Computational Prediction of Protein-Protein Interactions On
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Protein Identities in Evs Isolated from U87-MG GBM Cells As Determined by NG LC-MS/MS
Protein identities in EVs isolated from U87-MG GBM cells as determined by NG LC-MS/MS. No. Accession Description Σ Coverage Σ# Proteins Σ# Unique Peptides Σ# Peptides Σ# PSMs # AAs MW [kDa] calc. pI 1 A8MS94 Putative golgin subfamily A member 2-like protein 5 OS=Homo sapiens PE=5 SV=2 - [GG2L5_HUMAN] 100 1 1 7 88 110 12,03704523 5,681152344 2 P60660 Myosin light polypeptide 6 OS=Homo sapiens GN=MYL6 PE=1 SV=2 - [MYL6_HUMAN] 100 3 5 17 173 151 16,91913397 4,652832031 3 Q6ZYL4 General transcription factor IIH subunit 5 OS=Homo sapiens GN=GTF2H5 PE=1 SV=1 - [TF2H5_HUMAN] 98,59 1 1 4 13 71 8,048185945 4,652832031 4 P60709 Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 - [ACTB_HUMAN] 97,6 5 5 35 917 375 41,70973209 5,478027344 5 P13489 Ribonuclease inhibitor OS=Homo sapiens GN=RNH1 PE=1 SV=2 - [RINI_HUMAN] 96,75 1 12 37 173 461 49,94108966 4,817871094 6 P09382 Galectin-1 OS=Homo sapiens GN=LGALS1 PE=1 SV=2 - [LEG1_HUMAN] 96,3 1 7 14 283 135 14,70620005 5,503417969 7 P60174 Triosephosphate isomerase OS=Homo sapiens GN=TPI1 PE=1 SV=3 - [TPIS_HUMAN] 95,1 3 16 25 375 286 30,77169764 5,922363281 8 P04406 Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 - [G3P_HUMAN] 94,63 2 13 31 509 335 36,03039959 8,455566406 9 Q15185 Prostaglandin E synthase 3 OS=Homo sapiens GN=PTGES3 PE=1 SV=1 - [TEBP_HUMAN] 93,13 1 5 12 74 160 18,68541938 4,538574219 10 P09417 Dihydropteridine reductase OS=Homo sapiens GN=QDPR PE=1 SV=2 - [DHPR_HUMAN] 93,03 1 1 17 69 244 25,77302971 7,371582031 11 P01911 HLA class II histocompatibility antigen, -

Bppart and Bpmax: RNA-RNA Interaction Partition Function and Structure Prediction for the Base Pair Counting Model
BPPart and BPMax: RNA-RNA Interaction Partition Function and Structure Prediction for the Base Pair Counting Model Ali Ebrahimpour-Boroojeny, Sanjay Rajopadhye, and Hamidreza Chitsaz ∗ Department of Computer Science, Colorado State University Abstract A few elite classes of RNA-RNA interaction (RRI), with complex roles in cellular functions such as miRNA-target and lncRNAs in human health, have already been studied. Accordingly, RRI bioinfor- matics tools tailored for those elite classes have been proposed in the last decade. Interestingly, there are somewhat unnoticed mRNA-mRNA interactions in the literature with potentially drastic biological roles. Hence, there is a need for high-throughput generic RRI bioinformatics tools. We revisit our RRI partition function algorithm, piRNA, which happens to be the most comprehensive and computationally-intensive thermodynamic model for RRI. We propose simpler models that are shown to retain the vast majority of the thermodynamic information that piRNA captures. We simplify the energy model and instead consider only weighted base pair counting to obtain BPPart for Base-pair Partition function and BPMax for Base-pair Maximization which are 225 and 1350 faster ◦ × × than piRNA, with a correlation of 0.855 and 0.836 with piRNA at 37 C on 50,500 experimentally charac- terized RRIs. This correlation increases to 0.920 and 0.904, respectively, at 180◦C. − Finally, we apply our algorithm BPPart to discover two disease-related RNAs, SNORD3D and TRAF3, and hypothesize their potential roles in Parkinson's disease and Cerebral Autosomal Dominant Arteri- opathy with Subcortical Infarcts and Leukoencephalopathy (CADASIL). 1 Introduction Since mid 1990s with the advent of RNA interference discovery, RNA-RNA interaction (RRI) has moved to the spotlight in modern, post-genome biology. -

The Function of NM23-H1/NME1 and Its Homologs in Major Processes Linked to Metastasis
University of Dundee The Function of NM23-H1/NME1 and Its Homologs in Major Processes Linked to Metastasis Mátyási, Barbara; Farkas, Zsolt; Kopper, László; Sebestyén, Anna; Boissan, Mathieu; Mehta, Anil Published in: Pathology and Oncology Research DOI: 10.1007/s12253-020-00797-0 Publication date: 2020 Licence: CC BY Document Version Publisher's PDF, also known as Version of record Link to publication in Discovery Research Portal Citation for published version (APA): Mátyási, B., Farkas, Z., Kopper, L., Sebestyén, A., Boissan, M., Mehta, A., & Takács-Vellai, K. (2020). The Function of NM23-H1/NME1 and Its Homologs in Major Processes Linked to Metastasis. Pathology and Oncology Research, 26(1), 49-61. https://doi.org/10.1007/s12253-020-00797-0 General rights Copyright and moral rights for the publications made accessible in Discovery Research Portal are retained by the authors and/or other copyright owners and it is a condition of accessing publications that users recognise and abide by the legal requirements associated with these rights. • Users may download and print one copy of any publication from Discovery Research Portal for the purpose of private study or research. • You may not further distribute the material or use it for any profit-making activity or commercial gain. • You may freely distribute the URL identifying the publication in the public portal. Take down policy If you believe that this document breaches copyright please contact us providing details, and we will remove access to the work immediately and investigate your -

Identification of Novel Chemotherapeutic Strategies For
www.nature.com/scientificreports OPEN Identification of novel chemotherapeutic strategies for metastatic uveal melanoma Received: 17 November 2016 Paolo Fagone1, Rosario Caltabiano2, Andrea Russo3, Gabriella Lupo1, Accepted: 09 February 2017 Carmelina Daniela Anfuso1, Maria Sofia Basile1, Antonio Longo3, Ferdinando Nicoletti1, Published: 17 March 2017 Rocco De Pasquale4, Massimo Libra1 & Michele Reibaldi3 Melanoma of the uveal tract accounts for approximately 5% of all melanomas and represents the most common primary intraocular malignancy. Despite improvements in diagnosis and more effective local therapies for primary cancer, the rate of metastatic death has not changed in the past forty years. In the present study, we made use of bioinformatics to analyze the data obtained from three public available microarray datasets on uveal melanoma in an attempt to identify novel putative chemotherapeutic options for the liver metastatic disease. We have first carried out a meta-analysis of publicly available whole-genome datasets, that included data from 132 patients, comparing metastatic vs. non metastatic uveal melanomas, in order to identify the most relevant genes characterizing the spreading of tumor to the liver. Subsequently, the L1000CDS2 web-based utility was used to predict small molecules and drugs targeting the metastatic uveal melanoma gene signature. The most promising drugs were found to be Cinnarizine, an anti-histaminic drug used for motion sickness, Digitoxigenin, a precursor of cardiac glycosides, and Clofazimine, a fat-soluble iminophenazine used in leprosy. In vitro and in vivo validation studies will be needed to confirm the efficacy of these molecules for the prevention and treatment of metastatic uveal melanoma. Uveal melanoma is the most common primary intraocular cancer, and after the skin, the uveal tract is the second most common location for melanoma1. -

Development and Validation of a Protein-Based Risk Score for Cardiovascular Outcomes Among Patients with Stable Coronary Heart Disease
Supplementary Online Content Ganz P, Heidecker B, Hveem K, et al. Development and validation of a protein-based risk score for cardiovascular outcomes among patients with stable coronary heart disease. JAMA. doi: 10.1001/jama.2016.5951 eTable 1. List of 1130 Proteins Measured by Somalogic’s Modified Aptamer-Based Proteomic Assay eTable 2. Coefficients for Weibull Recalibration Model Applied to 9-Protein Model eFigure 1. Median Protein Levels in Derivation and Validation Cohort eTable 3. Coefficients for the Recalibration Model Applied to Refit Framingham eFigure 2. Calibration Plots for the Refit Framingham Model eTable 4. List of 200 Proteins Associated With the Risk of MI, Stroke, Heart Failure, and Death eFigure 3. Hazard Ratios of Lasso Selected Proteins for Primary End Point of MI, Stroke, Heart Failure, and Death eFigure 4. 9-Protein Prognostic Model Hazard Ratios Adjusted for Framingham Variables eFigure 5. 9-Protein Risk Scores by Event Type This supplementary material has been provided by the authors to give readers additional information about their work. Downloaded From: https://jamanetwork.com/ on 10/02/2021 Supplemental Material Table of Contents 1 Study Design and Data Processing ......................................................................................................... 3 2 Table of 1130 Proteins Measured .......................................................................................................... 4 3 Variable Selection and Statistical Modeling ........................................................................................ -

Assessing the Human Canonical Protein Count[Version 1; Peer Review
F1000Research 2017, 6:448 Last updated: 15 JUL 2020 REVIEW Last rolls of the yoyo: Assessing the human canonical protein count [version 1; peer review: 1 approved, 2 approved with reservations] Christopher Southan IUPHAR/BPS Guide to Pharmacology, Centre for Integrative Physiology, University of Edinburgh, Edinburgh, EH8 9XD, UK First published: 07 Apr 2017, 6:448 Open Peer Review v1 https://doi.org/10.12688/f1000research.11119.1 Latest published: 07 Apr 2017, 6:448 https://doi.org/10.12688/f1000research.11119.1 Reviewer Status Abstract Invited Reviewers In 2004, when the protein estimate from the finished human genome was 1 2 3 only 24,000, the surprise was compounded as reviewed estimates fell to 19,000 by 2014. However, variability in the total canonical protein counts version 1 (i.e. excluding alternative splice forms) of open reading frames (ORFs) in 07 Apr 2017 report report report different annotation portals persists. This work assesses these differences and possible causes. A 16-year analysis of Ensembl and UniProtKB/Swiss-Prot shows convergence to a protein number of ~20,000. The former had shown some yo-yoing, but both have now plateaued. Nine 1 Michael Tress, Spanish National Cancer major annotation portals, reviewed at the beginning of 2017, gave a spread Research Centre (CNIO), Madrid, Spain of counts from 21,819 down to 18,891. The 4-way cross-reference concordance (within UniProt) between Ensembl, Swiss-Prot, Entrez Gene 2 Elspeth A. Bruford , European Molecular and the Human Gene Nomenclature Committee (HGNC) drops to 18,690, Biology Laboratory, Hinxton, UK indicating methodological differences in protein definitions and experimental existence support between sources. -
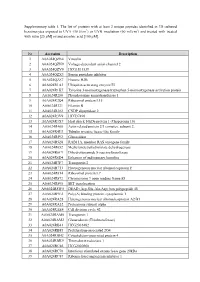
Supplementary Table 1. the List of Proteins with at Least 2 Unique
Supplementary table 1. The list of proteins with at least 2 unique peptides identified in 3D cultured keratinocytes exposed to UVA (30 J/cm2) or UVB irradiation (60 mJ/cm2) and treated with treated with rutin [25 µM] or/and ascorbic acid [100 µM]. Nr Accession Description 1 A0A024QZN4 Vinculin 2 A0A024QZN9 Voltage-dependent anion channel 2 3 A0A024QZV0 HCG1811539 4 A0A024QZX3 Serpin peptidase inhibitor 5 A0A024QZZ7 Histone H2B 6 A0A024R1A3 Ubiquitin-activating enzyme E1 7 A0A024R1K7 Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein 8 A0A024R280 Phosphoserine aminotransferase 1 9 A0A024R2Q4 Ribosomal protein L15 10 A0A024R321 Filamin B 11 A0A024R382 CNDP dipeptidase 2 12 A0A024R3V9 HCG37498 13 A0A024R3X7 Heat shock 10kDa protein 1 (Chaperonin 10) 14 A0A024R408 Actin related protein 2/3 complex, subunit 2, 15 A0A024R4U3 Tubulin tyrosine ligase-like family 16 A0A024R592 Glucosidase 17 A0A024R5Z8 RAB11A, member RAS oncogene family 18 A0A024R652 Methylenetetrahydrofolate dehydrogenase 19 A0A024R6C9 Dihydrolipoamide S-succinyltransferase 20 A0A024R6D4 Enhancer of rudimentary homolog 21 A0A024R7F7 Transportin 2 22 A0A024R7T3 Heterogeneous nuclear ribonucleoprotein F 23 A0A024R814 Ribosomal protein L7 24 A0A024R872 Chromosome 9 open reading frame 88 25 A0A024R895 SET translocation 26 A0A024R8W0 DEAD (Asp-Glu-Ala-Asp) box polypeptide 48 27 A0A024R9E2 Poly(A) binding protein, cytoplasmic 1 28 A0A024RA28 Heterogeneous nuclear ribonucleoprotein A2/B1 29 A0A024RA52 Proteasome subunit alpha 30 A0A024RAE4 Cell division cycle 42 31 -

Adenovirus Strategies for Altering the Cellular Environment in Favor of Infection
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations 2019 Adenovirus Strategies For Altering The Cellular Environment In Favor Of Infection Christin Herrmann University of Pennsylvania Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Allergy and Immunology Commons, Immunology and Infectious Disease Commons, Medical Immunology Commons, and the Virology Commons Recommended Citation Herrmann, Christin, "Adenovirus Strategies For Altering The Cellular Environment In Favor Of Infection" (2019). Publicly Accessible Penn Dissertations. 3568. https://repository.upenn.edu/edissertations/3568 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/3568 For more information, please contact [email protected]. Adenovirus Strategies For Altering The Cellular Environment In Favor Of Infection Abstract Viruses, as obligate intracellular pathogens, rely on their host cell for successful replication. Viruses have evolved different strategies to hijack and redirect cellular processes to benefit infection and overcome host immune responses. Understanding the mechanisms by which viruses exploit their host cells will reveal new targets for antiviral therapies. In addition, these studies can provide insights into the regulation of fundamental cellular processes. While much progress has been made in this area, many unexpected nuances of virus-host interaction are still being discovered. Here, we employed several strategies to uncover new aspects of viral manipulation of the host environment by adenovirus, a nuclear-replicating DNA virus that commonly infects humans. The first project focused on how viral histone-like proteins impact cellular chromatin. Adenovirus encodes the small, basic protein VII that coats and condenses viral genomes. The effect of this viral DNA-binding protein on host chromatin structure and function had remained unexplored. -

S41467-020-17157-W.Pdf
ARTICLE https://doi.org/10.1038/s41467-020-17157-w OPEN Transcriptional activity and strain-specific history of mouse pseudogenes Cristina Sisu 1,2,3,15, Paul Muir4,5,15, Adam Frankish6, Ian Fiddes7, Mark Diekhans 7, David Thybert6,8, Duncan T. Odom 9,10, Paul Flicek 6,10, Thomas M. Keane 6, Tim Hubbard 11, Jennifer Harrow12 & ✉ Mark Gerstein 1,2,5,13,14 Pseudogenes are ideal markers of genome remodelling. In turn, the mouse is an ideal plat- 1234567890():,; form for studying them, particularly with the recent availability of strain-sequencing and transcriptional data. Here, combining both manual curation and automatic pipelines, we present a genome-wide annotation of the pseudogenes in the mouse reference genome and 18 inbred mouse strains (available via the mouse.pseudogene.org resource). We also annotate 165 unitary pseudogenes in mouse, and 303, in human. The overall pseudogene repertoire in mouse is similar to that in human in terms of size, biotype distribution, and family composition (e.g. with GAPDH and ribosomal proteins being the largest families). Notable differences arise in the pseudogene age distribution, with multiple retro- transpositional bursts in mouse evolutionary history and only one in human. Furthermore, in each strain about a fifth of all pseudogenes are unique, reflecting strain-specific evolution. Finally, we find that ~15% of the mouse pseudogenes are transcribed, and that highly tran- scribed parent genes tend to give rise to many processed pseudogenes. 1 Program in Computational Biology and Bioinformatics, Yale University, New Haven, CT 06520, USA. 2 Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT 06520, USA. -

Assessment of Network Module Identification Across Complex Diseases
ANALYSIS https://doi.org/10.1038/s41592-019-0509-5 Assessment of network module identification across complex diseases Sarvenaz Choobdar1,2,20, Mehmet E. Ahsen3,117, Jake Crawford4,117, Mattia Tomasoni 1,2, Tao Fang5, David Lamparter1,2,6, Junyuan Lin7, Benjamin Hescott8, Xiaozhe Hu7, Johnathan Mercer9,10, Ted Natoli11, Rajiv Narayan11, The DREAM Module Identification Challenge Consortium12, Aravind Subramanian11, Jitao D. Zhang 5, Gustavo Stolovitzky 3,13, Zoltán Kutalik2,14, Kasper Lage 9,10,15, Donna K. Slonim 4,16, Julio Saez-Rodriguez 17,18, Lenore J. Cowen4,7, Sven Bergmann 1,2,19,21* and Daniel Marbach 1,2,5,21* Many bioinformatics methods have been proposed for reducing the complexity of large gene or protein networks into relevant subnetworks or modules. Yet, how such methods compare to each other in terms of their ability to identify disease-relevant modules in different types of network remains poorly understood. We launched the ‘Disease Module Identification DREAM Challenge’, an open competition to comprehensively assess module identification methods across diverse protein–protein interaction, signaling, gene co-expression, homology and cancer-gene networks. Predicted network modules were tested for association with complex traits and diseases using a unique collection of 180 genome-wide association studies. Our robust assessment of 75 module identification methods reveals top-performing algorithms, which recover complementary trait- associated modules. We find that most of these modules correspond to core disease-relevant pathways, which often com- prise therapeutic targets. This community challenge establishes biologically interpretable benchmarks, tools and guidelines for molecular network analysis to study human disease biology. omplex diseases involve many genes and molecules that inter- assessed on in silico generated benchmark graphs11. -

Downloaded Per Proteome Cohort Via the Web- Site Links of Table 1, Also Providing Information on the Deposited Spectral Datasets
www.nature.com/scientificreports OPEN Assessment of a complete and classifed platelet proteome from genome‑wide transcripts of human platelets and megakaryocytes covering platelet functions Jingnan Huang1,2*, Frauke Swieringa1,2,9, Fiorella A. Solari2,9, Isabella Provenzale1, Luigi Grassi3, Ilaria De Simone1, Constance C. F. M. J. Baaten1,4, Rachel Cavill5, Albert Sickmann2,6,7,9, Mattia Frontini3,8,9 & Johan W. M. Heemskerk1,9* Novel platelet and megakaryocyte transcriptome analysis allows prediction of the full or theoretical proteome of a representative human platelet. Here, we integrated the established platelet proteomes from six cohorts of healthy subjects, encompassing 5.2 k proteins, with two novel genome‑wide transcriptomes (57.8 k mRNAs). For 14.8 k protein‑coding transcripts, we assigned the proteins to 21 UniProt‑based classes, based on their preferential intracellular localization and presumed function. This classifed transcriptome‑proteome profle of platelets revealed: (i) Absence of 37.2 k genome‑ wide transcripts. (ii) High quantitative similarity of platelet and megakaryocyte transcriptomes (R = 0.75) for 14.8 k protein‑coding genes, but not for 3.8 k RNA genes or 1.9 k pseudogenes (R = 0.43–0.54), suggesting redistribution of mRNAs upon platelet shedding from megakaryocytes. (iii) Copy numbers of 3.5 k proteins that were restricted in size by the corresponding transcript levels (iv) Near complete coverage of identifed proteins in the relevant transcriptome (log2fpkm > 0.20) except for plasma‑derived secretory proteins, pointing to adhesion and uptake of such proteins. (v) Underrepresentation in the identifed proteome of nuclear‑related, membrane and signaling proteins, as well proteins with low‑level transcripts.