Original Article High Frequency of the SDK1:AMACR Fusion Transcript in Chinese Prostate Cancer
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Atlas Antibodies in Breast Cancer Research Table of Contents
ATLAS ANTIBODIES IN BREAST CANCER RESEARCH TABLE OF CONTENTS The Human Protein Atlas, Triple A Polyclonals and PrecisA Monoclonals (4-5) Clinical markers (6) Antibodies used in breast cancer research (7-13) Antibodies against MammaPrint and other gene expression test proteins (14-16) Antibodies identified in the Human Protein Atlas (17-14) Finding cancer biomarkers, as exemplified by RBM3, granulin and anillin (19-22) Co-Development program (23) Contact (24) Page 2 (24) Page 3 (24) The Human Protein Atlas: a map of the Human Proteome The Human Protein Atlas (HPA) is a The Human Protein Atlas consortium cell types. All the IHC images for Swedish-based program initiated in is mainly funded by the Knut and Alice the normal tissue have undergone 2003 with the aim to map all the human Wallenberg Foundation. pathology-based annotation of proteins in cells, tissues and organs expression levels. using integration of various omics The Human Protein Atlas consists of technologies, including antibody- six separate parts, each focusing on References based imaging, mass spectrometry- a particular aspect of the genome- 1. Sjöstedt E, et al. (2020) An atlas of the based proteomics, transcriptomics wide analysis of the human proteins: protein-coding genes in the human, pig, and and systems biology. mouse brain. Science 367(6482) 2. Thul PJ, et al. (2017) A subcellular map of • The Tissue Atlas shows the the human proteome. Science. 356(6340): All the data in the knowledge resource distribution of proteins across all eaal3321 is open access to allow scientists both major tissues and organs in the 3. -

Cell Adhesion and Specificity
RESEARCH ARTICLE Molecular basis of sidekick-mediated cell- cell adhesion and specificity Kerry M Goodman1†, Masahito Yamagata2,3†, Xiangshu Jin1,4‡, Seetha Mannepalli1, Phinikoula S Katsamba4,5, Go¨ ran Ahlse´ n4,5, Alina P Sergeeva4,5, Barry Honig1,4,5,6,7*, Joshua R Sanes2,3*, Lawrence Shapiro1,5,7* 1Department of Biochemistry and Molecular Biophysics, Columbia University, New York, United States; 2Department of Molecular and Cellular Biology, Harvard University, Cambridge, United States; 3Center for Brain Science, Harvard University, Cambridge, United States; 4Howard Hughes Medical Institute, Columbia University, New York, United States; 5Department of Systems Biology, Columbia University, New York, United States; 6Department of Medicine, Columbia University, New York, United States; 7Zuckerman Mind Brain and Behavior Institute, Columbia University, New York, United States *For correspondence: bh6@cumc. Abstract Sidekick (Sdk) 1 and 2 are related immunoglobulin superfamily cell adhesion proteins columbia.edu (BH); sanesj@mcb. required for appropriate synaptic connections between specific subtypes of retinal neurons. Sdks harvard.edu (JRS); shapiro@ mediate cell-cell adhesion with homophilic specificity that underlies their neuronal targeting convex.hhmi.columbia.edu (LS) function. Here we report crystal structures of Sdk1 and Sdk2 ectodomain regions, revealing similar †These authors contributed homodimers mediated by the four N-terminal immunoglobulin domains (Ig1–4), arranged in a equally to this work horseshoe conformation. These Ig1–4 horseshoes interact in a novel back-to-back orientation in both homodimers through Ig1:Ig2, Ig1:Ig1 and Ig3:Ig4 interactions. Structure-guided mutagenesis Present address: ‡Department results show that this canonical dimer is required for both Sdk-mediated cell aggregation (via trans of Chemistry, Michigan State interactions) and Sdk clustering in isolated cells (via cis interactions). -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -
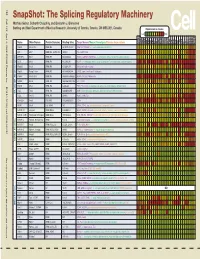
Snapshot: the Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J
192 Cell SnapShot: The Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J. Blencowe 133 Banting and Best Department of Medical Research, University of Toronto, Toronto, ON M5S 3E1, Canada Expression in mouse , April4, 2008©2008Elsevier Inc. Low High Name Other Names Protein Domains Binding Sites Target Genes/Mouse Phenotypes/Disease Associations Amy Ceb Hip Hyp OB Eye SC BM Bo Ht SM Epd Kd Liv Lu Pan Pla Pro Sto Spl Thy Thd Te Ut Ov E6.5 E8.5 E10.5 SRp20 Sfrs3, X16 RRM, RS GCUCCUCUUC SRp20, CT/CGRP; −/− early embryonic lethal E3.5 9G8 Sfrs7 RRM, RS, C2HC Znf (GAC)n Tau, GnRH, 9G8 ASF/SF2 Sfrs1 RRM, RS RGAAGAAC HipK3, CaMKIIδ, HIV RNAs; −/− embryonic lethal, cond. KO cardiomyopathy SC35 Sfrs2 RRM, RS UGCUGUU AChE; −/− embryonic lethal, cond. KO deficient T-cell maturation, cardiomyopathy; LS SRp30c Sfrs9 RRM, RS CUGGAUU Glucocorticoid receptor SRp38 Fusip1, Nssr RRM, RS ACAAAGACAA CREB, type II and type XI collagens SRp40 Sfrs5, HRS RRM, RS AGGAGAAGGGA HipK3, PKCβ-II, Fibronectin SRp55 Sfrs6 RRM, RS GGCAGCACCUG cTnT, CD44 DOI 10.1016/j.cell.2008.03.010 SRp75 Sfrs4 RRM, RS GAAGGA FN1, E1A, CD45; overexpression enhances chondrogenic differentiation Tra2α Tra2a RRM, RS GAAARGARR GnRH; overexpression promotes RA-induced neural differentiation SR and SR-Related Proteins Tra2β Sfrs10 RRM, RS (GAA)n HipK3, SMN, Tau SRm160 Srrm1 RS, PWI AUGAAGAGGA CD44 SWAP Sfrs8 RS, SWAP ND SWAP, CD45, Tau; possible asthma susceptibility gene hnRNP A1 Hnrnpa1 RRM, RGG UAGGGA/U HipK3, SMN2, c-H-ras; rheumatoid arthritis, systemic lupus -

Logistic Regression Analysis for the Identification of the Metastasis-Associated Signaling Pathways of Osteosarcoma
INTERNATIONAL JOURNAL OF MOLECULAR MEDICINE 41: 1233-1244, 2018 Logistic regression analysis for the identification of the metastasis-associated signaling pathways of osteosarcoma YANG LIU1, WEI SUN2, XIAOJUN MA2, YUEDONG HAO3, GANG LIU3, XIAOHUI HU3, HOULAI SHANG3, PENGFEI WU3, ZEXUE ZHAO3 and WEIDONG LIU3 1Department of Orthopedics, Affiliated Hospital of Inner Mongolia University for The Nationalities, Tongliao, Inner Mongolia 028007; 2Department of Orthopaedics, Shanghai General Hospital, School of Medicine, Shanghai Jiao Tong University, Shanghai 200080; 3Department of Orthopedics, Huai'an First People's Hospital, Nanjing Medical University, Huai'an, Jiangsu 223300, P.R. China Received February 20, 2017; Accepted September 26, 2017 DOI: 10.3892/ijmm.2018.3360 Abstract. Osteosarcoma (OS) is the most common histological migration were identified as the terms that were significantly type of primary bone cancer. The present study was designed different. ICAM1, PDGFB, PDGFRB and TGFB1 were iden- to identify the key genes and signaling pathways involved in the tified to be enriched in cell migration and regulation of cell metastasis of OS. Microarray data of GSE39055 were down- migration. Nucleotide excision repair and the Wnt signaling loaded from the Gene Expression Omnibus database, which pathway were the metastasis-associated pathways of OS. In included 19 OS biopsy specimens before metastasis (control addition, ICAM1, PDGFB, PDGFRB and TGFB1, which were group) and 18 OS biopsy specimens after metastasis (case involved in cell migration and regulation of cell migration may group). After the differentially expressed genes (DEGs) were affect the metastasis of OS. identified using the Linear Models for Microarray Analysis package, hierarchical clustering analysis and unsupervised Introduction clustering analysis were performed separately, using orange software and the self-organization map method. -

Analysis of Alternative Splicing Associated with Aging and Neurodegeneration in the Human Brain
Downloaded from genome.cshlp.org on October 2, 2021 - Published by Cold Spring Harbor Laboratory Press Research Analysis of alternative splicing associated with aging and neurodegeneration in the human brain James R. Tollervey,1 Zhen Wang,1 Tibor Hortoba´gyi,2 Joshua T. Witten,1 Kathi Zarnack,3 Melis Kayikci,1 Tyson A. Clark,4 Anthony C. Schweitzer,4 Gregor Rot,5 Tomazˇ Curk,5 Blazˇ Zupan,5 Boris Rogelj,2 Christopher E. Shaw,2 and Jernej Ule1,6 1MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB2 0QH, United Kingdom; 2MRC Centre for Neurodegeneration Research, King’s College London, Institute of Psychiatry, De Crespigny Park, London SE5 8AF, United Kingdom; 3EMBL–European Bioinformatics Institute, Wellcome Trust Genome Campus, Hinxton, Cambridge CB10 1SD, United Kingdom; 4Expression Research, Affymetrix, Inc., Santa Clara, California 95051, USA; 5Faculty of Computer and Information Science, University of Ljubljana, Trzˇasˇka 25, SI-1000 Ljubljana, Slovenia Age is the most important risk factor for neurodegeneration; however, the effects of aging and neurodegeneration on gene expression in the human brain have most often been studied separately. Here, we analyzed changes in transcript levels and alternative splicing in the temporal cortex of individuals of different ages who were cognitively normal, affected by frontotemporal lobar degeneration (FTLD), or affected by Alzheimer’s disease (AD). We identified age-related splicing changes in cognitively normal individuals and found that these were present also in 95% of individuals with FTLD or AD, independent of their age. These changes were consistent with increased polypyrimidine tract binding protein (PTB)– dependent splicing activity. We also identified disease-specific splicing changes that were present in individuals with FTLD or AD, but not in cognitively normal individuals. -

Dynamic Changes in RNA–Protein Interactions and RNA Secondary Structure in Mammalian Erythropoiesis
Published Online: 27 July, 2021 | Supp Info: http://doi.org/10.26508/lsa.202000659 Downloaded from life-science-alliance.org on 30 September, 2021 Resource Dynamic changes in RNA–protein interactions and RNA secondary structure in mammalian erythropoiesis Mengge Shan1,2 , Xinjun Ji3, Kevin Janssen5 , Ian M Silverman3 , Jesse Humenik3, Ben A Garcia5, Stephen A Liebhaber3,4, Brian D Gregory1,2 Two features of eukaryotic RNA molecules that regulate their and RNA secondary structure. These techniques generally either post-transcriptional fates are RNA secondary structure and RNA- use chemical probing agents or structure-specific RNases (single- binding protein (RBP) interaction sites. However, a comprehen- stranded RNases (ssRNases) and double-stranded RNases [dsRNa- sive global overview of the dynamic nature of these sequence ses]) to provide site-specific evidence for a region being in single- or features during erythropoiesis has never been obtained. Here, we double-stranded configurations (4, 5). use our ribonuclease-mediated structure and RBP-binding site To date, the known repertoire of RBP–RNA interaction sites has mapping approach to reveal the global landscape of RNA sec- been built on a protein-by-protein basis, with studies identifying ondary structure and RBP–RNA interaction sites and the dynamics the targets of a single protein of interest, often through the use of of these features during this important developmental process. techniques such as crosslinking and immunoprecipitation se- We identify dynamic patterns of RNA secondary structure and RBP quencing (CLIP-seq) (6). In CLIP-seq, samples are irradiated with UV binding throughout the process and determine a set of corre- to induce the cross-linking of proteins to their RNA targets. -

Large-Scale Analysis of Genome and Transcriptome Alterations in Multiple Tumors Unveils Novel Cancer-Relevant Splicing Networks
Downloaded from genome.cshlp.org on October 2, 2021 - Published by Cold Spring Harbor Laboratory Press Large-scale analysis of genome and transcriptome alterations in multiple tumors unveils novel cancer-relevant splicing networks Endre Sebestyén1,*, Babita Singh1,*, Belén Miñana1,2, Amadís Pagès1, Francesca Mateo3, Miguel Angel Pujana3, Juan Valcárcel1,2,4, Eduardo Eyras1,4,5 1Universitat Pompeu Fabra, Dr. Aiguader 88, E08003 Barcelona, Spain 2Centre for Genomic Regulation, Dr. Aiguader 88, E08003 Barcelona, Spain 3Program Against Cancer Therapeutic Resistance (ProCURE), Catalan Institute of Oncology (ICO), Bellvitge Institute for Biomedical Research (IDIBELL), E08908 L’Hospitalet del Llobregat, Spain. 4Catalan Institution for Research and Advanced Studies, Passeig Lluís Companys 23, E08010 Barcelona, Spain *Equal contribution 5Correspondence to: [email protected] Keywords: alternative splicing, RNA binding proteins, splicing networks, cancer 1 Downloaded from genome.cshlp.org on October 2, 2021 - Published by Cold Spring Harbor Laboratory Press Abstract Alternative splicing is regulated by multiple RNA-binding proteins and influences the expression of most eukaryotic genes. However, the role of this process in human disease, and particularly in cancer, is only starting to be unveiled. We systematically analyzed mutation, copy number and gene expression patterns of 1348 RNA-binding protein (RBP) genes in 11 solid tumor types, together with alternative splicing changes in these tumors and the enrichment of binding motifs in the alternatively spliced sequences. Our comprehensive study reveals widespread alterations in the expression of RBP genes, as well as novel mutations and copy number variations in association with multiple alternative splicing changes in cancer drivers and oncogenic pathways. Remarkably, the altered splicing patterns in several tumor types recapitulate those of undifferentiated cells. -

Longitudinal Genome-Wide Methylation Study of PTSD Treatment Using Prolonged Exposure and Hydrocortisone ✉ Ruoting Yang 1,4 , Changxin Xu2,4, Linda M
Translational Psychiatry www.nature.com/tp ARTICLE OPEN Longitudinal genome-wide methylation study of PTSD treatment using prolonged exposure and hydrocortisone ✉ Ruoting Yang 1,4 , Changxin Xu2,4, Linda M. Bierer2, Janine D. Flory2,3, Aarti Gautam1, Heather N. Bader2, Amy Lehrner2,3, Iouri Makotkine3, Frank Desarnaud3, Stacy A. Miller1, Marti Jett 1, Rasha Hammamieh1,5 and Rachel Yehuda2,3,5 © The Author(s) 2021 Epigenetic changes are currently invoked as explanations for both the chronicity and tenacity of post-traumatic stress disorder (PTSD), a heterogeneous condition showing varying, sometimes idiosyncratic responses to treatment. This study evaluated epigenetic markers in the context of a randomized clinical trial of PTSD patients undergoing prolonged-exposure psychotherapy with and without a hydrocortisone augmentation prior to each session. The purpose of the longitudinal epigenome-wide analyses was to identify predictors of recovery (from pretreatment data) or markers associated with symptom change (based on differences between pre- and post-therapy epigenetic changes). The results of these analyses identified the CREB–BDNF signaling pathway, previously linked to startle reaction and fear learning and memory processes, as a convergent marker predicting both symptom change and severity. Several previous-reported resilience markers were also identified (FKBP5, NR3C1, SDK1, and MAD1L1) to associate with PTSD recovery in this study. Especially, the methylation levels of FKBP5 in the gene body region decreased significantly as CAPS score decreased in responders, while no changes occurred in nonresponders. These biomarkers may have future utility in understanding clinical recovery in PTSD and potential applications in predicting treatment effects. Translational Psychiatry (2021) 11:398 ; https://doi.org/10.1038/s41398-021-01513-5 INTRODUCTION association with recovery, or that themselves change in associa- Plasticity of the epigenome is a mechanism through which tion with symptom severity. -

Overview of Research on Fusion Genes in Prostate Cancer
2011 Review Article Overview of research on fusion genes in prostate cancer Chunjiao Song1,2, Huan Chen3 1Medical Research Center, Shaoxing People’s Hospital, Shaoxing University School of Medicine, Shaoxing 312000, China; 2Shaoxing Hospital, Zhejiang University School of Medicine, Shaoxing 312000, China; 3Key Laboratory of Microorganism Technology and Bioinformatics Research of Zhejiang Province, Zhejiang Institute of Microbiology, Hangzhou 310000, China Contributions: (I) Conception and design: C Song; (II) Administrative support: Shaoxing Municipal Health and Family Planning Science and Technology Innovation Project (2017CX004) and Shaoxing Public Welfare Applied Research Project (2018C30058); (III) Provision of study materials or patients: None; (IV) Collection and assembly of data: C Song; (V) Data analysis and interpretation: H Chen; (VI) Manuscript writing: All authors; (VII) Final approval of manuscript: All authors. Correspondence to: Chunjiao Song. No. 568 Zhongxing Bei Road, Shaoxing 312000, China. Email: [email protected]. Abstract: Fusion genes are known to drive and promote carcinogenesis and cancer progression. In recent years, the rapid development of biotechnologies has led to the discovery of a large number of fusion genes in prostate cancer specimens. To further investigate them, we summarized the fusion genes. We searched related articles in PubMed, CNKI (Chinese National Knowledge Infrastructure) and other databases, and the data of 92 literatures were summarized after preliminary screening. In this review, we summarized approximated 400 fusion genes since the first specific fusion TMPRSS2-ERG was discovered in prostate cancer in 2005. Some of these are prostate cancer specific, some are high-frequency in the prostate cancer of a certain ethnic group. This is a summary of scientific research in related fields and suggests that some fusion genes may become biomarkers or the targets for individualized therapies. -

Epigenetic Biomarkers in Obesity, Weight Loss and Inflammation: a Role for Circadian Rhythm and Methyl Donors
Facultad de Farmacia y Nutrición Epigenetic biomarkers in obesity, weight loss and inflammation: a role for circadian rhythm and methyl donors Mirian Samblas García Pamplona, 2018 Facultad de Farmacia y Nutrición Memoria presentada por Dña. Mirian Samblas García para aspirar al grado de Doctor por la Universidad de Navarra. Mirian Samblas García El presente trabajo ha sido realizado bajo nuestra dirección en el Departamento de Ciencias de la Alimentación y Fisiología de la Facultad de Farmacia y Nutrición de la Universidad de Navarra y autorizamos su presentación ante el Tribunal que lo ha de juzgar. Pamplona, 26 de Febrero de 2018 VºBº Director VºBº Co-Director Dr. Fermín I. Milagro Yoldi Prof. J. Alfredo Martínez Hernández Este trabajo ha sido posible gracias a la financiación de diversas entidades: Ministerio de Economía y Competitividad (AGL2013-45554-R), Centro de Investigación Biomédica en Red de Fisiopatología de la Obesidad y Nutrición (CIBERObn), Instituo de Salud Carlos III (ISCIII), Centro de Investigación en Nutrición (Universidad de Navarra). La investigación que ha dado lugar a estos resultados ha sido impulsada por la beca predoctoral 2014-2015 del Centro de Investigación en Nutrición y las becas predoctoral 2015-2018 y de movilidad del Ministerio de Educación, Cultura y Deporte. “Necesitamos especialmente de la imaginación en las ciencias. No todo es matemáticas y no todo es simple lógica, también se trata de un poco de belleza y poesía” Maria Montessori Dedicado a las dos personas que me lo han dado todo, Mi aita Ray, Mi ama Ana Con especial cariño a, Mi hermana, Bea Mi sobrina, Aroa Mi abu, Esther Agradecimientos/Acknowledgements En estas líneas quisiera mostrar mi más sincero agradecimiento a todas aquellas personas e instituciones que han hecho posible la actual tesis. -

Genome-Wide Analysis of the Alternative Splicing Profiles
Journal of Cancer 2020, Vol. 11 1542 Ivyspring International Publisher Journal of Cancer 2020; 11(6): 1542-1554. doi: 10.7150/jca.36998 Research Paper Genome-wide Analysis of the Alternative Splicing Profiles Revealed Novel Prognostic Index for Kidney Renal Cell Clear Cell Carcinoma Li Gao1#, Rong-quan He2#, Zhi-guang Huang1, Yi-wu Dang1, Yong-yao Gu1, Hai-biao Yan3, Sheng-hua Li3, Gang Chen1 1. Department of Pathology, First Affiliated Hospital of Guangxi Medical University, Nanning, Guangxi Zhuang Autonomous Region 530021, P. R. China 2. Department of Medical Oncology, First Affiliated Hospital of Guangxi Medical University, Nanning, Guangxi Zhuang Autonomous Region 530021, P. R. China 3. Department of Urology, First Affiliated Hospital of Guangxi Medical University, Nanning, Guangxi Zhuang Autonomous Region 530021, P. R. China # Li Gao and Rong-quan He contributed equally to this study. Corresponding authors: Sheng-hua Li: Tel: +86 0771-5356533; Fax: 0086-771-5350031; Email: [email protected]; Gang Chen: Tel: +86 0771-5356534; Fax: 0086-771-5356534; Email: [email protected] © The author(s). This is an open access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/). See http://ivyspring.com/terms for full terms and conditions. Received: 2019.05.25; Accepted: 2019.12.13; Published: 2020.01.14 Abstract Alternative splicing (AS) is a major mechanism that greatly enhanced the diversity of proteome. Mounting evidence demonstrated that aberration of AS are important steps for the initiation and progression of human cancers. Here, we comprehensively investigated the association between whole landscape of AS profiles and the survival outcome of renal cell carcinoma (RCC) patients using RNA-seq data from TCGA SpliceSeq.