Publis19-Bgpi-028 Golyaev Plan
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Characterization of P1 Leader Proteases of the Potyviridae Family
Characterization of P1 leader proteases of the Potyviridae family and identification of the host factors involved in their proteolytic activity during viral infection Hongying Shan Ph.D. Dissertation Madrid 2018 UNIVERSIDAD AUTONOMA DE MADRID Facultad de Ciencias Departamento de Biología Molecular Characterization of P1 leader proteases of the Potyviridae family and identification of the host factors involved in their proteolytic activity during viral infection Hongying Shan This thesis is performed in Departamento de Genética Molecular de Plantas of Centro Nacional de Biotecnología (CNB-CSIC) under the supervision of Dr. Juan Antonio García and Dr. Bernardo Rodamilans Ramos Madrid 2018 Acknowledgements First of all, I want to express my appreciation to thesis supervisors Bernardo Rodamilans and Juan Antonio García, who gave the dedicated guidance to this thesis. I also want to say thanks to Carmen Simón-Mateo, Fabio Pasin, Raquel Piqueras, Beatriz García, Mingmin, Zhengnan, Wenli, Linlin, Ruiqiang, Runhong and Yuwei, who helped me and provided interesting suggestions for the thesis as well as technical support. Thanks to the people in the greenhouse (Tomás Heras, Alejandro Barrasa and Esperanza Parrilla), in vitro plant culture facility (María Luisa Peinado and Beatriz Casal), advanced light microscopy (Sylvia Gutiérrez and Ana Oña), photography service (Inés Poveda) and proteomics facility (Sergio Ciordia and María Carmen Mena). Thanks a lot to all the assistance from lab313 colleagues. Thanks a lot to the whole CNB. Thanks a lot to the Chinese Scholarship Council. Thanks a lot to all my friends. Thanks a lot to my family. Madrid 20/03/2018 Index I CONTENTS Abbreviations………………………………………….……………………….……...VII Viruses cited…………………………………………………………………..……...XIII Summary…………………………………………………………………...….…….XVII Resumen…………………………………………………………......…...…………..XXI I. -

Nucleotide Sequence and Phylogenetic Analysis of a Bamboo Mosaic Potexvirus Isolate from Common Bamboo (Bambusa Vulgaris Mcclure)
YangBot. Bull. et al. Acad. Nucleotide Sin. (1997) sequence 38: 77-84 of BaMV-V RNA 77 Nucleotide sequence and phylogenetic analysis of a bamboo mosaic potexvirus isolate from common bamboo (Bambusa vulgaris McClure) Chi-Chen Yang1,2, Jih-Shiou Liu1, Chan-Pin Lin2 and Na-Sheng Lin1,3 1Institute of Botany, Academia Sinica, Taipei, Taiwan 115, Republic of China 2Department of Plant Pathology and Entomology, National Taiwan University, Taipei, Taiwan, Republic of China (Received September 20, 1996; Accepted January 30, 1997) Abstract. The complete cDNA sequence corresponding to the genomic RNA of an isolate bamboo mosaic potexvirus (BaMV-V) from common bamboo (Bambusa vulgaris McClure) was determined. This isolate is the first potexvirus with which a satellite RNA has been associated. The genome organization of BaMV-V, similar to those of other potexviruses, contained five open reading frames (ORFs 15) coding for polypeptides with molecular weight of 156 kDa, 28 kDa, 13 kDa, 6 kDa, and 25 kDa, respectively. Nucleotide sequence analysis showed a 10.0% difference from the BaMV-O isolate previously described whereas the amino acid comparison showed a difference of 3.2%. When three conservative domains of RNA dependent RNA polymerase (RdRp) were used for phylogenetic analysis, the greatest variation between two strains of each virus was only 12.8% of that between the two closest members of the potexvirus group. The grouping of potexviruses distinct from other groups of plant viruses was also confirmed by a comparision of three conservative motifs of RdRp. Keywords: Bamboo mosaic virus; Nucleotide sequence; Potexvirus. Introduction virus M, PVM) (Zavriev et al., 1991), hordeivirus (e.g. -

Tamada-Text R3 HK-TT
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Okayama University Scientific Achievement Repository Tamada & Kondo - 1 Journal of General Plant Pathology Biological and genetic diversity of plasmodiophorid-transmitted viruses and their vectors Tetsuo Tamada* and Hideki Kondo Institute of Plant Science and Resources (IPSR), Okayama University, Kurashiki 710-0046, Japan. Current address for T. Tamada: Kita 10, Nishi 1, 13-2-606, Sapporo, 001-0010, Japan. *Corresponding author: Tetsuo Tamada; E-mail: [email protected] Total text pages 32 Word counts: 10 words (title); 147 words (Abstract); 6004 words (Main Text) Tables: 3; Figure: 1; Supplementary figure: 1 Tamada & Kondo - 2 Abstract About 20 species of viruses belonging to five genera, Benyvirus, Furovirus, Pecluvirus, Pomovirus and Bymovirus, are known to be transmitted by plasmodiophorids. These viruses have all positive-sense single-stranded RNA genomes that consist of two to five RNA components. Three species of plasmodiophorids are recognized as vectors: Polymyxa graminis, P. betae, and Spongospora subterranea. The viruses can survive in soil within the long-lived resting spores of the vector. There are biological and genetic variations in both virus and vector species. Many of the viruses have become the causal agents of important diseases in major crops, such as rice, wheat, barley, rye, sugar beet, potato, and groundnut. Control measure is dependent on the development of the resistant cultivars. During the last half a century, several virus diseases have been rapidly spread and distributed worldwide. For the six major virus diseases, their geographical distribution, diversity, and genetic resistance are addressed. -

The Development and Characterisation of Grapevine Virus-Based
The development and characterisation of grapevine virus-based expression vectors by Jacques du Preez Presented in partial fulfilment of the requirements for the degree Doctor of Philosophy at the Department of Genetics, Stellenbosch University March 2010 Supervisors: Prof JT Burger and Dr DE Goszczynski Study leader: Dr D Stephan Declaration I, the undersigned, hereby declare that the work contained in this thesis is my own original work and that I have not previously in its entirety or in part submitted it at any university for a degree. ______________________ Date: _______________ Jacques du Preez Copyright © 2010 Stellenbosch University All rights reserved ii Abstract Grapevine ( Vitis vinifera L.) is a very important agricultural commodity that needs to be protected. To achieve this several in vivo tools are needed for the study of this crop and the pathogens that infect it. Recently the grapevine genome has been sequenced and the next important step will be gene annotation and function using these in vivo tools. In this study the use of Grapevine virus A (GVA), genus Vitivirus , family Flexiviridae , as transient expression and VIGS vector for heterologous protein expression and functional genomics in Nicotiana benthamiana and V . vinifera were evaluated. Full-length genomic sequences of three South African variants of the virus (GTR11, GTG111 and GTR12) were generated and used in a molecular sequence comparison study. Results confirmed the separation of GVA variants into three groups, with group III (mild variants) being the most distantly related. It showed the high molecular heterogeneity of the virus and that ORF 2 was the most diverse. The GVA variants GTG111, GTR12 and GTR11 were placed in molecular groups I, II and III respectively. -

Disruption of Virus Movement Confers Broad-Spectrum Resistance Against
Proc. Nati. Acad. Sci. USA Vol. 91, pp. 10310-10314, October 1994 Plant Biology Disruption of virus movement confers broad-spectrum resistance against systemic infection by plant viruses with a triple gene block (trnenic plant/doinat negative mutaton/ n l movement proein) DAVID L. BECK, CRAIG J. VAN DOLLEWEERD, TONY J. LOUGH, EZEQUIEL BALMORI, DAVIN M. VOOT, MARK T. ANDERSEN, IONA E. W. O'BRIEN, AND RICHARD L. S. FORSTERt Molecular Genetics Group, The Horticultural and Food Research Institute of New Zealand Ltd., Private Bag 92169, Auckland, New Zealand Communicated by George Bruening, June 23, 1994 ABSTRACT White clover mosaic virus strain 0 (WCIMV- tially difficult. New forms ofresistance active against several 0), species of the Potexvirus genus, contains a set of three different viruses or groups of viruses are being sought. One partially overlapping genes (the triple gene block) that encodes such approach involved the introduction of the gene coding nonvirion proteins of 26 kDa, 13 kDa, and 7 kDa. These for rat 2'-5' oligoadenylate synthetase into the genome of proteins are necesy for cell-to-cell movement in plants but potato plants (4). Genetically engineering transgenic plants to not for replication. The WCIMV-O 13-kDa gene was mutated block virus movement, mimicking the mechanism of some (to 13*) in a region of the gene that is conserved in all viruses natural resistance genes, has been proposed (1, 5, 6) but known to possess triple-gene-block proteins. All 10 13* trans- remains mostly unexploited. genic lines of Nicodiana benthamiana designed to express the The movement function of plant viruses can be comple- mutated movement protein were shown to be resistant to mented by another, frequently unrelated, virus in double systemic infection by WCIMV-O at 1 jug ofWCIMV virions per infections (7). -
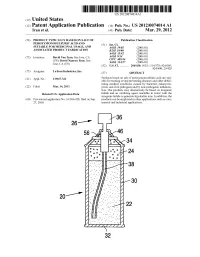
Us 2012/0074.014 A1 2 .. 1
US 2012O074O14A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2012/0074.014 A1 Tran et al. (43) Pub. Date: Mar. 29, 2012 (54) PRODUCT TYPICALLY BASED ON SALT OF Publication Classification PEROXYMONOSULFURIC ACID AND (51) Int. Cl SUITABLE FOR MEDICINAL USAGE, AND iBio/02 (2006.01) ASSOCATED PRODUCT FABRICATION B23P 19/00 (2006.01) A633/42 (2006.01) (75) Inventors: David Van Tran, San Jose, CA A6IR 9/14 (2006.01) (US); David Nguyen Tran, San CD7C 409/244 (2006.01) Jose, CA (US) A6II 3/327 (2006.01) s (52) U.S. Cl. ............. 206/438: 562/1; 514/578; 424/605; 424/400; 29/428 (73) Assignee: LuTran Industries, Inc. (57) ABSTRACT 21) Appl. No.: 13AO47,742 Products based on salt of peroxyperoxVmonosulfuric acid are suit (21) Appl. No 9 able for treating or/and preventing diseases and other debili tating medical conditions caused by bacterial, eukaryotic, (22) Filed: Mar. 14, 2011 prion, and viral pathogens and by non-pathogenic inflamma tion. The products may alternatively be based on inorganic Related U.S. Application Data halide and an oxidizing agent reactable in water with the inorganic halide to generate hypohalite ions. In addition, the (60) Provisional application No. 61/386,928, filed on Sep. products can be employed in other applications such as com 27, 2010. mercial and industrial applications. 5T. 2 .. 2. 1.4. J., Patent Application Publication Mar. 29, 2012 Sheet 1 of 2 US 2012/0074014 A1 cro Q-D 22 Fig. 2a Fig.2b Patent Application Publication Mar. 29, 2012 Sheet 2 of 2 US 2012/0074014 A1 90 92 US 2012/0074014 A1 Mar. -

Evidence to Support Safe Return to Clinical Practice by Oral Health Professionals in Canada During the COVID-19 Pandemic: a Repo
Evidence to support safe return to clinical practice by oral health professionals in Canada during the COVID-19 pandemic: A report prepared for the Office of the Chief Dental Officer of Canada. November 2020 update This evidence synthesis was prepared for the Office of the Chief Dental Officer, based on a comprehensive review under contract by the following: Paul Allison, Faculty of Dentistry, McGill University Raphael Freitas de Souza, Faculty of Dentistry, McGill University Lilian Aboud, Faculty of Dentistry, McGill University Martin Morris, Library, McGill University November 30th, 2020 1 Contents Page Introduction 3 Project goal and specific objectives 3 Methods used to identify and include relevant literature 4 Report structure 5 Summary of update report 5 Report results a) Which patients are at greater risk of the consequences of COVID-19 and so 7 consideration should be given to delaying elective in-person oral health care? b) What are the signs and symptoms of COVID-19 that oral health professionals 9 should screen for prior to providing in-person health care? c) What evidence exists to support patient scheduling, waiting and other non- treatment management measures for in-person oral health care? 10 d) What evidence exists to support the use of various forms of personal protective equipment (PPE) while providing in-person oral health care? 13 e) What evidence exists to support the decontamination and re-use of PPE? 15 f) What evidence exists concerning the provision of aerosol-generating 16 procedures (AGP) as part of in-person -

Molecular Analysis of Vanilla Mosaic Virus from the Cook Islands
Molecular Analysis of Vanilla mosaic virus from the Cook Islands Christopher Puli’uvea A thesis submitted to Auckland University of Technology in partial fulfilment of the requirements for the degree of Master of Science (MSc) 2017 School of Science I Abstract Vanilla was first introduced to French Polynesia in 1848 and from 1899-1966 was a major export for French Polynesia who then produced an average of 158 tonnes of cured Vanilla tahitensis beans annually. In 1967, vanilla production declined rapidly to a low of 0.6 tonnes by 1981, which prompted a nation-wide investigation with the aim of restoring vanilla production to its former levels. As a result, a mosaic-inducing virus was discovered infecting V. tahitensis that was distinct from Cymbidium mosaic virus (CyMV) and Odontoglossum ringspot virus (ORSV) but serologically related to dasheen mosaic virus (DsMV). The potyvirus was subsequently named vanilla mosaic virus (VanMV) and was later reported to infect V. tahitensis in the Cook Islands and V. planifolia in Fiji and Vanuatu. Attempts were made to mechanically inoculate VanMV to a number of plants that are susceptible to DsMV, but with no success. Based on a partial sequence analysis, VanMV-FP (French Polynesian isolate) and VanMV-CI (Cook Islands isolate) were later characterised as strains of DsMV exclusively infecting vanilla. Since its discovery, little information is known about how VanMV-CI acquired the ability to exclusively infect vanilla and lose its ability to infect natural hosts of DsMV or vice versa. The aims of this research were to characterise the VanMV genome and attempt to determine the molecular basis for host range specificity of VanMV-CI. -

Viroze Biljaka 2010
VIROZE BILJAKA Ferenc Bagi Stevan Jasnić Dragana Budakov Univerzitet u Novom Sadu, Poljoprivredni fakultet Novi Sad, 2016 EDICIJA OSNOVNI UDŽBENIK Osnivač i izdavač edicije Univerzitet u Novom Sadu, Poljoprivredni fakultet Trg Dositeja Obradovića 8, 21000 Novi Sad Godina osnivanja 1954. Glavni i odgovorni urednik edicije Dr Nedeljko Tica, redovni profesor Dekan Poljoprivrednog fakulteta Članovi komisije za izdavačku delatnost Dr Ljiljana Nešić, vanredni profesor – predsednik Dr Branislav Vlahović, redovni profesor – član Dr Milica Rajić, redovni profesor – član Dr Nada Plavša, vanredni profesor – član Autori dr Ferenc Bagi, vanredni profesor dr Stevan Jasnić, redovni profesor dr Dragana Budakov, docent Glavni i odgovorni urednik Dr Nedeljko Tica, redovni profesor Dekan Poljoprivrednog fakulteta u Novom Sadu Urednik Dr Vera Stojšin, redovni profesor Direktor departmana za fitomedicinu i zaštitu životne sredine Recenzenti Dr Vera Stojšin, redovni profesor, Univerzitet u Novom Sadu, Poljoprivredni fakultet Dr Mira Starović, naučni savetnik, Institut za zaštitu bilja i životnu sredinu, Beograd Grafički dizajn korice Lea Bagi Izdavač Univerzitet u Novom Sadu, Poljoprivredni fakultet, Novi Sad Zabranjeno preštampavanje i fotokopiranje. Sva prava zadržava izdavač. ISBN 978-86-7520-372-8 Štampanje ovog udžbenika odobrilo je Nastavno-naučno veće Poljoprivrednog fakulteta u Novom Sadu na sednici od 11. 07. 2016.godine. Broj odluke 1000/0102-797/9/1 Tiraž: 20 Mesto i godina štampanja: Novi Sad, 2016. CIP - Ʉɚɬɚɥɨɝɢɡɚɰɢʁɚɭɩɭɛɥɢɤɚɰɢʁɢ ȻɢɛɥɢɨɬɟɤɚɆɚɬɢɰɟɫɪɩɫɤɟɇɨɜɢɋɚɞ -

Progress and Prospects of Studies on Polymyxa Graminis and Its Transmitted Cereal Viruses in China*
PROGRESS IN NATURAL SCIENCE V ol .15 , No .6 , June 2005 REVIEW ARTICLE Progress and prospects of studies on Polymyxa graminis and its transmitted cereal viruses in China* CH EN Jianping ** (Virology and Biotechnology Institute , Zhejiang Academy of Agricultural Sciences, Key Laboratory of Plant Virology , Ministry of Agri- culture and Zhejiang Province, Hangzhou 310021, China) Received October 8 , 2004 ;revised October 25 , 2004 Abstract Polymyxa gram inis is a eukaryotic obligate biotrophic parasite of plant roots that belongs to a poorly studied discrete taxonomic unit informally called “ plasmodiophorids” .P .graminis is nonpathogenic , but has the ability to acquire and transmit nine plant viruses w hich belong to genera Bymovirus and Furovirus and cause serious diseases in cereal crop speciesand also result in significant yield reductions in China and elsew here.Genus Bymovirus contains barley yellow mosaic virus (BaYMV), barley mild mosaic virus (BaMMV), w heat yellow mosaic virus (WYMV), w heat spindle streak mosaic virus (WSS MV), and oat mosaic virus (OMV), and genus F urovirus contains soil-borne w heat mosaic virus(S BW MV), oat golden stripe virus(OGSV), and new ly identified Chinese w heat mosaic virus(CWM V)and soil-borne cereal mosaic virus(S BCMV).All these viruses have been sequenced and their w orldw ide distribu- tions have been studied .The viruses are protected by the environment w ithin P .gra minis resting spores that may remain dormant but vi- able for decades(probably until a suitable host plant is encountered).S pontaneous deletion mutants of S BW MV , OGS V and OMV are de- tected , and these deletion mutants are not transmissible by the fungus.The persistent, soil-borne nature of these diseases makes the use of virus-resistant crop varieties cu rren tly the only practical and environmentally friendly means to control them , and a large number of disease resistant germ plasms have been screened . -

(12) Patent Application Publication (10) Pub. No.: US 2009/0069256A1 Smith Et Al
US 20090069256A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2009/0069256A1 Smith et al. (43) Pub. Date: Mar. 12, 2009 (54) ENHANCING PROTEIN EXPRESSION Related U.S. Application Data (60) Provisional application No. 60/576,819, filed on Jun. (76) Inventors: Larry R. Smith, San Diego, CA 4, 2004. (US); Vafa Shahabi, Valley Forge, PA (US); Maninder K. Sidhu, New Publication Classification City, NY (US) (51) Int. Cl. A63L/7088 (2006.01) Correspondence Address: CI2N IS/II (2006.01) HUNTON & WILLIAMS LLP CI2N 15/87 (2006.01) INTELLECTUAL PROPERTY DEPARTMENT A6IP 43/00 (2006.01) 1900 KSTREET, N.W., SUITE 1200 CI2N IS/00 (2006.01) WASHINGTON, DC 20006-1109 (US) (52) U.S. Cl. ...... 514/44; 536/23.1; 536/23.53: 536/23.5; 536/23.6:536/23.7:536/23.72:536/23.74; 435/455; 435/320.1 (21) Appl. No.: 11/628,455 (57) ABSTRACT (22) PCT Fled: Jun. 6, 2005 Modified polynucleotide compositions providing enhanced gene expression and methods for preparing said compositions (86) PCT NO.: PCT/US05/19592 are disclosed. Methods of using the compositions, such as in screening assays, diagnostic tools, kits, etc. and for preven S371 (c)(1), tion and/or treatment of diseases and disorders are also dis (2), (4) Date: Nov. 8, 2007 closed. SWall (3733) DraI(3734) Stul (52) BclI (3512) ASCI (3496) ApaI (3482) BSInI (3370) BglII (457) 2->BGHpolyA BspEI (3271) BspMI (540) BseRI (3241) human IL-15 (BH15) Psp1406I (667) BamHI (3071) “HuigE leader PflMI (3056) kanamycin Eael (768) NsiI (791) (3737 bp) XmnI (806) BpmI (2976)1 AgeI (846) MsiI (2745) BSrFI (846) SnaBI (2722) Eam1105I (920) BSaAI (2722) DraII (1155) Asel (2390) Spel (2383) EcoRV (2291) ClaI (2285) NruI (2254) BSrBI (1965) AfilII (1896) MunI (2209) Patent Application Publication Mar. -

Multipartite Viruses: Organization, Emergence and Evolution
UNIVERSIDAD AUTÓNOMA DE MADRID FACULTAD DE CIENCIAS DEPARTAMENTO DE BIOLOGÍA MOLECULAR MULTIPARTITE VIRUSES: ORGANIZATION, EMERGENCE AND EVOLUTION TESIS DOCTORAL Adriana Lucía Sanz García Madrid, 2019 MULTIPARTITE VIRUSES Organization, emergence and evolution TESIS DOCTORAL Memoria presentada por Adriana Luc´ıa Sanz Garc´ıa Licenciada en Bioqu´ımica por la Universidad Autonoma´ de Madrid Supervisada por Dra. Susanna Manrubia Cuevas Centro Nacional de Biotecnolog´ıa (CSIC) Memoria presentada para optar al grado de Doctor en Biociencias Moleculares Facultad de Ciencias Departamento de Biolog´ıa Molecular Universidad Autonoma´ de Madrid Madrid, 2019 Tesis doctoral Multipartite viruses: Organization, emergence and evolution, 2019, Madrid, Espana. Memoria presentada por Adriana Luc´ıa-Sanz, licenciada en Bioqumica´ y con un master´ en Biof´ısica en la Universidad Autonoma´ de Madrid para optar al grado de doctor en Biociencias Moleculares del departamento de Biolog´ıa Molecular en la facultad de Ciencias de la Universidad Autonoma´ de Madrid Supervisora de tesis: Dr. Susanna Manrubia Cuevas. Investigadora Cient´ıfica en el Centro Nacional de Biotecnolog´ıa (CSIC), C/ Darwin 3, 28049 Madrid, Espana. to the reader CONTENTS Acknowledgments xi Resumen xiii Abstract xv Introduction xvii I.1 What is a virus? xvii I.2 What is a multipartite virus? xix I.3 The multipartite lifecycle xx I.4 Overview of this thesis xxv PART I OBJECTIVES PART II METHODOLOGY 0.5 Database management for constructing the multipartite and segmented datasets 3 0.6 Analytical