S41467-020-17157-W.Pdf
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Protein Identities in Evs Isolated from U87-MG GBM Cells As Determined by NG LC-MS/MS
Protein identities in EVs isolated from U87-MG GBM cells as determined by NG LC-MS/MS. No. Accession Description Σ Coverage Σ# Proteins Σ# Unique Peptides Σ# Peptides Σ# PSMs # AAs MW [kDa] calc. pI 1 A8MS94 Putative golgin subfamily A member 2-like protein 5 OS=Homo sapiens PE=5 SV=2 - [GG2L5_HUMAN] 100 1 1 7 88 110 12,03704523 5,681152344 2 P60660 Myosin light polypeptide 6 OS=Homo sapiens GN=MYL6 PE=1 SV=2 - [MYL6_HUMAN] 100 3 5 17 173 151 16,91913397 4,652832031 3 Q6ZYL4 General transcription factor IIH subunit 5 OS=Homo sapiens GN=GTF2H5 PE=1 SV=1 - [TF2H5_HUMAN] 98,59 1 1 4 13 71 8,048185945 4,652832031 4 P60709 Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 - [ACTB_HUMAN] 97,6 5 5 35 917 375 41,70973209 5,478027344 5 P13489 Ribonuclease inhibitor OS=Homo sapiens GN=RNH1 PE=1 SV=2 - [RINI_HUMAN] 96,75 1 12 37 173 461 49,94108966 4,817871094 6 P09382 Galectin-1 OS=Homo sapiens GN=LGALS1 PE=1 SV=2 - [LEG1_HUMAN] 96,3 1 7 14 283 135 14,70620005 5,503417969 7 P60174 Triosephosphate isomerase OS=Homo sapiens GN=TPI1 PE=1 SV=3 - [TPIS_HUMAN] 95,1 3 16 25 375 286 30,77169764 5,922363281 8 P04406 Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 - [G3P_HUMAN] 94,63 2 13 31 509 335 36,03039959 8,455566406 9 Q15185 Prostaglandin E synthase 3 OS=Homo sapiens GN=PTGES3 PE=1 SV=1 - [TEBP_HUMAN] 93,13 1 5 12 74 160 18,68541938 4,538574219 10 P09417 Dihydropteridine reductase OS=Homo sapiens GN=QDPR PE=1 SV=2 - [DHPR_HUMAN] 93,03 1 1 17 69 244 25,77302971 7,371582031 11 P01911 HLA class II histocompatibility antigen, -

Single Cell Mutational Profiling and Clonal Phylogeny in Cancer Nicola E
Downloaded from genome.cshlp.org on October 1, 2021 - Published by Cold Spring Harbor Laboratory Press For GENOME RESEARCH (Methods) Single cell mutational profiling and clonal phylogeny in cancer Nicola E Potter – Institute of Cancer Research, UK Luca Ermini – Institute of Cancer Research, UK Elli Papaemmanuil – Wellcome Trust Sanger Institute, UK Giovanni Cazzaniga - Clinica Pediatrica, Italy Gowri Vijayaraghavan – Institute of Cancer Research, UK Ian Titley – Institute of Cancer Research, UK Anthony Ford – Institute of Cancer Research, UK Peter Campbell - Wellcome Trust Sanger Institute, UK Lyndal Kearney – Institute of Cancer Research, UK Mel Greaves – Institute of Cancer Research, UK Corresponding Author - Professor Mel Greaves FRS, Head, Haemato-Oncology Research Unit, Division of Molecular Pathology, Brookes Lawley Building, 15 Cotswold Road, Sutton, Surrey SM2 5NG T +44 (0)20 8722 4073 (direct) E [email protected] Running title: Genetic analysis of single cancer cells Keywords: Heterogeneity, Single cells, Cancer, Phylogeny 1 Downloaded from genome.cshlp.org on October 1, 2021 - Published by Cold Spring Harbor Laboratory Press ABSTRACT The development of cancer is a dynamic evolutionary process in which intra-clonal, genetic diversity provides a substrate for clonal selection and a source of therapeutic escape. The complexity and topography of intra-clonal genetic architecture has major implications for biopsy-based prognosis and for targeted therapy. High depth, next generation sequencing (NGS) efficiently captures the mutational load of individual tumours or biopsies. But, being a snapshot portrait of total DNA, it disguises the fundamental features of sub-clonal variegation of genetic lesions and of clonal phylogeny. Single cell genetic profiling provides a potential resolution to this problem but methods developed to date all have limitations. -

Expression of the POTE Gene Family in Human Ovarian Cancer Carter J Barger1,2, Wa Zhang1,2, Ashok Sharma 1,2, Linda Chee1,2, Smitha R
www.nature.com/scientificreports OPEN Expression of the POTE gene family in human ovarian cancer Carter J Barger1,2, Wa Zhang1,2, Ashok Sharma 1,2, Linda Chee1,2, Smitha R. James3, Christina N. Kufel3, Austin Miller 4, Jane Meza5, Ronny Drapkin 6, Kunle Odunsi7,8,9, 2,10 1,2,3 Received: 5 July 2018 David Klinkebiel & Adam R. Karpf Accepted: 7 November 2018 The POTE family includes 14 genes in three phylogenetic groups. We determined POTE mRNA Published: xx xx xxxx expression in normal tissues, epithelial ovarian and high-grade serous ovarian cancer (EOC, HGSC), and pan-cancer, and determined the relationship of POTE expression to ovarian cancer clinicopathology. Groups 1 & 2 POTEs showed testis-specifc expression in normal tissues, consistent with assignment as cancer-testis antigens (CTAs), while Group 3 POTEs were expressed in several normal tissues, indicating they are not CTAs. Pan-POTE and individual POTEs showed signifcantly elevated expression in EOC and HGSC compared to normal controls. Pan-POTE correlated with increased stage, grade, and the HGSC subtype. Select individual POTEs showed increased expression in recurrent HGSC, and POTEE specifcally associated with reduced HGSC OS. Consistent with tumors, EOC cell lines had signifcantly elevated Pan-POTE compared to OSE and FTE cells. Notably, Group 1 & 2 POTEs (POTEs A/B/B2/C/D), Group 3 POTE-actin genes (POTEs E/F/I/J/KP), and other Group 3 POTEs (POTEs G/H/M) show within-group correlated expression, and pan-cancer analyses of tumors and cell lines confrmed this relationship. Based on their restricted expression in normal tissues and increased expression and association with poor prognosis in ovarian cancer, POTEs are potential oncogenes and therapeutic targets in this malignancy. -

The Function of NM23-H1/NME1 and Its Homologs in Major Processes Linked to Metastasis
University of Dundee The Function of NM23-H1/NME1 and Its Homologs in Major Processes Linked to Metastasis Mátyási, Barbara; Farkas, Zsolt; Kopper, László; Sebestyén, Anna; Boissan, Mathieu; Mehta, Anil Published in: Pathology and Oncology Research DOI: 10.1007/s12253-020-00797-0 Publication date: 2020 Licence: CC BY Document Version Publisher's PDF, also known as Version of record Link to publication in Discovery Research Portal Citation for published version (APA): Mátyási, B., Farkas, Z., Kopper, L., Sebestyén, A., Boissan, M., Mehta, A., & Takács-Vellai, K. (2020). The Function of NM23-H1/NME1 and Its Homologs in Major Processes Linked to Metastasis. Pathology and Oncology Research, 26(1), 49-61. https://doi.org/10.1007/s12253-020-00797-0 General rights Copyright and moral rights for the publications made accessible in Discovery Research Portal are retained by the authors and/or other copyright owners and it is a condition of accessing publications that users recognise and abide by the legal requirements associated with these rights. • Users may download and print one copy of any publication from Discovery Research Portal for the purpose of private study or research. • You may not further distribute the material or use it for any profit-making activity or commercial gain. • You may freely distribute the URL identifying the publication in the public portal. Take down policy If you believe that this document breaches copyright please contact us providing details, and we will remove access to the work immediately and investigate your -

Oncogene Associated Cdna Microarray Analysis Shows PRAME
J Clin Exp Hematopathol Vol. 49, No. 1, May 2009 Original Article Oncogene Associated cDNA Microarray Analysis Shows PRAME Gene Expression is a Marker for Response to Anthracycline Containing Chemotherapy in Patients with Diffuse Large B-cell Lymphoma Riko Kawano,1,3) Kennosuke Karube,3) Masahiro Kikuchi,2) Morishige Takeshita,2) Kazuo Tamura,1) Naokuni Uike,4) Tetsuya Eto,5) Koichi Ohshima,3) and Junji Suzumiya1,6) CHOP (cyclophosphamide, adriamycin, vincristine, and prednisolone) therapy achieves a response in more than 60% patients with diffuse large B-cell lymphomas (DLBCLs). However, DLBCL shows a heterogeneous response to chemotherapy, and some patients are refractory to CHOP therapy. This difference in response to therapy is most likely due to differences in biological characteristics. We used cDNA microarray analysis to identify genes differentially expressed in anthracycline containing chemotherapy-resistant DLBCLs (7 patients) compared with anthracycline containing chemotherapy-sensitive DLBCLs (6 patients). Nine genes on the cDNA chip showed increased expression in anthracycline containing chemotherapy- resistant patients. We chose the preferentially expressed antigen of melanoma (PRAME) gene because it showed the highest expression in anthracycline containing chemotherapy-resistant DLBCLs on the cDNA chip, and it has been linked to prognosis of hematological malignancies. We also examined the relationship between PRAME gene expression and progression-free survival (PFS) in 45 patients with DLBCL. The progression-free survival -

Short-Term Western-Style Diet Negatively Impacts Reproductive Outcomes in Primates
Short-term Western-style diet negatively impacts reproductive outcomes in primates Sweta Ravisankar, … , Shawn L. Chavez, Jon D. Hennebold JCI Insight. 2021;6(4):e138312. https://doi.org/10.1172/jci.insight.138312. Research Article Metabolism Reproductive biology A maternal Western-style diet (WSD) is associated with poor reproductive outcomes, but whether this is from the diet itself or underlying metabolic dysfunction is unknown. Here, we performed a longitudinal study using regularly cycling female rhesus macaques (n = 10) that underwent 2 consecutive in vitro fertilization (IVF) cycles, one while consuming a low-fat diet and another 6–8 months after consuming a high-fat WSD. Metabolic data were collected from the females prior to each IVF cycle. Follicular fluid (FF) and oocytes were assessed for cytokine/steroid levels and IVF potential, respectively. Although transition to a WSD led to weight gain and increased body fat, no difference in insulin levels was observed. A significant decrease in IL-1RA concentration and the ratio of cortisol/cortisone was detected in FF after WSD intake. Despite an increased probability of isolating mature oocytes, a 44% reduction in blastocyst number was observed with WSD consumption, and time-lapse imaging revealed delayed mitotic timing and multipolar divisions. RNA sequencing of blastocysts demonstrated dysregulation of genes involved in RNA binding, protein channel activity, mitochondrial function and pluripotency versus cell differentiation after WSD consumption. Thus, short-term WSD consumption promotes a proinflammatory intrafollicular microenvironment that is associated with impaired preimplantation development in the absence of large-scale metabolic changes. Find the latest version: https://jci.me/138312/pdf RESEARCH ARTICLE Short-term Western-style diet negatively impacts reproductive outcomes in primates Sweta Ravisankar,1,2 Alison Y. -

Development and Validation of a Protein-Based Risk Score for Cardiovascular Outcomes Among Patients with Stable Coronary Heart Disease
Supplementary Online Content Ganz P, Heidecker B, Hveem K, et al. Development and validation of a protein-based risk score for cardiovascular outcomes among patients with stable coronary heart disease. JAMA. doi: 10.1001/jama.2016.5951 eTable 1. List of 1130 Proteins Measured by Somalogic’s Modified Aptamer-Based Proteomic Assay eTable 2. Coefficients for Weibull Recalibration Model Applied to 9-Protein Model eFigure 1. Median Protein Levels in Derivation and Validation Cohort eTable 3. Coefficients for the Recalibration Model Applied to Refit Framingham eFigure 2. Calibration Plots for the Refit Framingham Model eTable 4. List of 200 Proteins Associated With the Risk of MI, Stroke, Heart Failure, and Death eFigure 3. Hazard Ratios of Lasso Selected Proteins for Primary End Point of MI, Stroke, Heart Failure, and Death eFigure 4. 9-Protein Prognostic Model Hazard Ratios Adjusted for Framingham Variables eFigure 5. 9-Protein Risk Scores by Event Type This supplementary material has been provided by the authors to give readers additional information about their work. Downloaded From: https://jamanetwork.com/ on 10/02/2021 Supplemental Material Table of Contents 1 Study Design and Data Processing ......................................................................................................... 3 2 Table of 1130 Proteins Measured .......................................................................................................... 4 3 Variable Selection and Statistical Modeling ........................................................................................ -

FARE2021WINNERS Sorted by Institute
FARE2021WINNERS Sorted By Institute Swati Shah Postdoctoral Fellow CC Radiology/Imaging/PET and Neuroimaging Characterization of CNS involvement in Ebola-Infected Macaques using Magnetic Resonance Imaging, 18F-FDG PET and Immunohistology The Ebola (EBOV) virus outbreak in Western Africa resulted in residual neurologic abnormalities in survivors. Many case studies detected EBOV in the CSF, suggesting that the neurologic sequelae in survivors is related to viral presence. In the periphery, EBOV infects endothelial cells and triggers a “cytokine stormâ€. However, it is unclear whether a similar process occurs in the brain, with secondary neuroinflammation, neuronal loss and blood-brain barrier (BBB) compromise, eventually leading to lasting neurological damage. We have used in vivo imaging and post-necropsy immunostaining to elucidate the CNS pathophysiology in Rhesus macaques infected with EBOV (Makona). Whole brain MRI with T1 relaxometry (pre- and post-contrast) and FDG-PET were performed to monitor the progression of disease in two cohorts of EBOV infected macaques from baseline to terminal endpoint (day 5-6). Post-necropsy, multiplex fluorescence immunohistochemical (MF-IHC) staining for various cellular markers in the thalamus and brainstem was performed. Serial blood and CSF samples were collected to assess disease progression. The linear mixed effect model was used for statistical analysis. Post-infection, we first detected EBOV in the serum (day 3) and CSF (day 4) with dramatic increases until euthanasia. The standard uptake values of FDG-PET relative to whole brain uptake (SUVr) in the midbrain, pons, and thalamus increased significantly over time (p<0.01) and positively correlated with blood viremia (p≤0.01). -

Assessing the Human Canonical Protein Count[Version 1; Peer Review
F1000Research 2017, 6:448 Last updated: 15 JUL 2020 REVIEW Last rolls of the yoyo: Assessing the human canonical protein count [version 1; peer review: 1 approved, 2 approved with reservations] Christopher Southan IUPHAR/BPS Guide to Pharmacology, Centre for Integrative Physiology, University of Edinburgh, Edinburgh, EH8 9XD, UK First published: 07 Apr 2017, 6:448 Open Peer Review v1 https://doi.org/10.12688/f1000research.11119.1 Latest published: 07 Apr 2017, 6:448 https://doi.org/10.12688/f1000research.11119.1 Reviewer Status Abstract Invited Reviewers In 2004, when the protein estimate from the finished human genome was 1 2 3 only 24,000, the surprise was compounded as reviewed estimates fell to 19,000 by 2014. However, variability in the total canonical protein counts version 1 (i.e. excluding alternative splice forms) of open reading frames (ORFs) in 07 Apr 2017 report report report different annotation portals persists. This work assesses these differences and possible causes. A 16-year analysis of Ensembl and UniProtKB/Swiss-Prot shows convergence to a protein number of ~20,000. The former had shown some yo-yoing, but both have now plateaued. Nine 1 Michael Tress, Spanish National Cancer major annotation portals, reviewed at the beginning of 2017, gave a spread Research Centre (CNIO), Madrid, Spain of counts from 21,819 down to 18,891. The 4-way cross-reference concordance (within UniProt) between Ensembl, Swiss-Prot, Entrez Gene 2 Elspeth A. Bruford , European Molecular and the Human Gene Nomenclature Committee (HGNC) drops to 18,690, Biology Laboratory, Hinxton, UK indicating methodological differences in protein definitions and experimental existence support between sources. -

The Tumor-Associated Antigen PRAME Is Universally Expressed in High-Stage Neuroblastoma and Associated with Poor Outcome
Vol. 10, 4307–4313, July 1, 2004 Clinical Cancer Research 4307 Featured Article The Tumor-Associated Antigen PRAME Is Universally Expressed in High-Stage Neuroblastoma and Associated with Poor Outcome Andre´Oberthuer, Barbara Hero, Ru¨diger Spitz, particularly attractive target for immunotherapeutic strat- Frank Berthold, and Matthias Fischer egies in neuroblastoma. Children’s Hospital, Department of Pediatric Oncology and Hematology, University of Cologne, Cologne, Germany INTRODUCTION Treatment of cancer by inducing a specific immune ABSTRACT response has been the focus of recent studies and might Purpose: The tumor-associated antigen PRAME, a po- provide an efficient therapeutic concept in the near future tential candidate for immunotherapeutic targeting, is fre- (1–3). However, suitable antigens for specific targeting are quently expressed in a variety of cancers. However, no still rare, and their identification remains a major task for information about its presence in neuroblastoma is available promoting further immunotherapeutic strategies. Cancer- to date. We therefore evaluated and quantified PRAME testis antigen(s) (CTA) belong to the group of tumor-associ- expression in a considerable number of neuroblastoma tu- ated antigens and represent possible target proteins for such mors and assessed its impact on the outcome of patients. approaches. Their expression is encountered in a variety of Experimental Design: Qualitative analysis of PRAME malignancies of different histological origin but is negligibly expression was assessed by reverse transcription (RT)-PCR low in healthy tissues, with male germinal cells usually being screening of 94 patients with primary neuroblastoma. The the sole exception (4). Because of this exclusive expression same cohort was used for semiquantitative determination of pattern, CTAs are considered to be promising candidates for transcript levels by Northern blotting, comparing the signal tumor immunotherapy, because they restrict the effects of an intensities of patients with those of testis total RNA. -
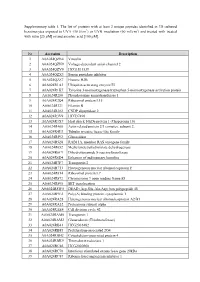
Supplementary Table 1. the List of Proteins with at Least 2 Unique
Supplementary table 1. The list of proteins with at least 2 unique peptides identified in 3D cultured keratinocytes exposed to UVA (30 J/cm2) or UVB irradiation (60 mJ/cm2) and treated with treated with rutin [25 µM] or/and ascorbic acid [100 µM]. Nr Accession Description 1 A0A024QZN4 Vinculin 2 A0A024QZN9 Voltage-dependent anion channel 2 3 A0A024QZV0 HCG1811539 4 A0A024QZX3 Serpin peptidase inhibitor 5 A0A024QZZ7 Histone H2B 6 A0A024R1A3 Ubiquitin-activating enzyme E1 7 A0A024R1K7 Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein 8 A0A024R280 Phosphoserine aminotransferase 1 9 A0A024R2Q4 Ribosomal protein L15 10 A0A024R321 Filamin B 11 A0A024R382 CNDP dipeptidase 2 12 A0A024R3V9 HCG37498 13 A0A024R3X7 Heat shock 10kDa protein 1 (Chaperonin 10) 14 A0A024R408 Actin related protein 2/3 complex, subunit 2, 15 A0A024R4U3 Tubulin tyrosine ligase-like family 16 A0A024R592 Glucosidase 17 A0A024R5Z8 RAB11A, member RAS oncogene family 18 A0A024R652 Methylenetetrahydrofolate dehydrogenase 19 A0A024R6C9 Dihydrolipoamide S-succinyltransferase 20 A0A024R6D4 Enhancer of rudimentary homolog 21 A0A024R7F7 Transportin 2 22 A0A024R7T3 Heterogeneous nuclear ribonucleoprotein F 23 A0A024R814 Ribosomal protein L7 24 A0A024R872 Chromosome 9 open reading frame 88 25 A0A024R895 SET translocation 26 A0A024R8W0 DEAD (Asp-Glu-Ala-Asp) box polypeptide 48 27 A0A024R9E2 Poly(A) binding protein, cytoplasmic 1 28 A0A024RA28 Heterogeneous nuclear ribonucleoprotein A2/B1 29 A0A024RA52 Proteasome subunit alpha 30 A0A024RAE4 Cell division cycle 42 31 -

PRAME Antibody Is a Rabbit Monoclonal Antibody Purified from Ascites
PRAME antibody is a rabbit monoclonal antibody purified from ascites. HRP is ® extracted from horseradish plant. The ihc Enhancer is a signal amplification ihcDirect PRAME Ab-Enh reagent. Novodiax ihc DAB 1:1 Kit or DAB Kit are recommended for use with the Anti-Human PRAME (Clone R1009) PRAME antibody conjugate. Ab: K32042-015 and K32042-030, 150 and 300 tissue stains* PRAME Ab-Enh Components (K47042-###): Ab K32042-005 and K32042-010 50 and 100 tissue stains* Ab-Enh: K47042-015 and K47042-030, 150 and 300 tissue stains* Reagent Description Component Part Numbers Sizes (mL) Ab-Enh: K47042-005 and K47042-010 50 and 100 tissue stains* PRAME pHRP H32042-R### (005, 015) 5, 15 Intended Use: For In Vitro Diagnostic Use ihc Enhancer D28020-R### (005, 015) 5, 15 Polymerized horseradish peroxidase (polyHRP)-labeled anti-PRAME (R1009) rabbit monoclonal antibody is intended for laboratory use to qualitatively identify PRAME Ab Components (K32042-###): by light microscopy the presence of PRAME protein in sections of formalin-fixed, paraffin-embedded (FFPE) and/or fresh frozen tissues using Reagent Description Component Part Numbers Sizes (mL) immunohistochemistry (IHC) test methods. The clinical interpretation of any PRAME pHRP H32042-R### (005, 015) 5, 15 staining or its absence should be complemented by morphological studies using proper controls and should be evaluated within the context of the patient’s clinical Ancillary Reagents for Use with PRAME pHRP Antibody: history and other diagnostic tests and proper controls interpreted by a qualified pathologist and/or physician. This conjugated antibody has been pre-diluted and Reagent Description Part Numbers Sizes (mL) optimized for IHC use without further dilution.