Oxygen and RNA in Stress-Induced Mutation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Phosphatidylinositol-3-Kinase Related Kinases (Pikks) in Radiation-Induced Dna Damage
Mil. Med. Sci. Lett. (Voj. Zdrav. Listy) 2012, vol. 81(4), p. 177-187 ISSN 0372-7025 DOI: 10.31482/mmsl.2012.025 REVIEW ARTICLE PHOSPHATIDYLINOSITOL-3-KINASE RELATED KINASES (PIKKS) IN RADIATION-INDUCED DNA DAMAGE Ales Tichy 1, Kamila Durisova 1, Eva Novotna 1, Lenka Zarybnicka 1, Jirina Vavrova 1, Jaroslav Pejchal 2, Zuzana Sinkorova 1 1 Department of Radiobiology, Faculty of Health Sciences in Hradec Králové, University of Defence in Brno, Czech Republic 2 Centrum of Advanced Studies, Faculty of Health Sciences in Hradec Králové, University of Defence in Brno, Czech Republic. Received 5 th September 2012. Revised 27 th November 2012. Published 7 th December 2012. Summary This review describes a drug target for cancer therapy, family of phosphatidylinositol-3 kinase related kinases (PIKKs), and it gives a comprehensive review of recent information. Besides general information about phosphatidylinositol-3 kinase superfamily, it characterizes a DNA-damage response pathway since it is monitored by PIKKs. Key words: PIKKs; ATM; ATR; DNA-PK; Ionising radiation; DNA-repair ABBREVIATIONS therapy and radiation play a pivotal role. Since cancer is one of the leading causes of death worldwide, it is DSB - double stand breaks, reasonable to invest time and resources in the enligh - IR - ionising radiation, tening of mechanisms, which underlie radio-resis - p53 - TP53 tumour suppressors, tance. PI - phosphatidylinositol. The aim of this review is to describe the family INTRODUCTION of phosphatidyinositol 3-kinases (PI3K) and its func - tional subgroup - phosphatidylinositol-3-kinase rela - An efficient cancer treatment means to restore ted kinases (PIKKs) and their relation to repairing of controlled tissue growth via interfering with cell sig - radiation-induced DNA damage. -

Collision with Duplex DNA Renders Escherichia Coli DNA Polymerase III
www.nature.com/scientificreports OPEN Collision with duplex DNA renders Escherichia coli DNA polymerase III holoenzyme susceptible to Received: 8 May 2017 Accepted: 18 September 2017 DNA polymerase IV-mediated Published: xx xx xxxx polymerase switching on the sliding clamp Thanh Thi Le, Asako Furukohri, Masahiro Tatsumi-Akiyama & Hisaji Maki Organisms possess multiple DNA polymerases (Pols) and use each for a different purpose. One of the five Pols inEscherichia coli, DNA polymerase IV (Pol IV), encoded by the dinB gene, is known to participate in lesion bypass at certain DNA adducts. To understand how cells choose Pols when the replication fork encounters an obstacle on template DNA, the process of polymerase exchange from the primary replicative enzyme DNA polymerase III (Pol III) to Pol IV was studied in vitro. Replicating Pol III forming a tight holoenzyme (Pol III HE) with the sliding clamp was challenged by Pol IV on a primed ssDNA template carrying a short inverted repeat. A rapid and lesion-independent switch from Pol III to Pol IV occurred when Pol III HE encountered a hairpin stem duplex, implying that the loss of Pol III-ssDNA contact induces switching to Pol IV. Supporting this idea, mutant Pol III with an increased affinity for ssDNA was more resistant to Pol IV than wild-type Pol III was. We observed that an exchange between Pol III and Pol IV also occurred when Pol III HE collided with primer/template duplex. Our data suggest that Pol III-ssDNA interaction may modulate the susceptibility of Pol III HE to Pol IV-mediated polymerase exchange. -

The Microbiota-Produced N-Formyl Peptide Fmlf Promotes Obesity-Induced Glucose
Page 1 of 230 Diabetes Title: The microbiota-produced N-formyl peptide fMLF promotes obesity-induced glucose intolerance Joshua Wollam1, Matthew Riopel1, Yong-Jiang Xu1,2, Andrew M. F. Johnson1, Jachelle M. Ofrecio1, Wei Ying1, Dalila El Ouarrat1, Luisa S. Chan3, Andrew W. Han3, Nadir A. Mahmood3, Caitlin N. Ryan3, Yun Sok Lee1, Jeramie D. Watrous1,2, Mahendra D. Chordia4, Dongfeng Pan4, Mohit Jain1,2, Jerrold M. Olefsky1 * Affiliations: 1 Division of Endocrinology & Metabolism, Department of Medicine, University of California, San Diego, La Jolla, California, USA. 2 Department of Pharmacology, University of California, San Diego, La Jolla, California, USA. 3 Second Genome, Inc., South San Francisco, California, USA. 4 Department of Radiology and Medical Imaging, University of Virginia, Charlottesville, VA, USA. * Correspondence to: 858-534-2230, [email protected] Word Count: 4749 Figures: 6 Supplemental Figures: 11 Supplemental Tables: 5 1 Diabetes Publish Ahead of Print, published online April 22, 2019 Diabetes Page 2 of 230 ABSTRACT The composition of the gastrointestinal (GI) microbiota and associated metabolites changes dramatically with diet and the development of obesity. Although many correlations have been described, specific mechanistic links between these changes and glucose homeostasis remain to be defined. Here we show that blood and intestinal levels of the microbiota-produced N-formyl peptide, formyl-methionyl-leucyl-phenylalanine (fMLF), are elevated in high fat diet (HFD)- induced obese mice. Genetic or pharmacological inhibition of the N-formyl peptide receptor Fpr1 leads to increased insulin levels and improved glucose tolerance, dependent upon glucagon- like peptide-1 (GLP-1). Obese Fpr1-knockout (Fpr1-KO) mice also display an altered microbiome, exemplifying the dynamic relationship between host metabolism and microbiota. -

Supplementary Information
Supplementary information (a) (b) Figure S1. Resistant (a) and sensitive (b) gene scores plotted against subsystems involved in cell regulation. The small circles represent the individual hits and the large circles represent the mean of each subsystem. Each individual score signifies the mean of 12 trials – three biological and four technical. The p-value was calculated as a two-tailed t-test and significance was determined using the Benjamini-Hochberg procedure; false discovery rate was selected to be 0.1. Plots constructed using Pathway Tools, Omics Dashboard. Figure S2. Connectivity map displaying the predicted functional associations between the silver-resistant gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S3. Connectivity map displaying the predicted functional associations between the silver-sensitive gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S4. Metabolic overview of the pathways in Escherichia coli. The pathways involved in silver-resistance are coloured according to respective normalized score. Each individual score represents the mean of 12 trials – three biological and four technical. Amino acid – upward pointing triangle, carbohydrate – square, proteins – diamond, purines – vertical ellipse, cofactor – downward pointing triangle, tRNA – tee, and other – circle. -
Q 297 Suppl USE
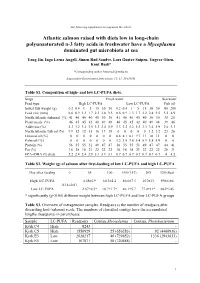
The following supplement accompanies the article Atlantic salmon raised with diets low in long-chain polyunsaturated n-3 fatty acids in freshwater have a Mycoplasma dominated gut microbiota at sea Yang Jin, Inga Leena Angell, Simen Rød Sandve, Lars Gustav Snipen, Yngvar Olsen, Knut Rudi* *Corresponding author: [email protected] Aquaculture Environment Interactions 11: 31–39 (2019) Table S1. Composition of high- and low LC-PUFA diets. Stage Fresh water Sea water Feed type High LC-PUFA Low LC-PUFA Fish oil Initial fish weight (g) 0.2 0.4 1 5 15 30 50 0.2 0.4 1 5 15 30 50 80 200 Feed size (mm) 0.6 0.9 1.3 1.7 2.2 2.8 3.5 0.6 0.9 1.3 1.7 2.2 2.8 3.5 3.5 4.9 North Atlantic fishmeal (%) 41 40 40 40 40 30 30 41 40 40 40 40 30 30 35 25 Plant meals (%) 46 45 45 42 40 49 48 46 45 45 42 40 49 48 39 46 Additives (%) 3.3 3.2 3.2 3.5 3.3 3.4 3.9 3.3 3.2 3.2 3.5 3.3 3.4 3.9 2.6 3.3 North Atlantic fish oil (%) 9.9 12 12 15 16 17 18 0 0 0 0 0 1.2 1.2 23 26 Linseed oil (%) 0 0 0 0 0 0 0 6.8 8.1 8.1 9.7 11 10 11 0 0 Palm oil (%) 0 0 0 0 0 0 0 3.2 3.8 3.8 5.4 5.9 5.8 5.9 0 0 Protein (%) 56 55 55 51 49 47 47 56 55 55 51 49 47 47 44 41 Fat (%) 16 18 18 21 22 22 22 16 18 18 21 22 22 22 28 31 EPA+DHA (% diet) 2.2 2.4 2.4 2.9 3.1 3.1 3.1 0.7 0.7 0.7 0.7 0.7 0.7 0.7 4 4.2 Table S2. -

Consolidated List of Up-Regulated Proteins Expressed at Different Cr (VI) Concentrations at Time Points
Electronic Supplementary Material (ESI) for Metallomics. -

Specificity of DNA Damage Inducible DNA Polymerase IV from Escherichia Coli
Specificity of DNA Damage Inducible DNA Polymerase IV from Escherichia coli A thesis presented by Jason M. Walsh to The Department of Chemistry and Chemical Biology in partial fulfillment of the requirements for the degree of Master of Science in the field of Chemistry Northeastern University Boston, Massachusetts December 2009 1 © 2009, Jason M Walsh All rights reserved. 2 Specificity of DNA Damage Inducible DNA Polymerase IV from Escherichia coli by Jason M. Walsh ABSTRACT OF THESIS Submitted in partial fulfillment of the requirements for the degree of Master of Science in Chemistry and Chemical Biology in the Graduate School of Arts and Sciences of Northeastern University, December 2009 3 ABSTRACT DNA polymerases are responsible for DNA replication during cell division. There are multiple families of polymerases (A, B, C, D, X) responsible for copying DNA during replication and repair. There is also a class of polymerases conserved throughout evolution, known as the Y family polymerases, that have reduced replication fidelity on undamaged DNA (Tang et al. 2000). However Y family DNA polymerases have the specialized property of replicating DNA by copying damaged DNA, a process known as translesion synthesis (TLS). Structural differences between Y family and replicative polymerases may account for the difference in enzymatic activity. However we demonstrate that the Klenow fragment (A family) can bypass a fluorescent cytosine analog known as 1, 3-diaza-2-oxophenothiazine (tC), that DinB, a Y family polymerase, cannot bypass. We show that DinB inserts dGTP faithfully, but cannot extend the DNA primer beyond that. Verifying which amino acid residues are responsible for both function and specificity of the Y family polymerases is accomplished by assessing the kinetic data of nucleotide incorporation events of DinB variants as compared to wild-type DinB. -

Supplementary Figure S1 Fluorescent Substrate Digestion in the Droplet
α-amylase Schematic image of the BODIPY-Starch Fig.1A Droplet with a better α- Droplet with a α-amylase Droplet with non-α- amylase producing cell producing cell amylase producing cell Supplementary Figure S1 Fluorescent substrate digestion in the droplet. Droplet with a better α-amylase producing cell will have higher α-amylase concentration inside the droplet and thus more BODIPY-starch can be degraded in the same unit of time. This will give rise to a stronger fluorescent signal that will be emitted upon excitation. A B Treshold: Treshold: 0.4 0.4 Sorted fraction: Sorted fraction: 0.24% 0.76% C D Threshold: Thresholds: 0.35‐0.37 0.5‐0.55 Sorted fraction: Sorted fraction 1% ≈3% Supplementary Figure S2 Histogram showing droplet fluorescence of yeast libraries in the second round of screening. (A) The Library MH34-11. (B) The Library MH34-14. (C) The Library MH34-8. (D) The Library MH23-11. Starting cells Sorted Waste cells cells MH34-11 MH34-14 MH34-8 MH23-11 Supplementary Figure S3 Fermentation of mixed colonies for evaluation of the titer and yield of α-amylase produced by different yeast populations before and after sorting. 50 colonies were randomly picked up from different populations of the library and mixed together for fermentation. A I I+IV:38% II III:60% III IV B Sorted Waste Sorted Waste All I Supplementary Figure S4 (A) 50 waste colonies from the library MH34-11 were tested in tube fermentation. (B) Average α-amylase titer and yield of sorted cells and waste cells from the library MH34-11. -

Involvement of Escherichia Coli DNA Polymerase IV in Tolerance of Cytotoxic Alkylating DNA Lesions in Vivo
Copyright Ó 2007 by the Genetics Society of America DOI: 10.1534/genetics.107.072405 Involvement of Escherichia coli DNA Polymerase IV in Tolerance of Cytotoxic Alkylating DNA Lesions in Vivo Ivana Bjedov,1 Chitralekha Nag Dasgupta,1 Dea Slade, Sophie Le Blastier, Marjorie Selva and Ivan Matic2 INSERM U571, Faculte´ de Me´decine, Universite´ Paris 5, 75730 Paris Cedex 15, France Manuscript received February 20, 2007 Accepted for publication May 3, 2007 ABSTRACT Escherichia coli PolIV, a DNA polymerase capable of catalyzing synthesis past replication-blocking DNA lesions, belongs to the most ubiquitous branch of Y-family DNA polymerases. The goal of this study is to identify spontaneous DNA damage that is bypassed specifically and accurately by PolIV in vivo. We in- creased the amount of spontaneous DNA lesions using mutants deficient for different DNA repair path- ways and measured mutation frequency in PolIV-proficient and -deficient backgrounds. We found that PolIV performs an error-free bypass of DNA damage that accumulates in the alkA tag genetic background. This result indicates that PolIV is involved in the error-free bypass of cytotoxic alkylating DNA lesions. When the amount of cytotoxic alkylating DNA lesions is increased by the treatment with chemical alkyl- ating agents, PolIV is required for survival in an alkA tag-proficient genetic background as well. Our study, together with the reported involvement of the mammalian PolIV homolog, Polk, in similar activity, indicates that Y-family DNA polymerases from the DinB branch can be added to the list of evolutionarily conserved molecular mechanisms that counteract cytotoxic effects of DNA alkylation. -

The Yeast Saccharomyces Cerevisiae DNA Polymerase IV: Possible Involvement in Double Strand Break DNA Repair
Q-DI 1994 Oxford University Press Nucleic Acids Research, 1994, Vol. 22, No. 15 3011 -3017 The yeast Saccharomyces cerevisiae DNA polymerase IV: possible involvement in double strand break DNA repair Sun-Hee Leem, Philip A.Ropp1 and Akio Sugino* Department of Molecular Immunology, Research Institute for Microbial Diseases, Osaka University, 3-1 Yamada-Oka, Suita, Osaka 565, Japan and 'Laboratory of Molecular Genetics, National Institute of Environmental Health Sciences, National Institutes of Health, PO Box 12233, Research Triangle Park, NC 27709, USA Received May 13, 1994; Revised and Accepted July 13, 1994 ABSTRACT We identified and purified a new DNA polymerase (DNA division-cycle (cdc) mutants, among which were alleles of the polymerase IV), which is similar to mammalian DNA genes now known to encode DNA polymerase a and 6 catalytic polymerase (, from Saccharomyces cerevisiae and polypeptides (7-10). DNA polymerase e is the DNA polymerase suggested that it is encoded by YCR14C (POLX) on that was last identified and purified in eukaryotes (1, 2). But, chromosome Ill. Here, we provided a direct evidence its yeast homolog, DNA polymerase II, was detected that the purified DNA polymerase IV is indeed encoded enzymatically in yeast cell extracts in early 1970 (11, 12). Its by POLX. Strains harboring a poI4 deletion mutation gene, POL2, was not found in any genetic screen, nor was its exhibit neither mitotic growth defect nor a meiosis mammalian counterpart identified in the SV40 in vitro system. defect, suggesting that DNA polymerase IV participates POL2 was eventually cloned by reverse genetics and has been in nonessential functions in DNA metabolism. -

New Insights Into Marine Group III Euryarchaeota, from Dark to Light
The ISME Journal (2017), 1–16 © 2017 International Society for Microbial Ecology All rights reserved 1751-7362/17 www.nature.com/ismej ORIGINAL ARTICLE New insights into marine group III Euryarchaeota, from dark to light Jose M Haro-Moreno1,3, Francisco Rodriguez-Valera1, Purificación López-García2, David Moreira2 and Ana-Belen Martin-Cuadrado1,3 1Evolutionary Genomics Group, Departamento de Producción Vegetal y Microbiología, Universidad Miguel Hernández, Alicante, Spain and 2Unité d’Ecologie, Systématique et Evolution, UMR CNRS 8079, Université Paris-Sud, Orsay Cedex, France Marine Euryarchaeota remain among the least understood major components of marine microbial communities. Marine group II Euryarchaeota (MG-II) are more abundant in surface waters (4–20% of the total prokaryotic community), whereas marine group III Euryarchaeota (MG-III) are generally considered low-abundance members of deep mesopelagic and bathypelagic communities. Using genome assembly from direct metagenome reads and metagenomic fosmid clones, we have identified six novel MG-III genome sequence bins from the photic zone (Epi1–6) and two novel bins from deep-sea samples (Bathy1–2). Genome completeness in those genome bins varies from 44% to 85%. Photic-zone MG-III bins corresponded to novel groups with no similarity, and significantly lower GC content, when compared with previously described deep-MG-III genome bins. As found in many other epipelagic microorganisms, photic-zone MG-III bins contained numerous photolyase and rhodopsin genes, as well as genes for peptide and lipid uptake and degradation, suggesting a photoheterotrophic lifestyle. Phylogenetic analysis of these photolyases and rhodopsins as well as their genomic context suggests that these genes are of bacterial origin, supporting the hypothesis of an MG-III ancestor that lived in the dark ocean. -

Eukaryotic DNA Polymerases in Homologous Recombination Mitch Mcvey, Varandt Y
GE50CH18-Heyer ARI 28 October 2016 10:25 ANNUAL REVIEWS Further Click here to view this article's online features: • Download figures as PPT slides • Navigate linked references • Download citations Eukaryotic DNA Polymerases • Explore related articles • Search keywords in Homologous Recombination Mitch McVey,1 Varandt Y. Khodaverdian,1 Damon Meyer,2,4 Paula Gonc¸alves Cerqueira,2 and Wolf-Dietrich Heyer2,3 1Department of Biology, Tufts University, Medford, Massachusetts 02155; email: [email protected] 2Department of Microbiology and Molecular Genetics, University of California, Davis, California 95616; email: [email protected] 3Department of Molecular and Cellular Biology, University of California, Davis, California 95616 4College of Health Sciences, California Northstate University, Rancho Cordova, California 95670 Annu. Rev. Genet. 2016. 50:393–421 Keywords The Annual Review of Genetics is online at DNA synthesis, genome stability, mutagenesis, template switching genet.annualreviews.org This article’s doi: Abstract 10.1146/annurev-genet-120215-035243 Homologous recombination (HR) is a central process to ensure genomic Copyright c 2016 by Annual Reviews. stability in somatic cells and during meiosis. HR-associated DNA synthe- All rights reserved Annu. Rev. Genet. 2016.50:393-421. Downloaded from www.annualreviews.org sis determines in large part the fidelity of the process. A number of recent Access provided by University of California - Davis on 11/30/16. For personal use only. studies have demonstrated that DNA synthesis during HR is conservative, less processive, and more mutagenic than replicative DNA synthesis. In this review, we describe mechanistic features of DNA synthesis during different types of HR-mediated DNA repair, including synthesis-dependent strand annealing, break-induced replication, and meiotic recombination.