And Mushroom-Associated Alkaloids from Two Behavior Modifying
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
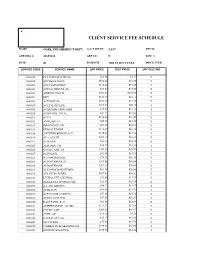
Oakland Sheriff's Dept Fee Schedule 02.09
CLIENT SERVICE FEE SCHEDULE NAME: OAKLAND SHERIFF'S DEPT ACCT SETUP: 2/1/87 REP ID: GRP BILL #: 22857610 GRP CD: N DISC %: FS ID: 01 FS DESCR: 2003 CLIENT FEES PRICE TYPE: SERVICE CODE SERVICE NAME UPF PRICE TEST PRICE UPF DISC IND 0000018 PLT SODIUM CITRATE $12.86 $4.37 Y 0000022 APC RESISTANCE $101.62 $34.55 Y 0000201 ACETAMINOPHEN $110.24 $37.48 Y 0000202 ACETALDEHYDE (B) $73.00 $73.00 N 0000203 ARSENIC,TISSUE $302.00 $302.00 N 0000204 DHT $182.70 $62.12 Y 0000205 ACETONE (B) $102.20 $34.75 Y 0000206 ACETYLCHOLINE $165.00 $56.10 Y 0000208 ACID PHOS, PROS, IMM $35.60 $12.10 Y 0000210 ACID PHOS, TOTAL $37.22 $12.65 Y 0000211 ACTH $110.00 $37.40 Y 0000212 AMYLASE (U) $45.45 $15.45 Y 0000213 IMMUNOFIX, UR $67.39 $22.91 Y 0000214 ETHOSUXIMIDE $112.07 $38.10 Y 0000216 ANTITHROMBIN III ACT $170.46 $57.96 Y 0000219 ALA, QUANT $107.10 $36.41 Y 0000223 ALBUMIN $22.88 $22.88 N 0000224 ALBUMIN, CSF $36.25 $12.33 Y 0000225 CYCLIC AMP, UR $205.80 $69.97 Y 0000227 ALDOLASE $32.08 $10.91 Y 0000228 A-2-MACROGLOB. $78.75 $26.78 Y 0000229 ALDOSTERONE (U) $251.58 $85.54 Y 0000230 ALDOSTERONE $207.10 $70.41 Y 0000231 ALK PHOS ISOENZYMES $61.34 $20.86 Y 0000232 AFP, FLUID W/RFX $107.40 $36.52 Y 0000233 LEUKOCYTE ALK PHOS $34.60 $11.76 Y 0000234 ALKALINE PHOSPHATASE $22.88 $22.88 N 0000235 A-1-ANTITRYPSN $40.22 $13.67 Y 0000236 AMIKACIN $99.91 $33.97 Y 0000237 AFP,TUMOR (CHIRON) $53.86 $18.31 Y 0000238 AMINO ACID SCR $87.15 $29.63 Y 0000240 RAST PANEL #103 $62.00 $62.00 N 0000241 AMPHETAMINE GC/MS $111.70 $37.98 Y 0000242 CYCLIC AMP $205.80 $69.97 Y 0000243 AMYLASE $16.42 $5.58 Y 0000248 HACKBERRY IGE $35.40 $35.40 N 0000249 ANA W/RFX $55.00 $18.70 Y 0000250 *AMIKACIN,PEAK&TROUGH $199.83 $67.94 Y 0000251 ANDROSTENEDIONE $180.65 $61.42 Y 0000252 ANTI-DIUR. -

Endophytic Fungi: Treasure for Anti-Cancerous Compounds
International Journal of Pharmacy and Pharmaceutical Sciences ISSN- 0975-1491 Vol 8, Issue 8, 2016 Review Article ENDOPHYTIC FUNGI: TREASURE FOR ANTI-CANCEROUS COMPOUNDS ANAND DILIP FIRODIYAa*, RAJESH KUMAR TENGURIAb aCSRD, Peoples University, Bhopal 462037, Madhya Pradesh, India, bDepartment of Botany, Govt. PG College, Rajgarh 496551, Madhya Pradesh, India Email: [email protected] Received: 22 Apr 2016 Revised and Accepted: 20 June 2016 ABSTRACT Endophytic fungi that live asymptomatically inside the plant tissues have novel bioactive metabolites exhibiting a variety of biological activities, especially against cancer. This review highlights the research progress on the production of anticancer compounds by endophytic fungi from 1990- 2015. Anticancer activity is generally associated with the cytotoxicity of the compounds present in the endophytic fungi. The ubiquitous nature of endophytic fungi synthesise diverse chemicals with promising anticancer activity from either their original host or related species. Modification in fermentation parameters and genetic insight of endophytes may produce novel anti-cancerous compounds. Keywords: Cancer, Medicinal plants, Secondary metabolites © 2016 The Authors. Published by Innovare Academic Sciences Pvt Ltd. This is an open access article under the CC BY license (http://creativecommons.org/licenses/by/4.0/) INTRODUCTION endophytic fungi detectable by high-performance liquid chromate- graphy, nuclear magnetic resonance, mass spectrophotometer and The interest in the biogenic medicines has revived throughout the X-ray crystallography and its cytotoxicity of the bioactive world, as the increase in awareness of the health hazards and compounds against cancer cell lines. The compounds with potential toxicity associated with the random use of synthetic drugs and application were also considered in the selection of antitumor antibiotics [1]. -

Evidence for Paternal Leakage in Hybrid Periodical Cicadas (Hemiptera: Magicicada Spp.) Kathryn M
Evidence for Paternal Leakage in Hybrid Periodical Cicadas (Hemiptera: Magicicada spp.) Kathryn M. Fontaine¤, John R. Cooley*, Chris Simon Department of Ecology and Evolutionary Biology, University of Connecticut, Storrs, Connecticut, United States of America Mitochondrial inheritance is generally assumed to be maternal. However, there is increasing evidence of exceptions to this rule, especially in hybrid crosses. In these cases, mitochondria are also inherited paternally, so ‘‘paternal leakage’’ of mitochondria occurs. It is important to understand these exceptions better, since they potentially complicate or invalidate studies that make use of mitochondrial markers. We surveyed F1 offspring of experimental hybrid crosses of the 17-year periodical cicadas Magicicada septendecim, M. septendecula, and M. cassini for the presence of paternal mitochondrial markers at various times during development (1-day eggs; 3-, 6-, 9-week eggs; 16-month old 1st and 2nd instar nymphs). We found evidence of paternal leakage in both reciprocal hybrid crosses in all of these samples. The relative difficulty of detecting paternal mtDNA in the youngest eggs and ease of detecting leakage in older eggs and in nymphs suggests that paternal mitochondria proliferate as the eggs develop. Our data support recent theoretical predictions that paternal leakage may be more common than previously estimated. Citation: Fontaine KM, Cooley JR, Simon C (2007) Evidence for Paternal Leakage in Hybrid Periodical Cicadas (Hemiptera: Magicicada spp.). PLoS ONE 2(9): e892. doi:10.1371/journal.pone.0000892 INTRODUCTION The seven currently-recognized 13- and 17-year periodical Although mitochondrial DNA (mtDNA) exhibits a variety of cicada species (Magicicada septendecim {17}, M. tredecim {13}, M. -
Fungal Endophytes As Efficient Sources of Plant-Derived Bioactive
microorganisms Review Fungal Endophytes as Efficient Sources of Plant-Derived Bioactive Compounds and Their Prospective Applications in Natural Product Drug Discovery: Insights, Avenues, and Challenges Archana Singh 1,2, Dheeraj K. Singh 3,* , Ravindra N. Kharwar 2,* , James F. White 4,* and Surendra K. Gond 1,* 1 Department of Botany, MMV, Banaras Hindu University, Varanasi 221005, India; [email protected] 2 Department of Botany, Institute of Science, Banaras Hindu University, Varanasi 221005, India 3 Department of Botany, Harish Chandra Post Graduate College, Varanasi 221001, India 4 Department of Plant Biology, Rutgers University, New Brunswick, NJ 08901, USA * Correspondence: [email protected] (D.K.S.); [email protected] (R.N.K.); [email protected] (J.F.W.); [email protected] (S.K.G.) Abstract: Fungal endophytes are well-established sources of biologically active natural compounds with many producing pharmacologically valuable specific plant-derived products. This review details typical plant-derived medicinal compounds of several classes, including alkaloids, coumarins, flavonoids, glycosides, lignans, phenylpropanoids, quinones, saponins, terpenoids, and xanthones that are produced by endophytic fungi. This review covers the studies carried out since the first report of taxol biosynthesis by endophytic Taxomyces andreanae in 1993 up to mid-2020. The article also highlights the prospects of endophyte-dependent biosynthesis of such plant-derived pharma- cologically active compounds and the bottlenecks in the commercialization of this novel approach Citation: Singh, A.; Singh, D.K.; Kharwar, R.N.; White, J.F.; Gond, S.K. in the area of drug discovery. After recent updates in the field of ‘omics’ and ‘one strain many Fungal Endophytes as Efficient compounds’ (OSMAC) approach, fungal endophytes have emerged as strong unconventional source Sources of Plant-Derived Bioactive of such prized products. -

The Evolutionary Relationships of 17-Year and 13-Year Cicadas, and Three New Species (Hornoptera, Cicadidae, Magicicada)
MISCELLANEOUS PUBLICATIONS MUSEUM OF ZOOLOGY, UNIVERSITY OF MICHIGAN, NO. 121 The Evolutionary Relationships of 17-Year and 13-Year Cicadas, and Three New Species (Hornoptera, Cicadidae, Magicicada) BY KlCHAKD D. ALEXANDEK AND THOMAS E. MOORE ANN ARBOR MUSEUM OF ZOOLOGY, UNIVERSITY OF MICHIGAN JULY 24, 1962 MISCELLANEOUS PUBLICATIONS MUSEUM OF ZOOLOGY, UNIVERSITY OF MICHIGAN The publications of the Museum of Zoology, University of Michigan, consist of two series-the Occasional Papers and the Miscellaneous Publications. Both series were founded by Dr. Bryant Walker, Mr. Bradsliaw H. Swales, and Dr. W. W. Newcomb. The Occasional Papers, publication d which was begun in 1913, serve as a medium for original studies based principally upon the collections in the Museum. They are issued separately. When a sufficient number of pages has been printed to make a volume. a title page, table of contents, and an index are supplied to libraries and indi- viduals on the mailing list for the series. The Miscellaneous Publications, which include papers on field and museum tech- niques, monographic studies, and other contributions not within the scope of the Occasional Papers, are published separately. It is not intended that they be grouped into volumes. Each number has a title page and, when necessary, a table of contents. A complete list of publications on Birds, Fishes, Insects, Mammals, Mollusks, and Reptiles and Amphibians is available. Address inquiries to the Director, Museum of Zoology, Ann Arbor, Michigan. No. 1. Directions for collecting and preserving specimens of dragonflies for museum purposes. By E. B. WILLIAMSON.(1916) 15 pp., 3 figs. .......... $0.25 No. -

The Effects of a Periodic Cicada Emergence on Forest Birds and the Ecology of Cerulean Warblers at Big Oaks National Wildlife Refuge, Indiana
University of Tennessee, Knoxville TRACE: Tennessee Research and Creative Exchange Masters Theses Graduate School 8-2006 The Effects of a Periodic Cicada Emergence on Forest Birds and the Ecology of Cerulean Warblers at Big Oaks National Wildlife Refuge, Indiana Dustin W. Varble University of Tennessee, Knoxville Follow this and additional works at: https://trace.tennessee.edu/utk_gradthes Part of the Animal Sciences Commons Recommended Citation Varble, Dustin W., "The Effects of a Periodic Cicada Emergence on Forest Birds and the Ecology of Cerulean Warblers at Big Oaks National Wildlife Refuge, Indiana. " Master's Thesis, University of Tennessee, 2006. https://trace.tennessee.edu/utk_gradthes/4519 This Thesis is brought to you for free and open access by the Graduate School at TRACE: Tennessee Research and Creative Exchange. It has been accepted for inclusion in Masters Theses by an authorized administrator of TRACE: Tennessee Research and Creative Exchange. For more information, please contact [email protected]. To the Graduate Council: I am submitting herewith a thesis written by Dustin W. Varble entitled "The Effects of a Periodic Cicada Emergence on Forest Birds and the Ecology of Cerulean Warblers at Big Oaks National Wildlife Refuge, Indiana." I have examined the final electronic copy of this thesis for form and content and recommend that it be accepted in partial fulfillment of the equirr ements for the degree of Master of Science, with a major in Wildlife and Fisheries Science. David A. Buehler, Major Professor We have read this thesis and recommend its acceptance: Frank Van Manen, Roger Tankersley Jr. Joseph R. Robb Accepted for the Council: Carolyn R. -

Fungal Endophytes: Promising Tools for Pharmaceutical Science
Int. J. Pharm. Sci. Rev. Res., 25(2), Mar – Apr 2014; Article No. 25, Pages: 128-138 ISSN 0976 – 044X Research Article Fungal Endophytes: Promising Tools for Pharmaceutical Science Pramod Kumar Pandey1,2, Siddhartha Singh1, Raj Naraian Singh Yadav2, Amit Kumar Singh1*, M. Chandra Kumar Singh1 1Department of Basic Science and Humanities, College of Horticulture and Forestry, Central Agricultural University, Pasighat, Arunachal Pradesh, India. 2Centre for Studies in Biotechnology, Dibrugarh University, Dibrugarh, Assam, India. *Corresponding author’s E-mail: [email protected] Accepted on: 27-01-2014; Finalized on: 31-03-2014. ABSTRACT Fungal endophytes are microorganisms that internally infect living plant tissues without causing any visible symptom of infection, and live in mutualistic relationship with plants for at least a part of their life cycle. Every plant in the world is reservoir of one or more number of endophytes. In recent year’s special attention have been made towards endophytic fungi because of their ability to synthesize several novel bioactive compounds not previously known to biological system which are important for pharmaceutical, agricultural and industrial sector. This review describes information on endophyte diversity, as well as production of secondary metabolites with special emphasis on anti cancerous, antimicrobial, antiviral, antibiotics along with huge number of other secondary metabolites for commercial exploitation in pharmaceutical and medical field. Furthermore, the chemical potential of endophytic fungi for drug discovery will be discussed with focus on the detection of pharmaceutically valuable plant constituents as products of fungal biosynthesis in recent years. At present a huge world population is suffering from the problem caused by drug resistant microbes (bacteria, parasitic protozoans and fungus) which decreases the efficiency of synthetic drugs. -

COMPARATIVE BIOLOGICAL EVALUATION of SIX ENDOPHYTIC FUNGI ISOLATED from VINCA ROSEA LEAVES Ahmed M
Az. J. Pharm Sci. Vol. 59, March, 2019. 137 COMPARATIVE BIOLOGICAL EVALUATION OF SIX ENDOPHYTIC FUNGI ISOLATED FROM VINCA ROSEA LEAVES Ahmed M. Metwaly*1, Mohamed A. Ashour1, Shabana Khan2, Guoyi Ma2, Hazem A. Kadry1, Atef A. El-Hela1, Abd-Elsalam I. Mohammad1, Stephen J. Cutler3 and Samir A. Ross2 1Department of Pharmacognosy, Faculty of Pharmacy, Al-Azhar University, Cairo, Egypt 2 Department of Biomolecular Science, National Center for Natural Products Research, University of Mississippi, Oxford, MS 38677, USA 3Department of Medicinal Chemistry, School of Pharmacy, the University of Mississippi, Oxford, MS 38677, USA *Corresponding authors: [email protected] Abstract In this study, a total of six endophytic fungi have been isolated from Vinca rosea (Apocynaceae) leaves growing in Egypt. The isolated fungi were identified morphologically and microscopically up to species to be; Alternaria phragmospora, Aspergillus awamori, Penicillium duclauxii, Penicillium melinii, Nigrospora sphaerica and Mucor ramosissimus. The extracts of the all identified fungi were screened biologically for antileukemic, cytotoxic, antimalarial, antileishmanial, antimicrobial, antioxidant and anti-inflammatory activities as well as for cannabinoid and opioid receptor binding affinities. Out of the six examined fungal extracts four exhibited promising antimalarial activities, four showed modeate antileukemic activities and two of them exhibited cytotoxic activities, three showed antioxidant activities, three exhibited anti-inflammatory activities, and the only one showed an antifungal activity. Alternaria phragmospora was the most active antimalarial agent inhibiting Plasmodium falciparum D6 and W2 clones with IC50 values of 1.9 and 2.1 μg/mL, respectively. Alternaria phragmospora extract was subjected to liquid-liquid partition using 90% MeOH, hexane then BuOH and H2O. -

Isolation and Characterization of Fungal Endophytes Isolated from Medicinal Plant Ephedra Pachyclada As Plant Growth-Promoting
biomolecules Article Isolation and Characterization of Fungal Endophytes Isolated from Medicinal Plant Ephedra pachyclada as Plant Growth-Promoting Ahmed Mohamed Aly Khalil 1,2, Saad El-Din Hassan 1,* , Sultan M. Alsharif 3, Ahmed M. Eid 1 , Emad El-Din Ewais 1, Ehab Azab 4,5 , Adil A. Gobouri 6, Amr Elkelish 7 and Amr Fouda 1,* 1 Department of Botany and Microbiology, Faculty of Science, Al-Azhar University, Nasr City, Cairo 11884, Egypt; [email protected] (A.M.A.K.); [email protected] (A.M.E.); [email protected] (E.E.-D.E.) 2 Biology Department, College of Science, Taibah University, Yanbu 41911, Saudi Arabia 3 Biology Department, Faculty of Science, Taibah University, Al Madinah P.O. Box 887, Saudi Arabia; [email protected] 4 Department of Biotechnology, College of Science, Taif University, P.O. Box 11099, Taif 21944, Saudi Arabia; [email protected] 5 Botany and Microbiology Department, Faculty of Science, Zagazig University, Zagazig 44519, Sharkia, Egypt 6 Department of Chemistry, College of Science, Taif University, P.O. Box 11099, Taif 21944, Saudi Arabia; [email protected] 7 Botany Department, Faculty of Science, Suez Canal University, Ismailia 41522, Egypt; [email protected] * Correspondence: [email protected] (S.E.-D.H.); [email protected] (A.F.); Citation: Khalil, A.M.A.; Tel.: +20-102-3884804 (S.E.-D.H.); +20-111-3351244 (A.F.) Hassan, S.E.-D.; Alsharif, S.M.; Eid, A.M.; Ewais, E.E.-D.; Azab, E.; Abstract: Endophytic fungi are widely present in internal plant tissues and provide different ben- Gobouri, A.A.; Elkelish, A.; Fouda, A. -

And 17-Y Life Cycles Among Three Periodical Cicada Lineages
Independent divergence of 13- and 17-y life cycles SEE COMMENTARY among three periodical cicada lineages Teiji Sotaa,1, Satoshi Yamamotob, John R. Cooleyc, Kathy B. R. Hillc, Chris Simonc, and Jin Yoshimurad,e,f aDepartment of Zoology, Graduate School of Science, Kyoto University, Sakyo, Kyoto 606-8502, Japan; bGraduate School of Global Environmental Studies, Kyoto University, Sakyo, Kyoto 606-8501, Japan; cDepartment of Ecology and Evolutionary Biology, University of Connecticut, Storrs, CT 06268-3043; dDepartment of Systems Engineering, Shizuoka University, Hamamatsu 432-8561, Japan; eMarine Biosystems Research Center, Chiba University, Kamogawa 299-5502, Chiba, Japan; and fDepartment of Environmental and Forest Biology, State University of New York College of Environmental Science and Forestry, Syracuse, NY 13210 Edited by Mary Jane West-Eberhard, Smithsonian Tropical Research Institute, Ciudad Universitaria, Costa Rica, and approved February 22, 2013 (received for review November 16, 2012) The evolution of 13- and 17-y periodical cicadas (Magicicada)is group (6, 14–18), and only rough divergence times among species enigmatic because at any given location, up to three distinct spe- have been inferred (8, 10). Thus, a comprehensive molecular cies groups (Decim, Cassini, Decula) with synchronized life cycles phylogeny covering all of the extant broods, their phylogeography, are involved. Each species group is divided into one 13- and one and divergence time has been lacking until now. 17-y species with the exception of the Decim group, which con- We conducted molecular phylogenetic and population genetic tains two 13-y species—13-y species are Magicicada tredecim, analyses by using nuclear and mitochondrial DNA markers for Magicicada neotredecim, Magicicada tredecassini, and Magicicada samples collected over a 30-y period (1978–2008). -

RSC COFI Prelims 1..4
The Chemistry of Fungi James R. Hanson Department of Chemistry, University of Sussex, Brighton, UK ISBN: 978-0-85404-136-7 A catalogue record for this book is available from the British Library r James R. Hanson, 2008 All rights reserved Apart from fair dealing for the purposes of research for non-commercial purposes or for private study, criticism or review, as permitted under the Copyright, Designs and Patents Act 1988 and the Copyright and Related Rights Regulations 2003, this publication may not be reproduced, stored or transmitted, in any form or by any means, without the prior permission in writing of The Royal Society of Chemistry or the copyright owner, or in the case of reproduction in accordance with the terms of licences issued by the Copyright Licensing Agency in the UK, or in accordance with the terms of the licences issued by the appropriate Reproduction Rights Organization outside the UK. Enquiries concerning reproduction outside the terms stated here should be sent to The Royal Society of Chemistry at the address printed on this page. Published by The Royal Society of Chemistry, Thomas Graham House, Science Park, Milton Road, Cambridge CB4 0WF, UK Registered Charity Number 207890 For further information see our web site at www.rsc.org Preface The diverse structures, biosyntheses and biological activities of fungal meta- bolites have attracted chemists for many years. This book is an introduction to the chemistry of fungal metabolites. The aim is to illustrate, within the context of fungal metabolites, the historical progression from chemical to spectroscopic methods of structure elucidation, the development in biosynthetic studies from establishing sequences and mechanisms to chemical enzymology and genetics and the increasing understanding of the biological roles of natural products. -

The Complete Genome Sequences of Two Species Of
F1000Research 2021, 10:215 Last updated: 09 SEP 2021 DATA NOTE The complete genome sequences of two species of seventeen- year cicadas: Magicicada septendecim and Magicicada septendecula [version 1; peer review: 2 approved with reservations] Harold B. White1, Stacy Pirro 2 1Department of Chemistry and Biochemistry, University of Delaware, Delaware, USA 2Department Biodiversity, Iridian Genomes, Bethesda, MD, 20817, USA v1 First published: 16 Mar 2021, 10:215 Open Peer Review https://doi.org/10.12688/f1000research.27309.1 Latest published: 16 Mar 2021, 10:215 https://doi.org/10.12688/f1000research.27309.1 Reviewer Status Invited Reviewers Abstract The genus Magicicada (Hemiptera: Cicadidae) includes the periodical 1 2 cicadas of Eastern North America. Spending the majority of their long lives underground, the adult cicadas emerge every 13 or 17 years to version 1 spend 4-6 weeks as adult to mate. We present the whole genome 16 Mar 2021 report report sequences of two species of 17-year cicadas, Magicicada septendecim and Magicicada septendecula. The reads were assembled by a de novo 1. Hu Li , China Agricultural University, method followed by alignments to related species. Annotation was performed by GeneMark-ES. The raw and assembled data is available Beijing, China via NCBI Short Read Archive and Assembly databases. 2. Shuai Zhan, Chinese Academy of Sciences, Keywords Shanghai, China Genome, assembly, arthropoda, insecta, hemiptera Any reports and responses or comments on the article can be found at the end of the article. This article is included in the Genome Sequencing gateway. Page 1 of 6 F1000Research 2021, 10:215 Last updated: 09 SEP 2021 Corresponding author: Stacy Pirro ([email protected]) Author roles: White HB: Resources; Pirro S: Data Curation, Investigation, Resources Competing interests: No competing interests were disclosed.