Study on Apoptosis in Human Deciduous Tooth Pulp Cells Original
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
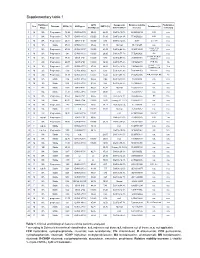
Supplementary Table 2 Supplementary Table 1
Supplementary table 1 Rai/ Binet IGHV Cytogenetic Relative viability Fludarabine- Sex Outcome CD38 (%) IGHV gene ZAP70 (%) Treatment (s) Stage identity (%) abnormalities* increase refractory 1 M 0/A Progressive 14,90 IGHV3-64*05 99,65 28,20 Del17p 18.0% 62,58322819 FCR n.a. 2 F 0/A Progressive 78,77 IGHV3-48*03 100,00 51,90 Del17p 24.8% 77,88052021 FCR n.a. 3 M 0/A Progressive 29,81 IGHV4-b*01 100,00 9,10 Del17p 12.0% 36,48 Len, Chl n.a. 4 M 1/A Stable 97,04 IGHV3-21*01 97,22 18,11 Normal 85,4191657 n.a. n.a. Chl+O, PCR, 5 F 0/A Progressive 87,00 IGHV4-39*07 100,00 43,20 Del13q 68.3% 35,23314039 n.a. HDMP+R 6 M 0/A Progressive 1,81 IGHV3-43*01 100,00 20,90 Del13q 77.7% 57,52490626 Chl n.a. Chl, FR, R-CHOP, 7 M 0/A Progressive 97,80 IGHV1-3*01 100,00 9,80 Del17p 88.5% 48,57389901 n.a. HDMP+R 8 F 2/B Progressive 69,07 IGHV5-a*03 100,00 16,50 Del17p 77.2% 107,9656878 FCR, BA No R-CHOP, FCR, 9 M 1/A Progressive 2,13 IGHV3-23*01 97,22 29,80 Del11q 16.3% 134,5866919 Yes Flavopiridol, BA 10 M 2/A Progressive 0,36 IGHV3-30*02 92,01 0,38 Del13q 81.9% 78,91844953 Unknown n.a. 11 M 2/B Progressive 15,17 IGHV3-20*01 100,00 13,20 Del11q 95.3% 75,52880995 FCR, R-CHOP, BR No 12 M 0/A Stable 0,14 IGHV3-30*02 90,62 7,40 Del13q 13.0% 13,0939004 n.a. -

The UVB-Induced Gene Expression Profile of Human Epidermis in Vivo Is Different from That of Cultured Keratinocytes
Oncogene (2006) 25, 2601–2614 & 2006 Nature Publishing Group All rights reserved 0950-9232/06 $30.00 www.nature.com/onc ORIGINAL ARTICLE The UVB-induced gene expression profile of human epidermis in vivo is different from that of cultured keratinocytes CD Enk1, J Jacob-Hirsch2, H Gal3, I Verbovetski4, N Amariglio2, D Mevorach4, A Ingber1, D Givol3, G Rechavi2 and M Hochberg1 1Department of Dermatology, The Hadassah-Hebrew University Medical Center, Jerusalem, Israel; 2Department of Pediatric Hemato-Oncology and Functional Genomics, Safra Children’s Hospital, Sheba Medical Center and Sackler School of Medicine, Tel-Aviv University,Tel Aviv, Israel; 3Department of Molecular Cell Biology, Weizmann Institute of Science, Rehovot, Israel and 4The Laboratory for Cellular and Molecular Immunology, Department of Medicine, The Hadassah-Hebrew University Medical Center, Jerusalem, Israel In order to obtain a comprehensive picture of the radiation. UVB, with a wavelength range between 290 molecular events regulating cutaneous photodamage of and 320 nm, represents one of the most important intact human epidermis, suction blister roofs obtained environmental hazards affectinghuman skin (Hahn after a single dose of in vivo ultraviolet (UV)B exposure and Weinberg, 2002). To protect itself against the were used for microarray profiling. We found a changed DNA-damaging effects of sunlight, the skin disposes expression of 619 genes. Half of the UVB-regulated genes over highly complicated cellular programs, including had returned to pre-exposure baseline levels at 72 h, cell-cycle arrest, DNA repair and apoptosis (Brash et al., underscoring the transient character of the molecular 1996). Failure in selected elements of these defensive cutaneous UVB response. -

Hsa-Mir-21-5P
Supporting Information An exosomal urinary miRNA signature for early diagnosis of renal fibrosis in Lupus Nephritis 1 2 2 1 1 Cristina Solé , Teresa Moliné , Marta Vidal , Josep Ordi-Ros , Josefina Cortés-Hernández 1Hospital Universitari Vall d’Hebron, Vall d’Hebron Research Institute (VHIR), Lupus Unit, Barcelona, Spain; [email protected] (C.S); [email protected] (J.O-R.); [email protected] (J.C-H.) 2 Hospital Universitari Vall d’Hebron, Department of Renal Pathology, Barcelona, Spain; [email protected] (T.M); [email protected] (M.V). INDEX 1. SI Materials and Methods......................................................................................3 2. SI References…………………………………………………………………………….7 2. SI Figures................................................................................................................8 4. SI Tables................................................................................................................21 2 SI Materials and Methods Patients and samples Patients with biopsy-proven active LN were recruited from the Lupus Unit at Vall d’Hebron Hospital (N=45). All patients fulfilled at least 4 of the American College of rheumatology (ACR) revised classification criteria for SLE [1]. Healthy donors were used as controls (N=20). Urine samples were collected from each patient 1 day before renal biopsy and processed immediately to be stored at -80ºC. Patients with urinary tract infection, diabetes mellitus, pregnancy, malignancy and non-lupus-related renal failure were excluded. In addition, key laboratory measurements were obtained including complement levels (C3 and C4), anti-double-stranded DNA (anti-dsDNA), 24-h proteinuria, blood urea nitrogen (BUN), serum creatinine and the estimated glomerular filtration ratio (eGFR) using the Chronic Kidney Disease Epidemiology Collaboration (CKD-EPI) formula [2]. SLE disease activity was assessed by the SLE Disease Activity Index 2000 update (SLEDAI-2Ks; range 0–105) [3]. -

Modes of Cell Death Induced by Photodynamic Therapy Using Zinc Phthalocyanine in Lung Cancer Cells Grown As a Monolayer and Three-Dimensional Multicellular Spheroids
molecules Article Modes of Cell Death Induced by Photodynamic Therapy Using Zinc Phthalocyanine in Lung Cancer Cells Grown as a Monolayer and Three-Dimensional Multicellular Spheroids Sello L. Manoto, Nicolette Houreld, Natasha Hodgkinson and Heidi Abrahamse * Laser Research Centre, Faculty of Health Sciences, University of Johannesburg, P.O. Box 17011, Doornfontein 2028, South Africa; [email protected] (S.L.M.); [email protected] (N.H.); [email protected] (N.H.) * Correspondence: [email protected]; Tel.: +27-11-559-6406; Fax: +27-11-559-6884 Academic Editors: Augusto C. Tomé, João Paulo C. Tomé and Michael Hanack Received: 8 March 2017; Accepted: 9 May 2017; Published: 16 May 2017 Abstract: Photodynamic therapy (PDT) involves interaction of a photosensitizer, light, and molecular oxygen which produces singlet oxygen and subsequent tumour eradication. The development of second generation photosensitizers, such as phthalocyanines, has improved this technology. Customary monolayer cell culture techniques are, unfortunately, too simple to replicate treatment effects in vivo. Multicellular tumour spheroids may provide a better alternative since they mimic aspects of the human tumour environment. This study aimed to profile 84 genes involved in apoptosis following treatment with PDT on lung cancer cells (A549) grown in a monolayer versus three-dimensional multicellular tumour spheroids (250 and 500 µm). Gene expression profiling 2 was performed 24 h post irradiation (680 nm; 5 J/cm ) with zinc sulfophthalocyanine (ZnPcSmix) to determine the genes involved in apoptotic cell death. In the monolayer cells, eight pro-apoptotic genes were upregulated, and two were downregulated. In the multicellular tumour spheroids (250 µm) there was upregulation of only 1 gene while there was downregulation of 56 genes. -

A Four-Gene Decision Tree Signature
Published OnlineFirst October 10, 2018; DOI: 10.1158/1535-7163.MCT-18-0243 Companion Diagnostic, Pharmacogenomic, and Cancer Biomarkers Molecular Cancer Therapeutics A Four-gene Decision Tree Signature Classification of Triple-negative Breast Cancer: Implications for Targeted Therapeutics Jelmar Quist1,2,3, Hasan Mirza1,2,3, Maggie C.U. Cheang4,5, Melinda L. Telli6, Joyce A. O'Shaughnessy7, Christopher J. Lord5,8, Andrew N.J. Tutt2,3,5, and Anita Grigoriadis1,2,3 Abstract The molecular complexity of triple-negative breast cancers in early diagnosed TNBCs. In TNBC patients with metastatic (TNBCs) provides a challenge for patient management. We set disease, the MC6 proportion was reduced to 25%, and was out to characterize this heterogeneous disease by combining independently associated with a higher response rate to transcriptomics and genomics data, with the aim of revealing platinum-based chemotherapy. In TNBC cell line data, convergent pathway dependencies with the potential for treat- platinum sensitivity was recapitulated, and a sensitivity ment intervention. A Bayesian algorithm was used to integrate to the inhibition of the phosphatase PPM1D was revealed. molecular profiles in two TNBC cohorts, followed by valida- Molecularly, MC6-TNBCs displayed high levels of telomeric þ þ tion using five independent cohorts (n ¼ 1,168), including allelic imbalances, enrichment of CD4 and CD8 immune three clinical trials. A four-gene decision tree signature was signatures, and reduced expression of genes negatively reg- identified, which robustly classified TNBCs into six subtypes. ulating the MAPK signaling pathway. These observations All four genes in the signature (EXO1, TP53BP2, FOXM1, and suggest that our integrative classification approach may RSU1) are associated with either genomic instability, malig- identify TNBC patients with discernible and theoretically nant growth, or treatment response. -

Rna-Sequencing Applications: Gene Expression Quantification and Methylator Phenotype Identification
The Texas Medical Center Library DigitalCommons@TMC The University of Texas MD Anderson Cancer Center UTHealth Graduate School of The University of Texas MD Anderson Cancer Biomedical Sciences Dissertations and Theses Center UTHealth Graduate School of (Open Access) Biomedical Sciences 8-2013 RNA-SEQUENCING APPLICATIONS: GENE EXPRESSION QUANTIFICATION AND METHYLATOR PHENOTYPE IDENTIFICATION Guoshuai Cai Follow this and additional works at: https://digitalcommons.library.tmc.edu/utgsbs_dissertations Part of the Bioinformatics Commons, Computational Biology Commons, and the Medicine and Health Sciences Commons Recommended Citation Cai, Guoshuai, "RNA-SEQUENCING APPLICATIONS: GENE EXPRESSION QUANTIFICATION AND METHYLATOR PHENOTYPE IDENTIFICATION" (2013). The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences Dissertations and Theses (Open Access). 386. https://digitalcommons.library.tmc.edu/utgsbs_dissertations/386 This Dissertation (PhD) is brought to you for free and open access by the The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences at DigitalCommons@TMC. It has been accepted for inclusion in The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences Dissertations and Theses (Open Access) by an authorized administrator of DigitalCommons@TMC. For more information, please contact [email protected]. RNA-SEQUENCING APPLICATIONS: GENE EXPRESSION QUANTIFICATION AND METHYLATOR PHENOTYPE IDENTIFICATION -

Gene Section Review
Atlas of Genetics and Cytogenetics in Oncology and Haematology OPEN ACCESS JOURNAL AT INIST-CNRS Gene Section Review TP53BP2 (tumor protein p53 binding protein, 2) Kathryn Van Hook, Zhiping Wang, Charles Lopez Department of Medicine, Division of Hematology and Medical Oncology, Oregon Health and Science University, Portland, OR, USA (KV, ZW, CL) Published in Atlas Database: January 2011 Online updated version : http://AtlasGeneticsOncology.org/Genes/TP53BP2ID42667ch1q42.html DOI: 10.4267/2042/46004 This work is licensed under a Creative Commons Attribution-Noncommercial-No Derivative Works 2.0 France Licence. © 2011 Atlas of Genetics and Cytogenetics in Oncology and Haematology Identity Pseudogene Not known. Other names: 53BP2; ASPP2; BBP; P53BP2; PPP1R13A Protein HGNC (Hugo): TP53BP2 Location: 1q41 Description ASPP2 is a pro-apoptotic protein with a predicted size DNA/RNA of approximately 135 kDa. It is the founding member of a family of ASPP proteins that all share the common Description motifs of four Ankyrin-repeats, a Src-homology 3 The TP53BP2 gene spans about 66 kb on chromosome (SH3) domain, and a Polyproline domain in their C- 1q42.1 on the minus strand (Yang et al., 1997). There terminus (Iwabuchi et al., 1994). The N-terminus of are two transcripts as a result of alternative splicing ASPP2 is thought to be important for regulating its (Takahashi et al., 2004). The transcript variant 1, which apoptotic function and contains a putative Ras- is shorter (4670 bp), does not contain exon 3 and gives association domain as well as a ubiquitin-like fold rise to a longer form of the protein named TP53BPL (Tidow et al., 2007). -

A Graph-Theoretic Approach to Model Genomic Data and Identify Biological Modules Asscociated with Cancer Outcomes
A Graph-Theoretic Approach to Model Genomic Data and Identify Biological Modules Asscociated with Cancer Outcomes Deanna Petrochilos A dissertation presented in partial fulfillment of the requirements for the degree of Doctor of Philosophy University of Washington 2013 Reading Committee: Neil Abernethy, Chair John Gennari, Ali Shojaie Program Authorized to Offer Degree: Biomedical Informatics and Health Education UMI Number: 3588836 All rights reserved INFORMATION TO ALL USERS The quality of this reproduction is dependent upon the quality of the copy submitted. In the unlikely event that the author did not send a complete manuscript and there are missing pages, these will be noted. Also, if material had to be removed, a note will indicate the deletion. UMI 3588836 Published by ProQuest LLC (2013). Copyright in the Dissertation held by the Author. Microform Edition © ProQuest LLC. All rights reserved. This work is protected against unauthorized copying under Title 17, United States Code ProQuest LLC. 789 East Eisenhower Parkway P.O. Box 1346 Ann Arbor, MI 48106 - 1346 ©Copyright 2013 Deanna Petrochilos University of Washington Abstract Using Graph-Based Methods to Integrate and Analyze Cancer Genomic Data Deanna Petrochilos Chair of the Supervisory Committee: Assistant Professor Neil Abernethy Biomedical Informatics and Health Education Studies of the genetic basis of complex disease present statistical and methodological challenges in the discovery of reliable and high-confidence genes that reveal biological phenomena underlying the etiology of disease or gene signatures prognostic of disease outcomes. This dissertation examines the capacity of graph-theoretical methods to model and analyze genomic information and thus facilitate using prior knowledge to create a more discrete and functionally relevant feature space. -

Kampa 2009 Cell Cycle.Pdf
EXTRA VIEW EXTRA VIEW Cell Cycle 8:18, 2871-2876; September 15, 2009; © 2009 Landes Bioscience New insights into the expanding complexity of the tumor suppressor ASPP2 Kerstin M. Kampa,1 Michael Bonin2 and Charles D. Lopez3,4,* 1Medizinische Universitätsklinik; Department of Hematology, Oncology, Rheumatology, Immunology and Pulmology; Universität Tübingen; Tübingen, Ger- many; 2Microarray Facility Tübingen; Universität Tübingen; Tübingen, Germany; 3Department of Medicine; Division of Hematology and Medical Oncology; Oregon Health and Science University; Portland, OR USA; 4The OHSU Knight Cancer Institute; Portland, OR USA poptosis Stimulating Protein of resolution by Gorina and Pavletich.4 Most A p53-2, ASPP2, aka 53BP2L, interestingly, the p53 “hotspot” muta- (encoded by TP53BP2) is a pro-apoptotic tions found in human cancers disrupt the member of a family of p53 binding pro- 53BP2/p53 interaction4-6—suggesting teins. ASPP2 expression is frequently sup- that this interaction plays an important pressed in human cancers and numerous role in p53 biology. Armed with this struc- studies have consistently demonstrated tural knowledge, the great challenge over that ASPP2 inhibits cell growth as well the last 15 years has been to understand as stimulates apoptosis—at least in part the function(s) of ASPP2 (and family through a p53-mediated pathway. Two members). independent mouse models have shown In 1996, during a yeast two-hybrid that ASPP2 is a haplo-insufficient tumor screen using Bcl-2 as bait, Naumovski suppressor and underscore the impor- and Cleary found that 53BP2 was actu- tance of the role of ASPP2 in human ally a partial clone of a transcript encod- cancer. -

Apoptosis-Stimulating Protein of P53 (ASPP2) Heterozygous Mice Are Tumor-Prone and Have Attenuated Cellular Damage–Response Thresholds
Apoptosis-stimulating protein of p53 (ASPP2) heterozygous mice are tumor-prone and have attenuated cellular damage–response thresholds Kerstin M. Kampaa,b, Jared D. Acobaa, Dexi Chena, Joel Gaya, Hunjoo Leea, Kelly Beemera, Emerson Padiernosa, Nataya Boonmarkc, Zhiyi Zhua, Alice C. Fanc, Alexis S. Baileya,d, William H. Fleminga,d, Christopher Corlessa, Dean W. Felsherc, Louie Naumovskic,1, and Charles D. Lopeza,2 aDepartment of Medicine, Division of Hematology and Medical Oncology, Oregon Health and Science University, Portland, OR 97239; bMedizinsche Universita¨tsklinik, Department of Hematology, Oncology, Rheumatology, Immunology, and Pulmology, Universita¨t Tu¨ bingen, Tu¨bingen 72076, Germany; cDepartments of Medicine and Pediatrics, Division of Hematology and Oncology, Stanford University, Stanford, CA 94305; and dOregon Stem Cell Center, Portland, OR 97239 Edited by Brian J. Druker, Oregon Health and Science University, Portland, OR, and approved December 30, 2008 (received for review September 11, 2008) The expression of ASPP2 (53BP2L), a proapoptotic member of a In this report, we targeted the ASPP2 allele in a mouse by family of p53-binding proteins, is frequently suppressed in many using homologous recombination to explore the in vivo con- human cancers. Accumulating evidence suggests that ASPP2 inhib- sequences of attenuated ASPP2 expression. We demonstrate its tumor growth; however, the mechanisms by which ASPP2 that reduced ASPP2 expression in heterozygous mice results suppresses tumor formation remain to be clarified. To study this, in: (i) an increased incidence of a variety of spontaneous we targeted the ASPP2 allele in a mouse by replacing exons 10–17 tumors, (ii) the accelerated formation of high-grade thymic T -with a neoR gene. -

Powerful Inhibition of Experimental Human Pancreatic Cancers by Receptor Targeted Cytotoxic LH-RH Analog AEZS-108
www.impactjournals.com/oncotarget/ Oncotarget, May, Vol.4, No 5 Powerful Inhibition of Experimental Human Pancreatic Cancers by Receptor Targeted Cytotoxic LH-RH analog AEZS-108 Karoly Szepeshazi1,2, Andrew V. Schally1,2,3,4,5, Norman L. Block1,2,3,4, Gabor Halmos1,2,3,6, Mehrdad Nadji3, Luca Szalontay1,2, Irving Vidaurre1,2, Andrew Abi- Chaker1,2,3, Ferenc G. Rick1,2,7 1 Veterans Affairs Medical Center, Miami, FL 2 South Florida VA Foundation for Research and Education, Miami, FL 3 Department of Pathology, University of Miami, Miller School of Medicine, Miami, FL 4 Division of Hematology/Oncology, University of Miami, Miller School of Medicine, Miami, FL 5 Division of Endocrinology, Department of Medicine, University of Miami, Miller School of Medicine, Miami, FL 6 Department of Biopharmacy, School of Pharmacy, University of Debrecen, Hungary; 7 Department of Urology, Florida International University, Herbert Wertheim College of Medicine, Miami, FL Correspondence to: Andrew V Schally, email: [email protected] Correspondence to: Ferenc G. Rick, email: [email protected] Keywords: pancreatic carcinoma, targeted therapy, LH-RH receptor, cytotoxic LHRH analog, GnRH, doxorubicin, peptide therapy Received: May 20, 2013 Accepted: June 1, 2013 Published: June 3, 2013 This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT: Pancreatic carcinoma is one of the cancers with the worse prognosis, thus any therapeutic improvement is imperative. Cytotoxic LH-RH analog, AN-152 (proprietary designation, AEZS-108), consisting of doxorubicin (DOX) conjugated to D-Lys6LH-RH, is now in clinical trials for targeted therapy of several sex hormone-dependent tumors that express LH-RH receptors. -

Bioinformatics Mining of the Dark Matter Proteome For
BIOINFORMATICS MINING OF THE DARK MATTER PROTEOME FOR CANCER TARGETS DISCOVERY by Ana Paula Delgado A Thesis Submitted to the Faculty of The Charles E. Schmidt College of Science In Partial Fulfillment of the Requirements for the Degree of Master of Science Florida Atlantic University Boca Raton, Florida May 2015 Copyright 2015 by Ana Paula Delgado ii ACKNOWLEDGEMENTS I would first like to thank Dr. Narayanan for his continuous encouragement, guidance, and support during the past two years of my graduate education. It has truly been an unforgettable experience working in his laboratory. I also want to express gratitude to my external advisor Professor Van de Ven from the University of Leuven, Belgium for his constant involvement and assistance on my project. Moreover, I would like to thank Dr. Binninger and Dr. Dawson-Scully for their advice and for agreeing to serve on my thesis committee. I also thank provost Dr. Perry for his involvement in my project. I thank Jeanine Narayanan for editorial assistance with the publications and with this dissertation. It has been a pleasure working with various undergraduate students some of whom became lab mates including Pamela Brandao, Maria Julia Chapado and Sheilin Hamid. I thank them for their expert help in the projects we were involved in. Lastly, I want to express my profound thanks to my parents and brother for their unconditional love, support and guidance over the last couple of years. They were my rock when I was in doubt and never let me give up. I would also like to thank my boyfriend Spencer Daniel and best friends for being part of an incredible support system.