Anti-GCLC (Aa 528-637) Polyclonal Antibody (DPAB-DC1367) This Product Is for Research Use Only and Is Not Intended for Diagnostic Use
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Targeting MYCN in Neuroblastoma by BET Bromodomain Inhibition
Published OnlineFirst February 21, 2013; DOI: 10.1158/2159-8290.CD-12-0418 RESEARCH ARTICLE Targeting MYCN in Neuroblastoma by BET Bromodomain Inhibition Alexandre Puissant1,3, Stacey M. Frumm1,3, Gabriela Alexe1,3,5,6, Christopher F. Bassil1,3, Jun Qi2, Yvan H. Chanthery8, Erin A. Nekritz8, Rhamy Zeid2, William Clay Gustafson8, Patricia Greninger7, Matthew J. Garnett10, Ultan McDermott10, Cyril H. Benes7, Andrew L. Kung1,3, William A. Weiss8,9, James E. Bradner2,4, and Kimberly Stegmaier1,3,6 Downloaded from cancerdiscovery.aacrjournals.org on October 2, 2021. © 2013 American Association for Cancer Research. 15-CD-12-0418_p308-323.indd 1 22/02/13 12:15 AM Published OnlineFirst February 21, 2013; DOI: 10.1158/2159-8290.CD-12-0418 A BSTRACT Bromodomain inhibition comprises a promising therapeutic strategy in cancer, particularly for hematologic malignancies. To date, however, genomic biomarkers to direct clinical translation have been lacking. We conducted a cell-based screen of genetically defined cancer cell lines using a prototypical inhibitor of BET bromodomains. Integration of genetic features with chemosensitivity data revealed a robust correlation between MYCN amplification and sensitivity to bromodomain inhibition. We characterized the mechanistic and translational significance of this finding in neuroblastoma, a childhood cancer with frequent amplification of MYCN. Genome-wide expression analysis showed downregulation of the MYCN transcriptional program accompanied by suppression of MYCN transcription. Functionally, bromodomain-mediated inhibition of MYCN impaired growth and induced apoptosis in neuroblastoma. BRD4 knockdown phenocopied these effects, establishing BET bromodomains as transcriptional regulators of MYCN. BET inhibition conferred a significant survival advantage in 3 in vivo neuroblastoma models, providing a compelling rationale for developing BET bro- modomain inhibitors in patients with neuroblastoma. -

The Role of Nuclear Factor-E2-Related Factor 1 in the Oxidative Stress Response in MC3T3-E1 Osteoblastic Cells
Original Endocrinol Metab 2016;31:336-342 http://dx.doi.org/10.3803/EnM.2016.31.2.336 Article pISSN 2093-596X · eISSN 2093-5978 The Role of Nuclear Factor-E2-Related Factor 1 in the Oxidative Stress Response in MC3T3-E1 Osteoblastic Cells So Young Park1, Sung Hoon Kim1, Hyun Koo Yoon1, Chang Hoon Yim1, Sung-Kil Lim2 1Department of Internal Medicine, Cheil General Hospital & Women’s Healthcare Center, Dankook University College of Medicine, Seoul; 2Division of Endocrinology and Metabolism, Department of Internal Medicine, Yonsei University College of Medicine, Seoul, Korea Background: Reactive oxygen species (ROS) and antioxidants are associated with maintenance of cellular function and metabo- lism. Nuclear factor-E2-related factor 1 (NFE2L1, Nrf1) is known to regulate the expression of a number of genes involved in oxidative stress and inflammation. The purpose of this study was to examine the effects of NFE2L1 on the response to oxidative stress in osteoblastic MC3T3-E1 cells. Methods: The murine calvaria-derived MC3T3-E1 cell line was exposed to lipopolysaccharide (LPS) for oxidative stress induc- tion. NFE2L1 effects were evaluated using small interfering RNA (siRNA) for NFE2L1 mRNA. ROS generation and the levels of known antioxidant enzyme genes were assayed. Results: NFE2L1 expression was significantly increased 2.4-fold compared to the control group at 10 μg/mL LPS in MC3T3-E1 cells (P<0.05). LPS increased formation of intracellular ROS in MC3T3-E1 cells. NFE2L1 knockdown led to an additional in- crease of ROS (20%) in the group transfected with NFE2L1 siRNA compared with the control group under LPS stimulation (P<0.05). -

University of Cincinnati
UNIVERSITY OF CINCINNATI _____________ , 20 _____ I,______________________________________________, hereby submit this as part of the requirements for the degree of: ________________________________________________ in: ________________________________________________ It is entitled: ________________________________________________ ________________________________________________ ________________________________________________ ________________________________________________ Approved by: ________________________ ________________________ ________________________ ________________________ ________________________ Role of Glutamate-Cysteine Ligase in Maintaining Glutathione Homeostasis and Protecting against Oxidative Stress A dissertation submitted to the Division of Research and Advanced Studies University of Cincinnati in partial fulfillment of the requirements for the degree of DOCTORATE OF PHILOSOPHY (Ph.D.) in the Department of Environmental Health of the College of Medicine 2003 by Yi Yang M.D., Sun Yat-sen University of Medical Sciences, 1995 M.S., Sun Yat-sen University of Medical Sciences, 1997 Committee Chair: Daniel W. Nebert, M.D. Professor Department of Environmental Health University of Cincinnati Abstract Glutamate-cysteine ligase (GCL) is the rate-limiting enzyme catalyzing the first step of glutathione (GSH) biosynthesis. In higher eukaryotes, this enzyme is a heterodimer comprising a catalytic subunit (GCLC) and a modifier subunit (GCLM); the latter changes the catalytic characteristics of the holoenzyme. In the first part -

Distinct Isoforms of Nrf1 Diversely Regulate Different Subsets of Its Cognate Target Genes
bioRxiv preprint doi: https://doi.org/10.1101/356071; this version posted June 28, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Distinct isoforms of Nrf1 diversely regulate different subsets of its cognate target genes Meng Wang, Lu Qiu, Xufang Ru, Yijiang Song, Yiguo Zhang* The Laboratory of Cell Biochemistry and Topogenetic Regulation, College of Bioengineering and Faculty of Sciences, Chongqing University, No. 174 Shazheng Street, Shapingba District, Chongqing 400044, China *Correspondence should be addressed to Yiguo Zhang (email: [email protected] or [email protected]). Abstract The single Nrf1 gene has capability to be differentially transcripted alongside with alternative mRNA-splicing and subsequent translation through different initiation signals so as to yield distinct lengths of polypeptide isoforms. Amongst them, three of the most representatives are Nrf1α, Nrf1β and Nrf1γ, but the putative specific contribution of each isoform to regulating ARE-driven target genes remains unknown. To address this, we have here established three cell lines on the base of the Flp-In™ T-REx™ system, which are allowed for tetracycline-inducibly stable expression of Nrf1α, Nrf1β and Nrf1γ. The RNA-Sequencing results have demonstrated that a vast majority of differentially expressed genes (i.e. 90 DEGs detected) were dominantly up-regulated by Nrf1α and/or Nrf1β following induction by tetracycline. By contrast, other DEGs regulated by Nrf1γ were far less than those regulated by Nrf1α/β (i.e. ~11 of Nrf1α and 7 of Nrf1β). Further transcriptomic analysis revealed that tetracycline-induced expression of Nrf1γ significantly increased the percentage of down-regulated genes in total DEGs. -
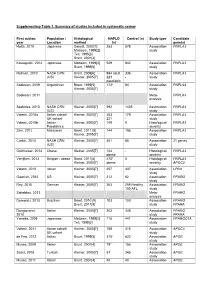
Supplementary Table 1. Summary of Studies Included in Systematic Review
Supplementary Table 1. Summary of studies included in systematic review First author, Population / Histological NAFLD Control (n) Study type Candidate year Location method (n) gene(s) Hotta, 2010 Japanese Sanyal, 2002[1] 253 578 Association PNPLA3 Matteoni, 1999[2] study Teli, 1995[3] Brunt, 2001[4] Kawaguchi, 2012 Japanese Matteoni, 1999[2] 529 942 Association PNPLA3 Brunt, 1999[5] study Rotman, 2010 NASH CRN Brunt, 2009[6] 894 adult 336 Association PNPLA3 (US) Kleiner, 2005[7] 223 - study paediatric Sookoian, 2009 Argentinian Brunt, 1999[5] 172* 94 Association PNPLA3 Kleiner, 2005[7] study Sookoian, 2011 Meta- PNPLA3 analysis Speliotes, 2010 NASH CRN Kleiner, 2005[7] 592 1405 Association PNPLA3 (US) study Valenti, 2010a Italian cohort/ Kleiner, 2005[7] 253 179 Association PNPLA3 UK cohort 321 - study Valenti, 2010b Italian - Kleiner, 2005[7] 149 0 Histological PNPLA3 Paediatrics severity Zain, 2012 Malaysian Brunt, 2011 [8] 144 198 Association PNPLA3 Kleiner, 2005[7] study Corbin, 2013 NASH CRN Kleiner, 2005[7] 361 85 Association 21 genes (US) study Guichelaar, 2013 Obese Kleiner, 2005[7] 144 - Histological PNPLA3 obese severity Verrijken, 2013 Belgian - obese Brunt, 2011[8] 470* 0 Histological PNPLA3 Kleiner, 2005[7] obese severity APOC3 Valenti, 2012 Italian Kleiner, 2005[7] 257 337 Association LPIN1 study Gawrieh, 2012 US Kleiner, 2005[7] 212 62 Association PPARG study Rey, 2010 German Kleiner, 2005[7] 263 259 Healthy Association PPARG 100 AFL study Sahebkar, 2013 Meta- PPARG analysis Domenici, 2013 Brazilian Brunt, 2010 [9] 103 103 -

Snps in Genes Coding for ROS Metabolism and Signalling in Association with Docetaxel Clearance
The Pharmacogenomics Journal (2010) 10, 513–523 & 2010 Macmillan Publishers Limited. All rights reserved 1470-269X/10 www.nature.com/tpj ORIGINAL ARTICLE SNPs in genes coding for ROS metabolism and signalling in association with docetaxel clearance H Edvardsen1,2, PF Brunsvig3, The dose of docetaxel is currently calculated based on body surface area 1,4 5 and does not reflect the pharmacokinetic, metabolic potential or genetic H Solvang , A Tsalenko , background of the patients. The influence of genetic variation on the 6 7 A Andersen , A-C Syvanen , clearance of docetaxel was analysed in a two-stage analysis. In step one, 583 Z Yakhini5, A-L Børresen-Dale1,2, single-nucleotide polymorphisms (SNPs) in 203 genes were genotyped on H Olsen6, S Aamdal3 and samples from 24 patients with locally advanced non-small cell lung cancer. 1,2 We found that many of the genes harbour several SNPs associated with VN Kristensen clearance of docetaxel. Most notably these were four SNPs in EGF, three SNPs 1Department of Genetics, Institute of Cancer in PRDX4 and XPC, and two SNPs in GSTA4, TGFBR2, TNFAIP2, BCL2, DPYD Research, Oslo University Hospital Radiumhospitalet, and EGFR. The multiple SNPs per gene suggested the existence of common Oslo, Norway; 2Institute of Clinical Medicine, haplotypes associated with clearance. These were confirmed with detailed 3 University of Oslo, Oslo, Norway; Cancer Clinic, haplotype analysis. On the basis of analysis of variance (ANOVA), quantitative Oslo University Hospital Radiumhospitalet, Oslo, Norway; 4Institute of -

NRF2-Dependent Gene Expression Promotes Ciliogenesis And
www.nature.com/scientificreports OPEN NRF2-dependent gene expression promotes ciliogenesis and Hedgehog signaling Received: 8 April 2019 Ana Martin-Hurtado1,2, Raquel Martin-Morales1,2, Natalia Robledinos-Antón1,2,3, Accepted: 11 September 2019 Ruth Blanco1,2,3, Ines Palacios-Blanco1,2, Isabel Lastres-Becker1,2,3, Antonio Cuadrado1,2,3 & Published: xx xx xxxx Francesc R. Garcia-Gonzalo 1,2 The transcription factor NRF2 is a master regulator of cellular antioxidant and detoxifcation responses, but it also regulates other processes such as autophagy and pluripotency. In human embryonic stem cells (hESCs), NRF2 antagonizes neuroectoderm diferentiation, which only occurs after NRF2 is repressed via a Primary Cilia-Autophagy-NRF2 (PAN) axis. However, the functional connections between NRF2 and primary cilia, microtubule-based plasma membrane protrusions that function as cellular antennae, remain poorly understood. For instance, nothing is known about whether NRF2 afects cilia, or whether cilia regulation of NRF2 extends beyond hESCs. Here, we show that NRF2 and primary cilia reciprocally regulate each other. First, we demonstrate that fbroblasts lacking primary cilia have higher NRF2 activity, which is rescued by autophagy-activating mTOR inhibitors, indicating that the PAN axis also operates in diferentiated cells. Furthermore, NRF2 controls cilia formation and function. NRF2-null cells grow fewer and shorter cilia and display impaired Hedgehog signaling, a cilia- dependent pathway. These defects are not due to increased oxidative stress or ciliophagy, but rather to NRF2 promoting expression of multiple ciliogenic and Hedgehog pathway genes. Among these, we focused on GLI2 and GLI3, the transcription factors controlling Hh pathway output. Both their mRNA and protein levels are reduced in NRF2-null cells, consistent with their gene promoters containing consensus ARE sequences predicted to bind NRF2. -

Global Analysis of Somatic Structural Genomic Alterations and Their Impact on Gene Expression in Diverse Human Cancers
Global analysis of somatic structural genomic alterations and their impact on gene expression in diverse human cancers Babak Alaei-Mahabadia, Joydeep Bhaduryb, Joakim W. Karlssona, Jonas A. Nilssonb, and Erik Larssona,1 aDepartment of Medical Biochemistry and Cell Biology, Institute of Biomedicine, The Sahlgrenska Academy, University of Gothenburg, SE-405 30 Gothenburg, Sweden; and bDepartment of Surgery, Sahlgrenska Cancer Center, Institute of Clinical Sciences, University of Gothenburg, SE-405 30 Gothenburg, Sweden Edited by Mary-Claire King, University of Washington, Seattle, WA, and approved October 21, 2016 (received for review April 19, 2016) Tumor genomes are mosaics of somatic structural variants (SVs) segments (14). Several factors complicate the analysis, in particular that may contribute to the activation of oncogenes or inactivation mappability issues due to repetitive sequence regions (15). Indeed, of tumor suppressors, for example, by altering gene copy number it has become clear that the results produced by different methods amplitude. However, there are multiple other ways in which SVs are not consistent, and some studies have intersected multiple ap- can modulate transcription, but the general impact of such events proaches to provide a presumed high-confidence set of predictions on tumor transcriptional output has not been systematically de- (16, 17). Adding to the challenges is the difficulty of assessing termined. Here we use whole-genome sequencing data to map SVs performance: True positive sets have thus far been obtained across 600 tumors and 18 cancers, and investigate the relationship through simulated genomic sequences (18), but this will not re- between SVs, copy number alterations (CNAs), and mRNA expression. -

Table S1. 103 Ferroptosis-Related Genes Retrieved from the Genecards
Table S1. 103 ferroptosis-related genes retrieved from the GeneCards. Gene Symbol Description Category GPX4 Glutathione Peroxidase 4 Protein Coding AIFM2 Apoptosis Inducing Factor Mitochondria Associated 2 Protein Coding TP53 Tumor Protein P53 Protein Coding ACSL4 Acyl-CoA Synthetase Long Chain Family Member 4 Protein Coding SLC7A11 Solute Carrier Family 7 Member 11 Protein Coding VDAC2 Voltage Dependent Anion Channel 2 Protein Coding VDAC3 Voltage Dependent Anion Channel 3 Protein Coding ATG5 Autophagy Related 5 Protein Coding ATG7 Autophagy Related 7 Protein Coding NCOA4 Nuclear Receptor Coactivator 4 Protein Coding HMOX1 Heme Oxygenase 1 Protein Coding SLC3A2 Solute Carrier Family 3 Member 2 Protein Coding ALOX15 Arachidonate 15-Lipoxygenase Protein Coding BECN1 Beclin 1 Protein Coding PRKAA1 Protein Kinase AMP-Activated Catalytic Subunit Alpha 1 Protein Coding SAT1 Spermidine/Spermine N1-Acetyltransferase 1 Protein Coding NF2 Neurofibromin 2 Protein Coding YAP1 Yes1 Associated Transcriptional Regulator Protein Coding FTH1 Ferritin Heavy Chain 1 Protein Coding TF Transferrin Protein Coding TFRC Transferrin Receptor Protein Coding FTL Ferritin Light Chain Protein Coding CYBB Cytochrome B-245 Beta Chain Protein Coding GSS Glutathione Synthetase Protein Coding CP Ceruloplasmin Protein Coding PRNP Prion Protein Protein Coding SLC11A2 Solute Carrier Family 11 Member 2 Protein Coding SLC40A1 Solute Carrier Family 40 Member 1 Protein Coding STEAP3 STEAP3 Metalloreductase Protein Coding ACSL1 Acyl-CoA Synthetase Long Chain Family Member 1 Protein -

New Genetic Loci Link Adipose and Insulin Biology to Body Fat Distribution
Europe PMC Funders Group Author Manuscript Nature. Author manuscript; available in PMC 2015 August 12. Published in final edited form as: Nature. 2015 February 12; 518(7538): 187–196. doi:10.1038/nature14132. Europe PMC Funders Author Manuscripts New genetic loci link adipose and insulin biology to body fat distribution A full list of authors and affiliations appears at the end of the article. # These authors contributed equally to this work. Abstract Body fat distribution is a heritable trait and a well-established predictor of adverse metabolic outcomes, independent of overall adiposity. To increase our understanding of the genetic basis of body fat distribution and its molecular links to cardiometabolic traits, we conducted genome-wide association meta-analyses of waist and hip circumference-related traits in up to 224,459 individuals. We identified 49 loci (33 new) associated with waist-to-hip ratio adjusted for body mass index (WHRadjBMI) and an additional 19 loci newly associated with related waist and hip circumference measures (P<5×10−8). Twenty of the 49 WHRadjBMI loci showed significant sexual dimorphism, 19 of which displayed a stronger effect in women. The identified loci were enriched for genes expressed in adipose tissue and for putative regulatory elements in adipocytes. Pathway analyses implicated adipogenesis, angiogenesis, transcriptional regulation, and insulin resistance as processes affecting fat distribution, providing insight into potential pathophysiological mechanisms. Europe PMC Funders Author Manuscripts Depot-specific accumulation of fat, particularly in the central abdomen, confers an elevated risk of metabolic and cardiovascular diseases and mortality1. An easily accessible measure of body fat distribution is waist-to-hip ratio (WHR), a comparison of waist and hip circumferences. -

Defining the Functional Targets of Cap'n'collar Transcription Factors NRF1
antioxidants Article Defining the Functional Targets of Cap‘n’collar Transcription Factors NRF1, NRF2, and NRF3 Lara Ibrahim 1,2 , Jaleh Mesgarzadeh 1, Ian Xu 1 , Evan T. Powers 2, R. Luke Wiseman 1 and Michael J. Bollong 2,* 1 Department of Molecular Medicine, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA; [email protected] (L.I.); [email protected] (J.M.); [email protected] (I.X.); [email protected] (R.L.W.) 2 Department of Chemistry, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA; [email protected] * Correspondence: [email protected] Received: 19 September 2020; Accepted: 19 October 2020; Published: 21 October 2020 Abstract: The NRF transcription factors NRF1, NRF2, and NRF3, are a subset of Cap‘n’collar transcriptional regulators which modulate the expression of genes harboring antioxidant-response element (ARE) sequences within their genomic loci. Despite the emerging physiological importance of NRF family members, the repertoire of their genetic targets remains incompletely defined. Here we use RNA-sequencing-based transcriptional profiling and quantitative proteomics to delineate the overlapping and differential genetic programs effected by the three NRF transcription factors. We then create consensus target gene sets regulated by NRF1, NRF2, and NRF3 and define the integrity of these gene sets for probing NRF activity in mammalian cell culture and human tissues. Together, our data provide a quantitative assessment of how NRF family members sculpt proteomes and transcriptomes, providing a framework to understand the critical physiological importance of NRF transcription factors and to establish pharmacologic approaches for therapeutically activating these transcriptional programs in disease. -

Systematic Analysis, Comparison, and Integration of Disease Based Human
Zhang et al. BMC Medical Genomics 2010, 3:1 http://www.biomedcentral.com/1755-8794/3/1 RESEARCH ARTICLE Open Access Systematic analysis, comparison, and integration of disease based human genetic association data and mouse genetic phenotypic information Yonqing Zhang1†, Supriyo De1†, John R Garner1, Kirstin Smith1, S Alex Wang2, Kevin G Becker1* Abstract Background: The genetic contributions to human common disorders and mouse genetic models of disease are complex and often overlapping. In common human diseases, unlike classical Mendelian disorders, genetic factors generally have small effect sizes, are multifactorial, and are highly pleiotropic. Likewise, mouse genetic models of disease often have pleiotropic and overlapping phenotypes. Moreover, phenotypic descriptions in the literature in both human and mouse are often poorly characterized and difficult to compare directly. Methods: In this report, human genetic association results from the literature are summarized with regard to replication, disease phenotype, and gene specific results; and organized in the context of a systematic disease ontology. Similarly summarized mouse genetic disease models are organized within the Mammalian Phenotype ontology. Human and mouse disease and phenotype based gene sets are identified. These disease gene sets are then compared individually and in large groups through dendrogram analysis and hierarchical clustering analysis. Results: Human disease and mouse phenotype gene sets are shown to group into disease and phenotypically relevant groups at both a coarse and fine level based on gene sharing. Conclusion: This analysis provides a systematic and global perspective on the genetics of common human disease as compared to itself and in the context of mouse genetic models of disease.