Datasheet for Protein Phosphatase 1 (PP1) (P0754; Lot 0121306)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

ATP-Induced Focal Adhesion Kinase Activity Is Negatively Modulated by Phospholipase D2 in PC12 Cells
EXPERIMENTAL and MOLECULAR MEDICINE, Vol. 33, No. 3, 150-155, September 2001 ATP-induced focal adhesion kinase activity is negatively modulated by phospholipase D2 in PC12 cells Yoe-Sik Bae1 and Sung Ho Ryu1,2 Introduction 1 Division of Molecular and Life Sciences, Pohang University of Purinergic receptors have been reported to play impor- Science and Technology, Pohang 790-784, Korea tant roles on the regulation of neuronal cell functions 2 Corresponding author: Tel, +82-54-279-2292; (Communi et al., 2000; Di Iorio et al., 1998). ATP, a Fax, +82-54-279-2199; E-mail, [email protected] ligand for the receptors modulate various cellular re- sponses such as mitogenic and morphogenic activity in Accepted 18 September 2001 PC12 rat pheochromocytoma cells (Neary et al., 1996; Soltoff et al., 1998; Schindelholz et al., 2000). Stimu- Abbreviations: Fak, focal adhesion kinase; PLD, phospholipase D; lation of cells with ATP induces tyrosine phosphorylation PA, phosphatidic acid; PC, phosphatidylcholine; DAG, diacylglyc- of several cytoskeletal proteins and focal adhesion erol; PBt, phosphatidylbutanol; PKC, protein kinase C; PAP, phos- molecules such as focal adhesion kinase (Fak), proline- phatidic acid phosphohydrolase rich tyrosine kinase (Pyk2), and paxillin (Soltoff et al., 1998; Schindelholz et al., 2000). Since these cytosk- eleton-associated proteins have been regarded as important factors for the regulation of neuronal cell Abstract functions, the study on the regulatory mechanism for the proteins remains an important issue. Extracellular ATP has been known to modulate vari- Phospholipase D (PLD) catalyzes the hydrolysis of ous cellular responses including mitogenesis, secre- phosphatidylcholine (PC) into phosphatidic acid (PA) tion and morphogenic activity in neuronal cells. -

Role of Phospholipases in Adrenal Steroidogenesis
229 1 W B BOLLAG Phospholipases in adrenal 229:1 R29–R41 Review steroidogenesis Role of phospholipases in adrenal steroidogenesis Wendy B Bollag Correspondence should be addressed Charlie Norwood VA Medical Center, One Freedom Way, Augusta, GA, USA to W B Bollag Department of Physiology, Medical College of Georgia, Augusta University (formerly Georgia Regents Email University), Augusta, GA, USA [email protected] Abstract Phospholipases are lipid-metabolizing enzymes that hydrolyze phospholipids. In some Key Words cases, their activity results in remodeling of lipids and/or allows the synthesis of other f adrenal cortex lipids. In other cases, however, and of interest to the topic of adrenal steroidogenesis, f angiotensin phospholipases produce second messengers that modify the function of a cell. In this f intracellular signaling review, the enzymatic reactions, products, and effectors of three phospholipases, f phospholipids phospholipase C, phospholipase D, and phospholipase A2, are discussed. Although f signal transduction much data have been obtained concerning the role of phospholipases C and D in regulating adrenal steroid hormone production, there are still many gaps in our knowledge. Furthermore, little is known about the involvement of phospholipase A2, Endocrinology perhaps, in part, because this enzyme comprises a large family of related enzymes of that are differentially regulated and with different functions. This review presents the evidence supporting the role of each of these phospholipases in steroidogenesis in the Journal Journal of Endocrinology adrenal cortex. (2016) 229, R1–R13 Introduction associated GTP-binding protein exchanges a bound GDP for a GTP. The G protein with GTP bound can then Phospholipids serve a structural function in the cell in that activate the enzyme, phospholipase C (PLC), that cleaves they form the lipid bilayer that maintains cell integrity. -
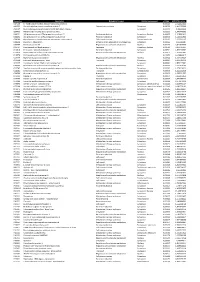
Accession Description Biological Process Cellular Component P-Value SCZ/CTRL Ration A4FU69 EF-Hand Calcium-Binding Domain-Contai
Accession Description Biological Process Cellular component p-Value SCZ/CTRL ration A4FU69 EF-hand calcium-binding domain-containing protein 5 0,001101 2,724427411 A4UGR9 Xin actin-binding repeat-containing protein 2 Cytoskeletal anchoring Cytoplasm 0,006756 1,413953388 A6NCE7 Microtubule-associated proteins 1A/1B light chain 3 beta 2 Cytoplasm 0,001417 1,99612969 B2RPK0 Putative high mobility group protein B1-like 1 0,032314 1,590743352 O00231 26S proteasome non-ATPase regulatory subunit 11 Protein metabolism Cytoplasm; Nucleus 0,000029 1,428664042 O00232 26S proteasome non-ATPase regulatory subunit 12 Protein metabolism Cytoplasm 0,008566 1,544545922 O00264 Membrane-associated progesterone receptor component 1 Cell communication Plasma membrane 0,001459 2,322924147 O00429 Dynamin-1-like protein Mitochondrion organization and biogenesis Cytoplasm 0,006560 0,06391487 O00567 Nucleolar protein 56 Regulation of nucleotide metabolism Nucleus 0,007330 3,842896338 O14737 Programmed cell death protein 5 Apoptosis Cytoplasm; Nucleus 0,006358 3,836727237 O14818 Proteasome subunit alpha type-7 Protein metabolism Cytoplasm 0,030521 1,893928387 O14979 Heterogeneous nuclear ribonucleoprotein D-like Regulation of nucleotide metabolism Nucleus 0,000637 2,150005885 O15078 Centrosomal protein of 290 kDa 0,015359 14,26648619 O15347 High mobility group protein B3 Regulation of nucleotide metabolism Nucleus 0,005500 1,364309014 O15540 Fatty acid-binding protein_ brain Transport Cytoplasm 0,000087 3,125786118 O43237 Cytoplasmic dynein 1 light intermediate -

The Regulatory Roles of Phosphatases in Cancer
Oncogene (2014) 33, 939–953 & 2014 Macmillan Publishers Limited All rights reserved 0950-9232/14 www.nature.com/onc REVIEW The regulatory roles of phosphatases in cancer J Stebbing1, LC Lit1, H Zhang, RS Darrington, O Melaiu, B Rudraraju and G Giamas The relevance of potentially reversible post-translational modifications required for controlling cellular processes in cancer is one of the most thriving arenas of cellular and molecular biology. Any alteration in the balanced equilibrium between kinases and phosphatases may result in development and progression of various diseases, including different types of cancer, though phosphatases are relatively under-studied. Loss of phosphatases such as PTEN (phosphatase and tensin homologue deleted on chromosome 10), a known tumour suppressor, across tumour types lends credence to the development of phosphatidylinositol 3--kinase inhibitors alongside the use of phosphatase expression as a biomarker, though phase 3 trial data are lacking. In this review, we give an updated report on phosphatase dysregulation linked to organ-specific malignancies. Oncogene (2014) 33, 939–953; doi:10.1038/onc.2013.80; published online 18 March 2013 Keywords: cancer; phosphatases; solid tumours GASTROINTESTINAL MALIGNANCIES abs in sera were significantly associated with poor survival in Oesophageal cancer advanced ESCC, suggesting that they may have a clinical utility in Loss of PTEN (phosphatase and tensin homologue deleted on ESCC screening and diagnosis.5 chromosome 10) expression in oesophageal cancer is frequent, Cao et al.6 investigated the role of protein tyrosine phosphatase, among other gene alterations characterizing this disease. Zhou non-receptor type 12 (PTPN12) in ESCC and showed that PTPN12 et al.1 found that overexpression of PTEN suppresses growth and protein expression is higher in normal para-cancerous tissues than induces apoptosis in oesophageal cancer cell lines, through in 20 ESCC tissues. -

Inhibition of Protein Phosphatase 1 Translation by Preventing JNK
Active ERK Contributes to Protein Translation by Preventing JNK-Dependent Inhibition of Protein Phosphatase 1 This information is current as Martha M. Monick, Linda S. Powers, Thomas J. Gross, of September 28, 2021. Dawn M. Flaherty, Christopher W. Barrett and Gary W. Hunninghake J Immunol 2006; 177:1636-1645; ; doi: 10.4049/jimmunol.177.3.1636 http://www.jimmunol.org/content/177/3/1636 Downloaded from References This article cites 59 articles, 43 of which you can access for free at: http://www.jimmunol.org/content/177/3/1636.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication by guest on September 28, 2021 *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2006 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Active ERK Contributes to Protein Translation by Preventing JNK-Dependent Inhibition of Protein Phosphatase 11 Martha M. Monick,2 Linda S. Powers, Thomas J. Gross, Dawn M. Flaherty, Christopher W. -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

Glycogen Degradation
Chapter 12 – Gluconeogenesis, the Pentose Phosphate Pathway and Glycogen Metabolism GLYCOGEN METABOLISM Glycogen stored in muscle and liver cells. Important in maintaining blood glucose levels. Glycogen structure: 1,4 glycosidic linkages with 1,6 branches. Branches give multiple free ends for quicker breakdown or for more places to add additional units. Glycogen Degradation Glucose residues of starch and glycogen released through enzymes called starch phosphorylases and glycogen phosphorylases. Catalyze phosphorolosis: polysaccharide +Pi ---> polysaccharide(n-1) + glucose 1-phosphate Pyridoxal phosphate (PLP) is prosthetic group in active site of enzyme; serves as a proton donor in active site. Allosterically inhibited by high [ATP] and high [glucose 6-phosphate]. Allosterically activated by high [AMP]. Sequentially removes glucose residues from nonreducing ends of glycogen, but stops 4 glucose residues from branch point --> leaves a limit dextran. Limit dextran further degraded by glycogen-debranching enzyme (glucanotransferase activity) which relocated the chain to a free hydroxyl end. Amylo-1,6-glucosidase activity of debranching enzyme removes remaining residues of chain. This leaves substrate for glycogen phosphorylase. Each glucose molecule released from glycogen by debranching enzyme will yield 3 ATPs in glycolysis. Each glucose molecule released by glycogen phosphorylase will yield 2 ATPs in glycolysis. Why? . ATP not needed in first step because glucose 1-phosphate already formed. phosphoglucomutase glucose 1-phosphate ----------------------> glucose 6-phosphate 1) In liver, kidney, pancreas, small intestine, glucose 6-phosphatase glucose 6-phosphate --------------------------> glucose + Pi 2 Glycogen Synthesis Not reverse of glycogen degradation because different enzymes are used. About 2/3 of glucose ingested during a meal is converted to glycogen. -

Adeno-Associated Virus Rep Proteins Antagonize Phosphatase PP1 To
Adeno-associated virus Rep proteins antagonize PNAS PLUS phosphatase PP1 to counteract KAP1 repression of the latent viral genome Sarah Smith-Moorea, Stuart J. D. Neila, Cornel Fraefelb, R. Michael Lindena, Mathieu Bollenc, Helen M. Rowed, and Els Henckaertsa,1 aDepartment of Infectious Diseases, School of Immunology and Microbial Sciences, King’s College London, SE1 9RT London, United Kingdom; bInstitute of Virology, University of Zurich, 8006 Zurich, Switzerland; cDepartment of Cellular and Molecular Medicine, Katholieke Universiteit Leuven, B-3000 Leuven, Belgium; and dDivision of Infection and Immunity, University College London, WC1E 6BT London, United Kingdom Edited by Thomas E. Shenk, Princeton University, Princeton, NJ, and approved March 7, 2018 (received for review December 15, 2017) Adeno-associated virus (AAV) is a small human Dependovirus of various aspects of basic AAV biology. Given that AAV vectors whose low immunogenicity and capacity for long-term persistence likely mimic the latent phase of the viral life cycle, defining the have led to its widespread use as vector for gene therapy. Despite mechanisms involved in the regulation of AAV latency is of great recent successes in AAV-based gene therapy, further im- particular importance for the future design and safety of im- provements in vector technology may be hindered by an inade- proved vectors. In this study, we sought to gain insight into the quate understanding of various aspects of basic AAV biology. AAV regulation of AAV latency by using a screening approach known is unique in that its replication is largely dependent on a helper as “BioID” (17) to identify interaction partners for the AAV2 virus and cellular factors. -

Cell-Specific Bioorthogonal Tagging of Glycoproteins in Co- Culture
Cell-specific Bioorthogonal Tagging of Glycoproteins in Co- culture Supporting Information Supporting Figures Experimentals Anna Ciocea,b, Beatriz Callea,b, Andrea Marchesia,b, Ganka Bineva-Toddb, Helen Flynnc, Zhen Lia,b, Omur Y. Tastanb, Chloe Roustand, Tessa Keenane, Peter Bothf, Kun Huangf, Fabio Parmeggianif, Ambrosius P. Snijdersc, Svend Kjaerd, Martin A. Fascionee, Sabine Flitschf, Benjamin Schumanna,b,* aDepartment of Chemistry, Imperial College London, 80 Wood Lane, W12 0BZ, London, United Kingdom. bThe Chemical Glycobiology Laboratory, The Francis Crick Institute, 1 Midland Rd, NW1 1AT London, United Kingdom. cProteomics Science Technology Platform, The Francis Crick Institute, NW1 1AT London, United Kingdom. dStructural Biology Science Technology Platform, The Francis Crick Institute, NW1 1AT London, United Kingdom. eDepartment of Chemistry, University of York, YO10 5DD York, United Kingdom. fManchester Institute of Biotechnology & School of Chemistry, The University of Manchester, M1 7DN Manchester, United Kingdom. *Correspondence should be addressed to: [email protected]. Supporting Figures Fig. S1: Active site architectures of human enzymes of the GalNAc salvage pathway. In AGX1, the N-acyl side chain in UDP-GalNAc is in proximity to Phe381 and Phe383. In GALK2, the N-acyl side chain of GalNAc-1-phosphate is in proximity with amino acids forming a hydrogen network (Glu179, Ser147 and Ser148). 2 Fig. S2: Evaluation of enzymatic turnover of GalN6yne-Based metabolites. A, in vitro UDP- sugar formation by AGX1 after 2 h incubation, as assessed by LC-MS. Data are means ± SD from three independent experiments. B, in vitro GalN6yne-1-phosphate formation by human and bacterial GalNAc kinases, as assessed by LC-MS and integrated ion count. -

Papadaki Et Al., 2009 Supplementary
Papadaki et al., 2009 Supplementary Supplemental Data Index x Supplemental Figures 1-6 x Supplemental Tables 1a, 1b, 2 Papadaki et al., 2009 Supplementary Supplemental Figure 1. Thymocyte restricted inactivation of the Elavl1 locus. + fl (A) Diagrammatic representation of the wild-type (Elavl1P P), floxed (Elavl1P P) and Cre- - recombined (Elavl1P P) Elavl1/HuR loci on mouse chromosome 8; Noted are the loxP sequences (triangles) flanking the selection marker (neo) used in gene targeting and the ATG containing exon 2 (white box); (H) denotes restriction sites for loci mapping. (B) Detection of native (+), targeted (fl) and Cre-recombinant (-) loci in thymocyte DNA extracts from control and test mice following HindIII digestion and Southern blotting. (C) Western blot of total thymic protein extracts probed with ĮHuR Ab + fl/fl indicating the loss of HuR protein in LckCreP PElavl1P P thymi. Į-actin is shown for quantitation. (D) Flow cytometric detection of intracellular mHuR protein in + fl/+ LckCreP PElavl1P P thymocytes (open histogram), and its respective loss in + fl/fl LckCreP PElavl1P P thymocytes (shaded histogram). The dotted histogram depicts the + isotype-matched background staining. (E) Flow cytometric detection of HuRP P or - + + + fl/+ HuRP P cells in gated splenic CD4P Por CD8P P T-cells from 8 week old LckCreP PElavl1P + fl/fl - Pand LckCreP PElavl1P P mice respectively. (F) Enumeration of HuRP P cells in + fl/fl LckCreP PElavl1P P thymocyte subsets and splenic T-cells; Data are percentages (+SEM) derived from the flow cytometric detection of HuR- cells in CD4/CD8/DP and DN gated populations (n=12-15) at 8-10 weeks of age. -

Phosphatases Page 1
Phosphatases esiRNA ID Gene Name Gene Description Ensembl ID HU-05948-1 ACP1 acid phosphatase 1, soluble ENSG00000143727 HU-01870-1 ACP2 acid phosphatase 2, lysosomal ENSG00000134575 HU-05292-1 ACP5 acid phosphatase 5, tartrate resistant ENSG00000102575 HU-02655-1 ACP6 acid phosphatase 6, lysophosphatidic ENSG00000162836 HU-13465-1 ACPL2 acid phosphatase-like 2 ENSG00000155893 HU-06716-1 ACPP acid phosphatase, prostate ENSG00000014257 HU-15218-1 ACPT acid phosphatase, testicular ENSG00000142513 HU-09496-1 ACYP1 acylphosphatase 1, erythrocyte (common) type ENSG00000119640 HU-04746-1 ALPL alkaline phosphatase, liver ENSG00000162551 HU-14729-1 ALPP alkaline phosphatase, placental ENSG00000163283 HU-14729-1 ALPP alkaline phosphatase, placental ENSG00000163283 HU-14729-1 ALPPL2 alkaline phosphatase, placental-like 2 ENSG00000163286 HU-07767-1 BPGM 2,3-bisphosphoglycerate mutase ENSG00000172331 HU-06476-1 BPNT1 3'(2'), 5'-bisphosphate nucleotidase 1 ENSG00000162813 HU-09086-1 CANT1 calcium activated nucleotidase 1 ENSG00000171302 HU-03115-1 CCDC155 coiled-coil domain containing 155 ENSG00000161609 HU-09022-1 CDC14A CDC14 cell division cycle 14 homolog A (S. cerevisiae) ENSG00000079335 HU-11533-1 CDC14B CDC14 cell division cycle 14 homolog B (S. cerevisiae) ENSG00000081377 HU-06323-1 CDC25A cell division cycle 25 homolog A (S. pombe) ENSG00000164045 HU-07288-1 CDC25B cell division cycle 25 homolog B (S. pombe) ENSG00000101224 HU-06033-1 CDKN3 cyclin-dependent kinase inhibitor 3 ENSG00000100526 HU-02274-1 CTDSP1 CTD (carboxy-terminal domain, -

Supplementary Figures and Table
SUPPLEMENTARY DATA Supplementary Figure 1. ©2014 American Diabetes Association. Published online at http://diabetes.diabetesjournals.org/lookup/suppl/doi:10.2337/db141 -0066/-/DC1 SUPPLEMENTARY DATA Supplementary Figure 2. ©2014 American Diabetes Association. Published online at http://diabetes.diabetesjournals.org/lookup/suppl/doi:10.2337/db142 -0066/-/DC1 SUPPLEMENTARY DATA -/- Supplementary Table 1. Fold increase of Ser/Thr/Tyr phosphorylation in livers of MKP-3 male mice versus wild type male mice fed on a high fat diet (n=5 for each group). Symbol Name Phosphorylation KO/WT ratio Q Value sites Apoptosis ACIN1 Acin1 protein S64 11.4 0.02 T66 8.3 0.02 API5 Apoptosis inhibitor 5 S461 2.2 0.03 S462 1.8 0.03 AIFM3 Apoptosis-inducing factor 3 S30 7.4 0.03 TP53BP2 Apoptosis-stimulating of p53 protein 2 S479 3.7 0.02 ACIN1 Apoptotic chromatin condensation inducer S64S70 5.7 0.02 1 S208 7.1 0.02 S210 7.0 0.02 S479S482S491 105.7 0.03 S729 2.8 0.02 PEA15 Astrocytic phosphoprotein PEA-15 S116 10.8 0.02 BAG3 BAG family molecular chaperone regulator S179 3.3 0.02 3 S353S357 2.3 0.03 S360 2.3 0.03 S390 8.4 0.02 BNIP2 BCL2/adenovirus E1B 19 kDa-interacting S114 3.9 0.02 protein 2 alpha BNIP3 BCL2/adenovirus E1B 19 kDa protein- S60 19.8 0.03 interacting protein 3 S85T86 14.5 0.02 S88 6.1 0.02 BCL2L13 Bcl-2-like protein 13 S387 4.0 0.02 T389 3.1 0.02 CAAP1 Caspase activity and apoptosis inhibitor S183 2.3 0.03 CARD6 Card6 caspase recruitment domain family, S809 3.6 0.03 member 6 CASP8 Caspase-8 S188 2.2 0.02 DAP Death-associated protein S51 5.4 0.02 DAPK2 Death-associated protein kinase 2 S299 3.8 0.02 S349 3.5 0.02 FAF1 FAS-associated factor 1 S269 17.1 0.04 GAS2 Growth arrest-specific protein 2 T282 5.3 0.02 S283 7.4 0.02 S287 5.3 0.02 S289 7.4 0.02 GCH1 GTP cyclohydrolase 1 S24 3.9 0.02 HTT Huntingtin S398S409S411 9.7 0.02 KRT18 Keratin, type I cytoskeletal 18 T9 2.7 0.02 S31S32S35 2.8 0.02 S43S45 3.1 0.02 PDCD5 MCG128907 S119 10.7 0.02 Y126 4.0 0.02 BNIP3I MCG2480, isoform CRA_b S61S62 12.9 0.03 S63S64 8.1 0.02 ©2014 American Diabetes Association.