.2). for nearly one-third of known marine species and are important Comparing the new genomes with other metazoan sequences, we both ecologically and as experimental systems in classical embry- characterized 8,756 modern bilaterian gene families as likely to have ology1–3. Here we describe the draft genomes of the owl limpet arisen from single progenitor genes in the last common bilaterian (Lottia gigantea), a marine polychaete (Capitella teleta) and a ancestor (Supplementary Note 3.4). As gene loss is common and highly freshwater leech (Helobdella robusta), and compare them with diverged orthologues can be difficult to detect, this is a conservative other animal genomes to investigate the origin and diversifica- lower bound on the number of genes encoded by the last common tion of bilaterians from a genomic perspective. We find that the bilaterian ancestor. -

Animal Relationships Metazoa = Animalia Eumetazoa Bilateria
On the move, and Land Ho! 1. Enter answer text... Animal relationships Metazoa = Animalia Eumetazoa Bilateria Porifera Radiata (sponges) Cnidaria Themes: 1) Mobility 2) Land (again) Kingdom Animalia Multicellular, eukaryotic, and heterotrophic Porifera - Examples of early animals ! No true tissues ! Only a few cell types "Some totipotent ! Filter feeders "Full of holes " Create current with choanocytes See 32.10 #Very similar to choanoflagellates! Animal relationships Metazoa = Animalia Eumetazoa Bilateria Porifera Radiata (sponges) Cnidaria Themes: 1) Mobility 2) Land (again) Kingdom Animalia Multicellular, eukaryotic, and heterotrophic Eumetazoa ! Diploblastic ! Two cell layers "Endoderm (inside) "Ectoderm (outside) ! E.g. Cnidarians "Jellyfish, anemones, corals See Fig. 32.2 Eumetazoa ! Diploblastic ! Two cell layers "Endoderm (inside) "Ectoderm (outside) ! E.g. Cnidarians "Jellyfish, anemones, corals Eumetazoa ! Diploblastic ! Two cell layers "Endoderm (inside) "Ectoderm (outside) ! Gastrulation ! Cnidarians "Jellyfish, anemones, corals Cnidaria - Diploblastic animals ! Have a nervous system! ! Mobility! ! Radially symmetrical "Not very directional See Fig. 32.7 Animal relationships Metazoa = Animalia Eumetazoa Bilateria "Porifera" "Radiata"Cnidaria (sponges) Eumetazoa Gastrulation Nervous system, synpases Radial Symmetry Animal relationships The ctrouble with Ctenophores ! Beautiful, and way cool ! Superficially similar to Cnidarians ! Quite complicated, in some ways like Bilaterians Metazoa = Animalia ! Have been a problem Eumetazoa Bilateria phylogenetically for a long "Porifera" Cnidaria"Radiata" time! (sponges) Eumetazoa Gastrulation Nervous system, synpases Radial Symmetry Animal relationships The ctrouble with Ctenophores ! No buts about it, the butthole is one of the finest innovations in the past 540 million years of animal evolution. The first animals that arose seem to have literally had potty mouths: Their modern-day descendants, such as sea sponges, sea anemones, and jellyfish, all lack an anus and must eat and excrete through the same hole. -

Six Major Steps in Animal Evolution: Are We Derived Sponge Larvae?
EVOLUTION & DEVELOPMENT 10:2, 241–257 (2008) Six major steps in animal evolution: are we derived sponge larvae? Claus Nielsen Zoological Museum (The Natural History Museum of Denmark, University of Copenhagen), Universitetsparken 15, DK-2100 Copenhagen, Denmark Correspondence (email: [email protected]) SUMMARY A review of the old and new literature on animal became sexually mature, and the adult sponge-stage was morphology/embryology and molecular studies has led me to abandoned in an extreme progenesis. This eumetazoan the following scenario for the early evolution of the metazoans. ancestor, ‘‘gastraea,’’ corresponds to Haeckel’s gastraea. The metazoan ancestor, ‘‘choanoblastaea,’’ was a pelagic Trichoplax represents this stage, but with the blastopore spread sphere consisting of choanocytes. The evolution of multicellularity out so that the endoderm has become the underside of the enabled division of labor between cells, and an ‘‘advanced creeping animal. Another lineage developed a nervous system; choanoblastaea’’ consisted of choanocytes and nonfeeding cells. this ‘‘neurogastraea’’ is the ancestor of the Neuralia. Cnidarians Polarity became established, and an adult, sessile stage have retained this organization, whereas the Triploblastica developed. Choanocytes of the upper side became arranged in (Ctenophora1Bilateria), have developed the mesoderm. The a groove with the cilia pumping water along the groove. Cells bilaterians developed bilaterality in a primitive form in the overarched the groove so that a choanocyte chamber was Acoelomorpha and in an advanced form with tubular gut and formed, establishing the body plan of an adult sponge; the pelagic long Hox cluster in the Eubilateria (Protostomia1Deuterostomia). larval stage was retained but became lecithotrophic. The It is indicated that the major evolutionary steps are the result of sponges radiated into monophyletic Silicea, Calcarea, and suites of existing genes becoming co-opted into new networks Homoscleromorpha.