High-Resolution Structural Genomics Reveals New Therapeutic Vulnerabilities in Glioblastoma
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Nuclear and Mitochondrial Genome Defects in Autisms
UC Irvine UC Irvine Previously Published Works Title Nuclear and mitochondrial genome defects in autisms. Permalink https://escholarship.org/uc/item/8vq3278q Journal Annals of the New York Academy of Sciences, 1151(1) ISSN 0077-8923 Authors Smith, Moyra Spence, M Anne Flodman, Pamela Publication Date 2009 DOI 10.1111/j.1749-6632.2008.03571.x License https://creativecommons.org/licenses/by/4.0/ 4.0 Peer reviewed eScholarship.org Powered by the California Digital Library University of California THE YEAR IN HUMAN AND MEDICAL GENETICS 2009 Nuclear and Mitochondrial Genome Defects in Autisms Moyra Smith, M. Anne Spence, and Pamela Flodman Department of Pediatrics, University of California, Irvine, California In this review we will evaluate evidence that altered gene dosage and structure im- pacts neurodevelopment and neural connectivity through deleterious effects on synap- tic structure and function, and evidence that the latter are key contributors to the risk for autism. We will review information on alterations of structure of mitochondrial DNA and abnormal mitochondrial function in autism and indications that interactions of the nuclear and mitochondrial genomes may play a role in autism pathogenesis. In a final section we will present data derived using Affymetrixtm SNP 6.0 microar- ray analysis of DNA of a number of subjects and parents recruited to our autism spectrum disorders project. We include data on two sets of monozygotic twins. Col- lectively these data provide additional evidence of nuclear and mitochondrial genome imbalance in autism and evidence of specific candidate genes in autism. We present data on dosage changes in genes that map on the X chromosomes and the Y chro- mosome. -

Analysis of Gene Expression Data for Gene Ontology
ANALYSIS OF GENE EXPRESSION DATA FOR GENE ONTOLOGY BASED PROTEIN FUNCTION PREDICTION A Thesis Presented to The Graduate Faculty of The University of Akron In Partial Fulfillment of the Requirements for the Degree Master of Science Robert Daniel Macholan May 2011 ANALYSIS OF GENE EXPRESSION DATA FOR GENE ONTOLOGY BASED PROTEIN FUNCTION PREDICTION Robert Daniel Macholan Thesis Approved: Accepted: _______________________________ _______________________________ Advisor Department Chair Dr. Zhong-Hui Duan Dr. Chien-Chung Chan _______________________________ _______________________________ Committee Member Dean of the College Dr. Chien-Chung Chan Dr. Chand K. Midha _______________________________ _______________________________ Committee Member Dean of the Graduate School Dr. Yingcai Xiao Dr. George R. Newkome _______________________________ Date ii ABSTRACT A tremendous increase in genomic data has encouraged biologists to turn to bioinformatics in order to assist in its interpretation and processing. One of the present challenges that need to be overcome in order to understand this data more completely is the development of a reliable method to accurately predict the function of a protein from its genomic information. This study focuses on developing an effective algorithm for protein function prediction. The algorithm is based on proteins that have similar expression patterns. The similarity of the expression data is determined using a novel measure, the slope matrix. The slope matrix introduces a normalized method for the comparison of expression levels throughout a proteome. The algorithm is tested using real microarray gene expression data. Their functions are characterized using gene ontology annotations. The results of the case study indicate the protein function prediction algorithm developed is comparable to the prediction algorithms that are based on the annotations of homologous proteins. -

Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model
Downloaded from http://www.jimmunol.org/ by guest on September 25, 2021 T + is online at: average * The Journal of Immunology , 34 of which you can access for free at: 2016; 197:1477-1488; Prepublished online 1 July from submission to initial decision 4 weeks from acceptance to publication 2016; doi: 10.4049/jimmunol.1600589 http://www.jimmunol.org/content/197/4/1477 Molecular Profile of Tumor-Specific CD8 Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. Waugh, Sonia M. Leach, Brandon L. Moore, Tullia C. Bruno, Jonathan D. Buhrman and Jill E. Slansky J Immunol cites 95 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2016/07/01/jimmunol.160058 9.DCSupplemental This article http://www.jimmunol.org/content/197/4/1477.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 25, 2021. The Journal of Immunology Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. -
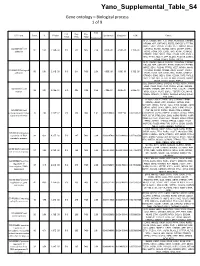
Supplemental Tables4.Pdf
Yano_Supplemental_Table_S4 Gene ontology – Biological process 1 of 9 Fold List Pop Pop GO Term Count % PValue Bonferroni Benjamini FDR Genes Total Hits Total Enrichment DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0007155~cell CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0022610~biological CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DCC, ENAH, PLXNA2, CAPZA2, ATP5B, ASTN1, PAX6, ZEB2, CDH2, CDH4, GLI3, CD24A, EPHB1, NRCAM, GO:0006928~cell CTTNBP2, EDNRB, APP, PTK2, ETV1, CLASP2, STRBP, 36 4.04 3.46E-07 510 205 7436 2.56 7.28E-04 3.64E-04 5.98E-04 motion NRG1, DCLK1, PLAT, SGPL1, TGFBR1, EVL, MYH9, YWHAE, NCKAP1, CTNNA2, SEMA6A, EPHA4, NDEL1, FYN, LRP6 PLXNA2, ADCY5, PAX6, GLI3, CTNNB1, LPHN2, EDNRB, LPHN3, APP, CSNK2A1, GPR45, NRG1, RAPGEF1, WWOX, SGPL1, TLE4, SPEN, NCAM1, DDR1, GRB10, GRM3, GNAQ, HIPK1, GNB1, HIPK2, PYGO1, GO:0007166~cell RNF138, ROR2, CNTN1, -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -
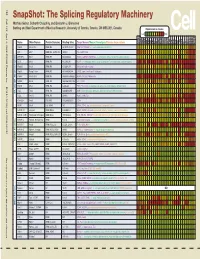
Snapshot: the Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J
192 Cell SnapShot: The Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J. Blencowe 133 Banting and Best Department of Medical Research, University of Toronto, Toronto, ON M5S 3E1, Canada Expression in mouse , April4, 2008©2008Elsevier Inc. Low High Name Other Names Protein Domains Binding Sites Target Genes/Mouse Phenotypes/Disease Associations Amy Ceb Hip Hyp OB Eye SC BM Bo Ht SM Epd Kd Liv Lu Pan Pla Pro Sto Spl Thy Thd Te Ut Ov E6.5 E8.5 E10.5 SRp20 Sfrs3, X16 RRM, RS GCUCCUCUUC SRp20, CT/CGRP; −/− early embryonic lethal E3.5 9G8 Sfrs7 RRM, RS, C2HC Znf (GAC)n Tau, GnRH, 9G8 ASF/SF2 Sfrs1 RRM, RS RGAAGAAC HipK3, CaMKIIδ, HIV RNAs; −/− embryonic lethal, cond. KO cardiomyopathy SC35 Sfrs2 RRM, RS UGCUGUU AChE; −/− embryonic lethal, cond. KO deficient T-cell maturation, cardiomyopathy; LS SRp30c Sfrs9 RRM, RS CUGGAUU Glucocorticoid receptor SRp38 Fusip1, Nssr RRM, RS ACAAAGACAA CREB, type II and type XI collagens SRp40 Sfrs5, HRS RRM, RS AGGAGAAGGGA HipK3, PKCβ-II, Fibronectin SRp55 Sfrs6 RRM, RS GGCAGCACCUG cTnT, CD44 DOI 10.1016/j.cell.2008.03.010 SRp75 Sfrs4 RRM, RS GAAGGA FN1, E1A, CD45; overexpression enhances chondrogenic differentiation Tra2α Tra2a RRM, RS GAAARGARR GnRH; overexpression promotes RA-induced neural differentiation SR and SR-Related Proteins Tra2β Sfrs10 RRM, RS (GAA)n HipK3, SMN, Tau SRm160 Srrm1 RS, PWI AUGAAGAGGA CD44 SWAP Sfrs8 RS, SWAP ND SWAP, CD45, Tau; possible asthma susceptibility gene hnRNP A1 Hnrnpa1 RRM, RGG UAGGGA/U HipK3, SMN2, c-H-ras; rheumatoid arthritis, systemic lupus -

Downloaded, Each with Over 20 Samples for AD-Specific Pathways, Biological Processes, and Driver Each Specific Brain Region in Each Condition
Xiang et al. BMC Medical Genomics 2018, 11(Suppl 6):115 https://doi.org/10.1186/s12920-018-0431-1 RESEARCH Open Access Condition-specific gene co-expression network mining identifies key pathways and regulators in the brain tissue of Alzheimer’s disease patients Shunian Xiang1,2, Zhi Huang4, Wang Tianfu1, Zhi Han3, Christina Y. Yu3,5, Dong Ni1*, Kun Huang3* and Jie Zhang2* From 29th International Conference on Genome Informatics Yunnan, China. 3-5 December 2018 Abstract Background: Gene co-expression network (GCN) mining is a systematic approach to efficiently identify novel disease pathways, predict novel gene functions and search for potential disease biomarkers. However, few studies have systematically identified GCNs in multiple brain transcriptomic data of Alzheimer’s disease (AD) patients and looked for their specific functions. Methods: In this study, we first mined GCN modules from AD and normal brain samples in multiple datasets respectively; then identified gene modules that are specific to AD or normal samples; lastly, condition-specific modules with similar functional enrichments were merged and enriched differentially expressed upstream transcription factors were further examined for the AD/normal-specific modules. Results: We obtained 30 AD-specific modules which showed gain of correlation in AD samples and 31 normal- specific modules with loss of correlation in AD samples compared to normal ones, using the network mining tool lmQCM. Functional and pathway enrichment analysis not only confirmed known gene functional categories related to AD, but also identified novel regulatory factors and pathways. Remarkably, pathway analysis suggested that a variety of viral, bacteria, and parasitic infection pathways are activated in AD samples. -

The Long Non-Coding RNA Gomafu Is Acutely Regulated in Response to Neuronal Activation and Involved in Schizophrenia-Associated Alternative Splicing
Molecular Psychiatry (2014) 19, 486–494 & 2014 Macmillan Publishers Limited All rights reserved 1359-4184/14 www.nature.com/mp ORIGINAL ARTICLE The long non-coding RNA Gomafu is acutely regulated in response to neuronal activation and involved in schizophrenia-associated alternative splicing G Barry1, JA Briggs2, DP Vanichkina1, EM Poth3, NJ Beveridge4,5, VS Ratnu6, SP Nayler2, K Nones7,JHu8, TW Bredy6, S Nakagawa9, F Rigo10, RJ Taft1, MJ Cairns4,5, S Blackshaw3, EJ Wolvetang2 and JS Mattick1,11,12 Schizophrenia (SZ) is a complex disease characterized by impaired neuronal functioning. Although defective alternative splicing has been linked to SZ, the molecular mechanisms responsible are unknown. Additionally, there is limited understanding of the early transcriptomic responses to neuronal activation. Here, we profile these transcriptomic responses and show that long non-coding RNAs (lncRNAs) are dynamically regulated by neuronal activation, including acute downregulation of the lncRNA Gomafu, previously implicated in brain and retinal development. Moreover, we demonstrate that Gomafu binds directly to the splicing factors QKI and SRSF1 (serine/arginine-rich splicing factor 1) and dysregulation of Gomafu leads to alternative splicing patterns that resemble those observed in SZ for the archetypal SZ-associated genes DISC1 and ERBB4. Finally, we show that Gomafu is downregulated in post-mortem cortical gray matter from the superior temporal gyrus in SZ. These results functionally link activity-regulated lncRNAs and alternative splicing in neuronal function and suggest that their dysregulation may contribute to neurological disorders. Molecular Psychiatry (2014) 19, 486–494; doi:10.1038/mp.2013.45; published online 30 April 2013 Keywords: alternative splicing; Gomafu; neuronal activation; quaking homolog; schizophrenia INTRODUCTION LncRNAs are arbitrarily defined as transcripts longer than 200 bp 10 Schizophrenia (SZ) disorder, a debilitating mental disorder that possess no protein-coding capacity. -

Rgma-Induced Neo1 Proteolysis Promotes Neural Tube Morphogenesis
The Journal of Neuroscience, September 18, 2019 • 39(38):7465–7484 • 7465 Development/Plasticity/Repair Rgma-Induced Neo1 Proteolysis Promotes Neural Tube Morphogenesis Sharlene Brown,* Pradeepa Jayachandran,* XMaraki Negesse, Valerie Olmo, Eudorah Vital, and XRachel Brewster Department of Biological Sciences, University of Maryland Baltimore County, Baltimore, Maryland 21250 Neuroepithelial cell (NEC) elongation is one of several key cell behaviors that mediate the tissue-level morphogenetic movements that shape the neural tube (NT), the precursor of the brain and spinal cord. However, the upstream signals that promote NEC elongation have been difficult to tease apart from those regulating apico-basal polarity and hingepoint formation, due to their confounding interdepen- dence. The Repulsive Guidance Molecule a (Rgma)/Neogenin 1 (Neo1) signaling pathway plays a conserved role in NT formation (neu- rulation) and is reported to regulate both NEC elongation and apico-basal polarity, through signal transduction events that have not been identified. We examine here the role of Rgma/Neo1 signaling in zebrafish (sex unknown), an organism that does not use hingepoints to shape its hindbrain, thereby enabling a direct assessment of the role of this pathway in NEC elongation. We confirm that Rgma/Neo1 signaling is required for microtubule-mediated NEC elongation, and demonstrate via cell transplantation that Neo1 functions cell autonomously to promote elongation. However, in contrast to previous findings, our data do not support a role for this pathway in establishing apical junctional complexes. Last, we provide evidence that Rgma promotes Neo1 glycosylation and intramembrane prote- olysis, resulting in the production of a transient, nuclear intracellular fragment (NeoICD). Partial rescue of Neo1a and Rgma knockdown embryos by overexpressing neoICD suggests that this proteolytic cleavage is essential for neurulation. -

Whole-Genome Sequencing of Finnish Type 1 Diabetic Siblings Discordant for Kidney Disease Reveals DNA Variants Associated with Diabetic Nephropathy
BASIC RESEARCH www.jasn.org Whole-Genome Sequencing of Finnish Type 1 Diabetic Siblings Discordant for Kidney Disease Reveals DNA Variants associated with Diabetic Nephropathy Jing Guo ,1,2 Owen J. L. Rackham ,2 Niina Sandholm ,3,4,5 Bing He ,1 Anne-May Österholm,1,2 Erkka Valo ,3,4,5 Valma Harjutsalo ,3,4,5,6 Carol Forsblom,3,4,5 Iiro Toppila,3,4,5 Maija Parkkonen,3,4,5 Qibin Li,7 Wenjuan Zhu,7 Nathan Harmston ,2,8 Sonia Chothani,2 Miina K. Öhman ,2 Eudora Eng,2 Yang Sun,2 Enrico Petretto ,2,9 Per-Henrik Groop,3,4,5,10 and Karl Tryggvason1,2,11 Due to the number of contributing authors, the affiliations are listed at the end of this article. ABSTRACT Background Several genetic susceptibility loci associated with diabetic nephropathy have been documen- ted, but no causative variants implying novel pathogenetic mechanisms have been elucidated. Methods We carried out whole-genome sequencing of a discovery cohort of Finnish siblings with type 1 diabetes who were discordant for the presence (case) or absence (control) of diabetic nephropathy. Con- trols had diabetes without complications for 15–37 years. We analyzed and annotated variants at genome, gene, and single-nucleotide variant levels. We then replicated the associated variants, genes, and regions in a replication cohort from the Finnish Diabetic Nephropathy study that included 3531 unrelated Finns with type 1 diabetes. Results We observed protein-altering variants and an enrichment of variants in regions associated with the presence or absence of diabetic nephropathy. The replication cohort confirmed variants in both regulatory and protein-coding regions. -

The Emerging Roles of the RNA Binding Protein QKI in Cardiovascular Development and Function
fcell-09-668659 June 14, 2021 Time: 13:17 # 1 MINI REVIEW published: 16 June 2021 doi: 10.3389/fcell.2021.668659 The Emerging Roles of the RNA Binding Protein QKI in Cardiovascular Development and Function Xinyun Chen1,2†, Jianwen Yin3†, Dayan Cao1, Deyong Xiao1, Zhongjun Zhou4, Ying Liu1* and Weinian Shou1* 1 Department of Pediatrics, Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN, United States, 2 Guangdong Key Laboratory for Genome Stability and Human Disease Prevention, Department of Biochemistry and Molecular Biology, School of Basic Medical Sciences, Shenzhen University, Shenzhen, China, 3 Department of Foot, Ankle and Hand Surgery, Shenzhen Second People’s Hospital, First Affiliated Hospital of Shenzhen University, Shenzhen, China, 4 Faculty of Medicine, School of Biomedical Sciences, The University of Hong Kong, Hong Kong Edited by: Janine M. Van Gils, Leiden University Medical Center, RNA binding proteins (RBPs) have a broad biological and physiological function and Netherlands are critical in regulating pre-mRNA posttranscriptional processing, intracellular migration, Reviewed by: Jianbo Wang, and mRNA stability. QKI, also known as Quaking, is a member of the signal transduction University of Alabama at Birmingham, and activation of RNA (STAR) family, which also belongs to the heterogeneous nuclear United States ribonucleoprotein K- (hnRNP K-) homology domain protein family. There are three Wuqiang Zhu, Mayo Clinic Arizona, United States major alternatively spliced isoforms, QKI-5, QKI-6, and QKI-7, differing in carboxy- *Correspondence: terminal domains. They share a common RNA binding property, but each isoform can Ying Liu regulate pre-mRNA splicing, transportation or stability differently in a unique cell type- [email protected] Weinian Shou specific manner. -

A Fast, Non-Radioactive Method to Identify in Vivo Targets of RNA-Binding Proteins
bioRxiv preprint doi: https://doi.org/10.1101/158410; this version posted July 1, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. uvCLAP: a fast, non-radioactive method to identify in vivo targets of RNA-binding proteins Daniel Maticzka1,4, Ibrahim Avsar Ilik2,4, Tugce Aktas2,4, Rolf Backofen1,3,# and Asifa Akhtar2,#* 1 Bioinformatics Group, Department of Computer Science, University of Freiburg, Freiburg, Germany 2 Max Planck Institute of Immunobiology and Epigenetics, 79108 Freiburg im Breisgau, Germany 3 Centre for Biological Signalling Studies (BIOSS), University of Freiburg, Freiburg, Germany 4 These authors contributed equally to this work # Corresponding author [email protected] Phone: +49 761 203 7461 Fax: +49 761 203 7462 # Corresponding author, * Lead contact [email protected] Phone: +49 (0)7615108565 Fax: +49 (0)7615108566 1 bioRxiv preprint doi: https://doi.org/10.1101/158410; this version posted July 1, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract RNA-binding proteins (RBPs) play important and essential roles in eukaryotic gene expression regulating splicing, localization, translation and stability of mRNAs. Understanding the exact contribution of RBPs to gene regulation is crucial as many RBPs are frequently mis-regulated in several neurological diseases and certain cancers. While recently developed techniques provide binding sites of RBPs, they are labor-intensive and generally rely on radioactive labeling of RNA. With more than 1,000 RBPs in a human cell, it is imperative to develop easy, robust, reproducible and high-throughput methods to determine in vivo targets of RBPs.