A Fast, Non-Radioactive Method to Identify in Vivo Targets of RNA-Binding Proteins
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Atlas Antibodies in Breast Cancer Research Table of Contents
ATLAS ANTIBODIES IN BREAST CANCER RESEARCH TABLE OF CONTENTS The Human Protein Atlas, Triple A Polyclonals and PrecisA Monoclonals (4-5) Clinical markers (6) Antibodies used in breast cancer research (7-13) Antibodies against MammaPrint and other gene expression test proteins (14-16) Antibodies identified in the Human Protein Atlas (17-14) Finding cancer biomarkers, as exemplified by RBM3, granulin and anillin (19-22) Co-Development program (23) Contact (24) Page 2 (24) Page 3 (24) The Human Protein Atlas: a map of the Human Proteome The Human Protein Atlas (HPA) is a The Human Protein Atlas consortium cell types. All the IHC images for Swedish-based program initiated in is mainly funded by the Knut and Alice the normal tissue have undergone 2003 with the aim to map all the human Wallenberg Foundation. pathology-based annotation of proteins in cells, tissues and organs expression levels. using integration of various omics The Human Protein Atlas consists of technologies, including antibody- six separate parts, each focusing on References based imaging, mass spectrometry- a particular aspect of the genome- 1. Sjöstedt E, et al. (2020) An atlas of the based proteomics, transcriptomics wide analysis of the human proteins: protein-coding genes in the human, pig, and and systems biology. mouse brain. Science 367(6482) 2. Thul PJ, et al. (2017) A subcellular map of • The Tissue Atlas shows the the human proteome. Science. 356(6340): All the data in the knowledge resource distribution of proteins across all eaal3321 is open access to allow scientists both major tissues and organs in the 3. -
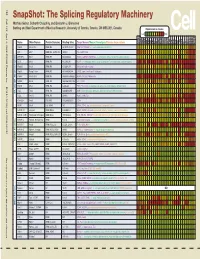
Snapshot: the Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J
192 Cell SnapShot: The Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J. Blencowe 133 Banting and Best Department of Medical Research, University of Toronto, Toronto, ON M5S 3E1, Canada Expression in mouse , April4, 2008©2008Elsevier Inc. Low High Name Other Names Protein Domains Binding Sites Target Genes/Mouse Phenotypes/Disease Associations Amy Ceb Hip Hyp OB Eye SC BM Bo Ht SM Epd Kd Liv Lu Pan Pla Pro Sto Spl Thy Thd Te Ut Ov E6.5 E8.5 E10.5 SRp20 Sfrs3, X16 RRM, RS GCUCCUCUUC SRp20, CT/CGRP; −/− early embryonic lethal E3.5 9G8 Sfrs7 RRM, RS, C2HC Znf (GAC)n Tau, GnRH, 9G8 ASF/SF2 Sfrs1 RRM, RS RGAAGAAC HipK3, CaMKIIδ, HIV RNAs; −/− embryonic lethal, cond. KO cardiomyopathy SC35 Sfrs2 RRM, RS UGCUGUU AChE; −/− embryonic lethal, cond. KO deficient T-cell maturation, cardiomyopathy; LS SRp30c Sfrs9 RRM, RS CUGGAUU Glucocorticoid receptor SRp38 Fusip1, Nssr RRM, RS ACAAAGACAA CREB, type II and type XI collagens SRp40 Sfrs5, HRS RRM, RS AGGAGAAGGGA HipK3, PKCβ-II, Fibronectin SRp55 Sfrs6 RRM, RS GGCAGCACCUG cTnT, CD44 DOI 10.1016/j.cell.2008.03.010 SRp75 Sfrs4 RRM, RS GAAGGA FN1, E1A, CD45; overexpression enhances chondrogenic differentiation Tra2α Tra2a RRM, RS GAAARGARR GnRH; overexpression promotes RA-induced neural differentiation SR and SR-Related Proteins Tra2β Sfrs10 RRM, RS (GAA)n HipK3, SMN, Tau SRm160 Srrm1 RS, PWI AUGAAGAGGA CD44 SWAP Sfrs8 RS, SWAP ND SWAP, CD45, Tau; possible asthma susceptibility gene hnRNP A1 Hnrnpa1 RRM, RGG UAGGGA/U HipK3, SMN2, c-H-ras; rheumatoid arthritis, systemic lupus -

Logistic Regression Analysis for the Identification of the Metastasis-Associated Signaling Pathways of Osteosarcoma
INTERNATIONAL JOURNAL OF MOLECULAR MEDICINE 41: 1233-1244, 2018 Logistic regression analysis for the identification of the metastasis-associated signaling pathways of osteosarcoma YANG LIU1, WEI SUN2, XIAOJUN MA2, YUEDONG HAO3, GANG LIU3, XIAOHUI HU3, HOULAI SHANG3, PENGFEI WU3, ZEXUE ZHAO3 and WEIDONG LIU3 1Department of Orthopedics, Affiliated Hospital of Inner Mongolia University for The Nationalities, Tongliao, Inner Mongolia 028007; 2Department of Orthopaedics, Shanghai General Hospital, School of Medicine, Shanghai Jiao Tong University, Shanghai 200080; 3Department of Orthopedics, Huai'an First People's Hospital, Nanjing Medical University, Huai'an, Jiangsu 223300, P.R. China Received February 20, 2017; Accepted September 26, 2017 DOI: 10.3892/ijmm.2018.3360 Abstract. Osteosarcoma (OS) is the most common histological migration were identified as the terms that were significantly type of primary bone cancer. The present study was designed different. ICAM1, PDGFB, PDGFRB and TGFB1 were iden- to identify the key genes and signaling pathways involved in the tified to be enriched in cell migration and regulation of cell metastasis of OS. Microarray data of GSE39055 were down- migration. Nucleotide excision repair and the Wnt signaling loaded from the Gene Expression Omnibus database, which pathway were the metastasis-associated pathways of OS. In included 19 OS biopsy specimens before metastasis (control addition, ICAM1, PDGFB, PDGFRB and TGFB1, which were group) and 18 OS biopsy specimens after metastasis (case involved in cell migration and regulation of cell migration may group). After the differentially expressed genes (DEGs) were affect the metastasis of OS. identified using the Linear Models for Microarray Analysis package, hierarchical clustering analysis and unsupervised Introduction clustering analysis were performed separately, using orange software and the self-organization map method. -

Analysis of Alternative Splicing Associated with Aging and Neurodegeneration in the Human Brain
Downloaded from genome.cshlp.org on October 2, 2021 - Published by Cold Spring Harbor Laboratory Press Research Analysis of alternative splicing associated with aging and neurodegeneration in the human brain James R. Tollervey,1 Zhen Wang,1 Tibor Hortoba´gyi,2 Joshua T. Witten,1 Kathi Zarnack,3 Melis Kayikci,1 Tyson A. Clark,4 Anthony C. Schweitzer,4 Gregor Rot,5 Tomazˇ Curk,5 Blazˇ Zupan,5 Boris Rogelj,2 Christopher E. Shaw,2 and Jernej Ule1,6 1MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB2 0QH, United Kingdom; 2MRC Centre for Neurodegeneration Research, King’s College London, Institute of Psychiatry, De Crespigny Park, London SE5 8AF, United Kingdom; 3EMBL–European Bioinformatics Institute, Wellcome Trust Genome Campus, Hinxton, Cambridge CB10 1SD, United Kingdom; 4Expression Research, Affymetrix, Inc., Santa Clara, California 95051, USA; 5Faculty of Computer and Information Science, University of Ljubljana, Trzˇasˇka 25, SI-1000 Ljubljana, Slovenia Age is the most important risk factor for neurodegeneration; however, the effects of aging and neurodegeneration on gene expression in the human brain have most often been studied separately. Here, we analyzed changes in transcript levels and alternative splicing in the temporal cortex of individuals of different ages who were cognitively normal, affected by frontotemporal lobar degeneration (FTLD), or affected by Alzheimer’s disease (AD). We identified age-related splicing changes in cognitively normal individuals and found that these were present also in 95% of individuals with FTLD or AD, independent of their age. These changes were consistent with increased polypyrimidine tract binding protein (PTB)– dependent splicing activity. We also identified disease-specific splicing changes that were present in individuals with FTLD or AD, but not in cognitively normal individuals. -

Dynamic Changes in RNA–Protein Interactions and RNA Secondary Structure in Mammalian Erythropoiesis
Published Online: 27 July, 2021 | Supp Info: http://doi.org/10.26508/lsa.202000659 Downloaded from life-science-alliance.org on 30 September, 2021 Resource Dynamic changes in RNA–protein interactions and RNA secondary structure in mammalian erythropoiesis Mengge Shan1,2 , Xinjun Ji3, Kevin Janssen5 , Ian M Silverman3 , Jesse Humenik3, Ben A Garcia5, Stephen A Liebhaber3,4, Brian D Gregory1,2 Two features of eukaryotic RNA molecules that regulate their and RNA secondary structure. These techniques generally either post-transcriptional fates are RNA secondary structure and RNA- use chemical probing agents or structure-specific RNases (single- binding protein (RBP) interaction sites. However, a comprehen- stranded RNases (ssRNases) and double-stranded RNases [dsRNa- sive global overview of the dynamic nature of these sequence ses]) to provide site-specific evidence for a region being in single- or features during erythropoiesis has never been obtained. Here, we double-stranded configurations (4, 5). use our ribonuclease-mediated structure and RBP-binding site To date, the known repertoire of RBP–RNA interaction sites has mapping approach to reveal the global landscape of RNA sec- been built on a protein-by-protein basis, with studies identifying ondary structure and RBP–RNA interaction sites and the dynamics the targets of a single protein of interest, often through the use of of these features during this important developmental process. techniques such as crosslinking and immunoprecipitation se- We identify dynamic patterns of RNA secondary structure and RBP quencing (CLIP-seq) (6). In CLIP-seq, samples are irradiated with UV binding throughout the process and determine a set of corre- to induce the cross-linking of proteins to their RNA targets. -

Large-Scale Analysis of Genome and Transcriptome Alterations in Multiple Tumors Unveils Novel Cancer-Relevant Splicing Networks
Downloaded from genome.cshlp.org on October 2, 2021 - Published by Cold Spring Harbor Laboratory Press Large-scale analysis of genome and transcriptome alterations in multiple tumors unveils novel cancer-relevant splicing networks Endre Sebestyén1,*, Babita Singh1,*, Belén Miñana1,2, Amadís Pagès1, Francesca Mateo3, Miguel Angel Pujana3, Juan Valcárcel1,2,4, Eduardo Eyras1,4,5 1Universitat Pompeu Fabra, Dr. Aiguader 88, E08003 Barcelona, Spain 2Centre for Genomic Regulation, Dr. Aiguader 88, E08003 Barcelona, Spain 3Program Against Cancer Therapeutic Resistance (ProCURE), Catalan Institute of Oncology (ICO), Bellvitge Institute for Biomedical Research (IDIBELL), E08908 L’Hospitalet del Llobregat, Spain. 4Catalan Institution for Research and Advanced Studies, Passeig Lluís Companys 23, E08010 Barcelona, Spain *Equal contribution 5Correspondence to: [email protected] Keywords: alternative splicing, RNA binding proteins, splicing networks, cancer 1 Downloaded from genome.cshlp.org on October 2, 2021 - Published by Cold Spring Harbor Laboratory Press Abstract Alternative splicing is regulated by multiple RNA-binding proteins and influences the expression of most eukaryotic genes. However, the role of this process in human disease, and particularly in cancer, is only starting to be unveiled. We systematically analyzed mutation, copy number and gene expression patterns of 1348 RNA-binding protein (RBP) genes in 11 solid tumor types, together with alternative splicing changes in these tumors and the enrichment of binding motifs in the alternatively spliced sequences. Our comprehensive study reveals widespread alterations in the expression of RBP genes, as well as novel mutations and copy number variations in association with multiple alternative splicing changes in cancer drivers and oncogenic pathways. Remarkably, the altered splicing patterns in several tumor types recapitulate those of undifferentiated cells. -

Overview of Research on Fusion Genes in Prostate Cancer
2011 Review Article Overview of research on fusion genes in prostate cancer Chunjiao Song1,2, Huan Chen3 1Medical Research Center, Shaoxing People’s Hospital, Shaoxing University School of Medicine, Shaoxing 312000, China; 2Shaoxing Hospital, Zhejiang University School of Medicine, Shaoxing 312000, China; 3Key Laboratory of Microorganism Technology and Bioinformatics Research of Zhejiang Province, Zhejiang Institute of Microbiology, Hangzhou 310000, China Contributions: (I) Conception and design: C Song; (II) Administrative support: Shaoxing Municipal Health and Family Planning Science and Technology Innovation Project (2017CX004) and Shaoxing Public Welfare Applied Research Project (2018C30058); (III) Provision of study materials or patients: None; (IV) Collection and assembly of data: C Song; (V) Data analysis and interpretation: H Chen; (VI) Manuscript writing: All authors; (VII) Final approval of manuscript: All authors. Correspondence to: Chunjiao Song. No. 568 Zhongxing Bei Road, Shaoxing 312000, China. Email: [email protected]. Abstract: Fusion genes are known to drive and promote carcinogenesis and cancer progression. In recent years, the rapid development of biotechnologies has led to the discovery of a large number of fusion genes in prostate cancer specimens. To further investigate them, we summarized the fusion genes. We searched related articles in PubMed, CNKI (Chinese National Knowledge Infrastructure) and other databases, and the data of 92 literatures were summarized after preliminary screening. In this review, we summarized approximated 400 fusion genes since the first specific fusion TMPRSS2-ERG was discovered in prostate cancer in 2005. Some of these are prostate cancer specific, some are high-frequency in the prostate cancer of a certain ethnic group. This is a summary of scientific research in related fields and suggests that some fusion genes may become biomarkers or the targets for individualized therapies. -

Genome-Wide Analysis of the Alternative Splicing Profiles
Journal of Cancer 2020, Vol. 11 1542 Ivyspring International Publisher Journal of Cancer 2020; 11(6): 1542-1554. doi: 10.7150/jca.36998 Research Paper Genome-wide Analysis of the Alternative Splicing Profiles Revealed Novel Prognostic Index for Kidney Renal Cell Clear Cell Carcinoma Li Gao1#, Rong-quan He2#, Zhi-guang Huang1, Yi-wu Dang1, Yong-yao Gu1, Hai-biao Yan3, Sheng-hua Li3, Gang Chen1 1. Department of Pathology, First Affiliated Hospital of Guangxi Medical University, Nanning, Guangxi Zhuang Autonomous Region 530021, P. R. China 2. Department of Medical Oncology, First Affiliated Hospital of Guangxi Medical University, Nanning, Guangxi Zhuang Autonomous Region 530021, P. R. China 3. Department of Urology, First Affiliated Hospital of Guangxi Medical University, Nanning, Guangxi Zhuang Autonomous Region 530021, P. R. China # Li Gao and Rong-quan He contributed equally to this study. Corresponding authors: Sheng-hua Li: Tel: +86 0771-5356533; Fax: 0086-771-5350031; Email: [email protected]; Gang Chen: Tel: +86 0771-5356534; Fax: 0086-771-5356534; Email: [email protected] © The author(s). This is an open access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/). See http://ivyspring.com/terms for full terms and conditions. Received: 2019.05.25; Accepted: 2019.12.13; Published: 2020.01.14 Abstract Alternative splicing (AS) is a major mechanism that greatly enhanced the diversity of proteome. Mounting evidence demonstrated that aberration of AS are important steps for the initiation and progression of human cancers. Here, we comprehensively investigated the association between whole landscape of AS profiles and the survival outcome of renal cell carcinoma (RCC) patients using RNA-seq data from TCGA SpliceSeq. -

The Long Non-Coding RNA Gomafu Is Acutely Regulated in Response to Neuronal Activation and Involved in Schizophrenia-Associated Alternative Splicing
Molecular Psychiatry (2014) 19, 486–494 & 2014 Macmillan Publishers Limited All rights reserved 1359-4184/14 www.nature.com/mp ORIGINAL ARTICLE The long non-coding RNA Gomafu is acutely regulated in response to neuronal activation and involved in schizophrenia-associated alternative splicing G Barry1, JA Briggs2, DP Vanichkina1, EM Poth3, NJ Beveridge4,5, VS Ratnu6, SP Nayler2, K Nones7,JHu8, TW Bredy6, S Nakagawa9, F Rigo10, RJ Taft1, MJ Cairns4,5, S Blackshaw3, EJ Wolvetang2 and JS Mattick1,11,12 Schizophrenia (SZ) is a complex disease characterized by impaired neuronal functioning. Although defective alternative splicing has been linked to SZ, the molecular mechanisms responsible are unknown. Additionally, there is limited understanding of the early transcriptomic responses to neuronal activation. Here, we profile these transcriptomic responses and show that long non-coding RNAs (lncRNAs) are dynamically regulated by neuronal activation, including acute downregulation of the lncRNA Gomafu, previously implicated in brain and retinal development. Moreover, we demonstrate that Gomafu binds directly to the splicing factors QKI and SRSF1 (serine/arginine-rich splicing factor 1) and dysregulation of Gomafu leads to alternative splicing patterns that resemble those observed in SZ for the archetypal SZ-associated genes DISC1 and ERBB4. Finally, we show that Gomafu is downregulated in post-mortem cortical gray matter from the superior temporal gyrus in SZ. These results functionally link activity-regulated lncRNAs and alternative splicing in neuronal function and suggest that their dysregulation may contribute to neurological disorders. Molecular Psychiatry (2014) 19, 486–494; doi:10.1038/mp.2013.45; published online 30 April 2013 Keywords: alternative splicing; Gomafu; neuronal activation; quaking homolog; schizophrenia INTRODUCTION LncRNAs are arbitrarily defined as transcripts longer than 200 bp 10 Schizophrenia (SZ) disorder, a debilitating mental disorder that possess no protein-coding capacity. -

RNA-Seq Analysis of Prostate Cancer in the Chinese Population Identifies Recurrent Gene Fusions, Cancer-Associated Long Noncodin
npg RNA-seq analysis of prostate cancer in Chinese population Cell Research (2012) 22:806-821. 806 © 2012 IBCB, SIBS, CAS All rights reserved 1001-0602/12 $ 32.00 npg ORIGINAL ARTICLE www.nature.com/cr RNA-seq analysis of prostate cancer in the Chinese population identifies recurrent gene fusions, cancer-associated long noncoding RNAs and aberrant alternative splicings Shancheng Ren1, *, Zhiyu Peng2, *, Jian-Hua Mao3, *, Yongwei Yu4, Changjun Yin5, Xin Gao6, Zilian Cui1, Jibin Zhang2, Kang Yi2, Weidong Xu1, Chao Chen2, Fubo Wang1, Xinwu Guo2, Ji Lu1, Jun Yang2, Min Wei1, Zhijian Tian2, Yinghui Guan3, Liang Tang1, Chuanliang Xu1, Linhui Wang1, Xu Gao1, Wei Tian2, Jian Wang2, Huanming Yang2, Jun Wang2, Yinghao Sun1 1Department of Urology, Shanghai Changhai Hospital, Second Military Medical University, Shanghai 200433, China; 2Beijing Ge- nomics Institute at Shenzhen, Shenzhen, Guangdong 518083, China; 3Life Sciences Division, Lawrence Berkeley National Labora- tory, One Cyclotron Road, Berkeley, CA 94720, USA; 4Department of Pathology, Shanghai Changhai Hospital, Second Military Medical University, Shanghai 200433, China; 5Department of Urology, Jiangsu Provincial People’s Hospital, Nanjing Medical University, Nanjing, Jiangsu 210029, China; 6Department of Urology, the Third Affiliated Hospital of Sun Yat Sen University, Guangzhou, Guangdong 510630, China There are remarkable disparities among patients of different races with prostate cancer; however, the mechanism underlying this difference remains unclear. Here, we present a comprehensive landscape of the transcriptome profiles of 14 primary prostate cancers and their paired normal counterparts from the Chinese population using RNA-seq, revealing tremendous diversity across prostate cancer transcriptomes with respect to gene fusions, long noncoding RNAs (long ncRNA), alternative splicing and somatic mutations. -

The Emerging Roles of the RNA Binding Protein QKI in Cardiovascular Development and Function
fcell-09-668659 June 14, 2021 Time: 13:17 # 1 MINI REVIEW published: 16 June 2021 doi: 10.3389/fcell.2021.668659 The Emerging Roles of the RNA Binding Protein QKI in Cardiovascular Development and Function Xinyun Chen1,2†, Jianwen Yin3†, Dayan Cao1, Deyong Xiao1, Zhongjun Zhou4, Ying Liu1* and Weinian Shou1* 1 Department of Pediatrics, Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN, United States, 2 Guangdong Key Laboratory for Genome Stability and Human Disease Prevention, Department of Biochemistry and Molecular Biology, School of Basic Medical Sciences, Shenzhen University, Shenzhen, China, 3 Department of Foot, Ankle and Hand Surgery, Shenzhen Second People’s Hospital, First Affiliated Hospital of Shenzhen University, Shenzhen, China, 4 Faculty of Medicine, School of Biomedical Sciences, The University of Hong Kong, Hong Kong Edited by: Janine M. Van Gils, Leiden University Medical Center, RNA binding proteins (RBPs) have a broad biological and physiological function and Netherlands are critical in regulating pre-mRNA posttranscriptional processing, intracellular migration, Reviewed by: Jianbo Wang, and mRNA stability. QKI, also known as Quaking, is a member of the signal transduction University of Alabama at Birmingham, and activation of RNA (STAR) family, which also belongs to the heterogeneous nuclear United States ribonucleoprotein K- (hnRNP K-) homology domain protein family. There are three Wuqiang Zhu, Mayo Clinic Arizona, United States major alternatively spliced isoforms, QKI-5, QKI-6, and QKI-7, differing in carboxy- *Correspondence: terminal domains. They share a common RNA binding property, but each isoform can Ying Liu regulate pre-mRNA splicing, transportation or stability differently in a unique cell type- [email protected] Weinian Shou specific manner. -

Alternatively Spliced Genes
1 Alternatively Spliced Genes Jane Y. Wu1,2,LiyaYuan1 and Necat Havlioglu1 1Washington University School of Medicine, St. Louis, MO, USA 2John F. Kennedy Center for Research on Human Development, Vanderbilt University Medical Center, Nashville, TN, USA 1 Pre-mRNA Splicing and Splicing Machinery 3 1.1 Splicing Machinery: Spliceosome 3 1.2 Splicing Signals 4 1.3 Spliceosomal UsnRNP Biogenesis 12 1.4 Spliceosome Assembly 13 1.5 Biochemical Mechanisms of pre-mRNA Splicing 17 2 Alternative pre-mRNA Splicing 17 2.1 Alternative Splicing and its Role in Regulating Gene Activities and Generating Genetic Diversity 17 2.1.1 Different Patterns of Alternative Splicing 17 2.1.2 Alternative Splicing and Genetic Diversity 18 2.2 Mechanisms Underlying Alternative Splicing Regulation 19 2.2.1 Splicing Signals and Splicing Regulatory Elements 20 2.2.2 Trans-acting Splicing Regulators 23 2.3 Tissue-specific and Developmentally Regulated Alternative Splicing 26 2.4 Regulation of Alternative Splicing in Response to Extracellular Stimuli 27 3 Pre-mRNA Splicing and Human Diseases 28 3.1 Splicing Defects in Human Diseases 28 3.2 Molecular Mechanisms Underlying Splicing Defects Associated with Disease 33 4 Perspectives on Diagnosis and Treatment of Diseases Caused by pre-mRNA Splicing Defects 36 4.1 Diagnosis of Human Diseases Caused by Splicing Defects 36 2 Alternatively Spliced Genes 4.2 Potential Therapeutic Approaches 37 4.2.1 Oligonucleotide-based Approaches: Antisense, RNAi, and Chimeric Molecules 37 4.2.2 Ribozymes 37 4.2.3 SMaRT 38 4.2.4 Chemical Compounds 38 5 Concluding Remarks 38 Acknowledgment 39 Bibliography 39 Books and Reviews 39 Keywords Pre-mRNA Nascent transcripts that are precursors of mature messenger RNAs.