PI 3-Kinase Isoforms at the Crossroads of Signalling and Vesicular Traffic
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Symbol Gene Description ACVR1B Activin a Receptor, Type IB
Table S1. Kinase clones included in human kinase cDNA library for yeast two-hybrid screening Gene Symbol Gene Description ACVR1B activin A receptor, type IB ADCK2 aarF domain containing kinase 2 ADCK4 aarF domain containing kinase 4 AGK multiple substrate lipid kinase;MULK AK1 adenylate kinase 1 AK3 adenylate kinase 3 like 1 AK3L1 adenylate kinase 3 ALDH18A1 aldehyde dehydrogenase 18 family, member A1;ALDH18A1 ALK anaplastic lymphoma kinase (Ki-1) ALPK1 alpha-kinase 1 ALPK2 alpha-kinase 2 AMHR2 anti-Mullerian hormone receptor, type II ARAF v-raf murine sarcoma 3611 viral oncogene homolog 1 ARSG arylsulfatase G;ARSG AURKB aurora kinase B AURKC aurora kinase C BCKDK branched chain alpha-ketoacid dehydrogenase kinase BMPR1A bone morphogenetic protein receptor, type IA BMPR2 bone morphogenetic protein receptor, type II (serine/threonine kinase) BRAF v-raf murine sarcoma viral oncogene homolog B1 BRD3 bromodomain containing 3 BRD4 bromodomain containing 4 BTK Bruton agammaglobulinemia tyrosine kinase BUB1 BUB1 budding uninhibited by benzimidazoles 1 homolog (yeast) BUB1B BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) C9orf98 chromosome 9 open reading frame 98;C9orf98 CABC1 chaperone, ABC1 activity of bc1 complex like (S. pombe) CALM1 calmodulin 1 (phosphorylase kinase, delta) CALM2 calmodulin 2 (phosphorylase kinase, delta) CALM3 calmodulin 3 (phosphorylase kinase, delta) CAMK1 calcium/calmodulin-dependent protein kinase I CAMK2A calcium/calmodulin-dependent protein kinase (CaM kinase) II alpha CAMK2B calcium/calmodulin-dependent -

AN INVESTIGATION of the ROLE of PAK6 in TUMORIGENESIS By
AN INVESTIGATION OF THE ROLE OF PAK6 IN TUMORIGENESIS by JoAnn Roberts A Thesis Submitted to the Faculty of The Charles E. Schmidt College of Medicine In Partial Fulfillment of the Requirements for the Degree of Master of Science Florida Atlantic University Boca Raton, Florida August 2012 ACKNOWLEDGMENTS This material is based upon work supported by the National Science Foundation under Grant No. DGE: 0638662. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation. I would like to thank and acknowledge my thesis advisor, Dr. Michael Lu, for his support and guidance throughout the writing of this thesis and design of experiments in this manuscript. I would also like to thank my colleagues for assistance in various trouble-shooting circumstances. Last, but certainly not least, I would like to thank my family and friends for their support in the pursuit of my graduate studies. iii ABSTRACT Author: JoAnn Roberts Title: An Investigation of the Role of PAK6 in Tumorigenesis Institution: Florida Atlantic University Thesis Advisor: Dr. Michael Lu Degree: Master of Science Year: 2012 The function and role of PAK6, a serine/threonine kinase, in cancer progression has not yet been clearly identified. Several studies reveal that PAK6 may participate in key changes contributing to cancer progression such as cell survival, cell motility, and invasiveness. Based on the membrane localization of PAK6 in prostate and breast cancer cells, we speculated that PAK6 plays a role in cancer progression cells by localizing on the membrane and modifying proteins linked to motility and proliferation. -

New Somatic Mutations and WNK1-B4GALNT3 Gene Fusion in Papillary Thyroid Carcinoma
www.impactjournals.com/oncotarget/ Oncotarget, Vol. 6, No. 13 New somatic mutations and WNK1-B4GALNT3 gene fusion in papillary thyroid carcinoma Valerio Costa1,*, Roberta Esposito1,*, Carmela Ziviello1, Romina Sepe2, Larissa Valdemarin Bim2, Nunzio Antonio Cacciola2, Myriam Decaussin-Petrucci3, Pierlorenzo Pallante2, Alfredo Fusco2,4 and Alfredo Ciccodicola1,5 1 Institute of Genetics and Biophysics “Adriano Buzzati-Traverso”, CNR, Naples, Italy 2 Istituto per l’Endocrinologia e l’Oncologia Sperimentale (IEOS), Consiglio Nazionale delle Ricerche (CNR), c/o Dipartimento di Medicina Molecolare e Biotecnologie Mediche (DMMBM), Università degli Studi di Napoli “Federico II”, Naples, Italy 3 Department of Pathology, Lyon Sud Hospital Center, Hospices Civils de Lyon, Pierre-Bénite, Lyon, France 4 Instituto Nacional de Câncer - INCA, Praça da Cruz Vermelha, Rio de Janeiro, RJ, Brazil 5 Department of Science and Technology, University “Parthenope” of Naples, Italy * These authors contributed equally to this article Correspondence to: Alfredo Fusco, email: [email protected] Correspondence to: Alfredo Ciccodicola, email: [email protected] Keywords: thyroid, papillary carcinomas, RNA-Sequencing, gene fusions, mutations Received: February 24, 2015 Accepted: February 25, 2015 Published: March 14, 2015 This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT Papillary thyroid carcinoma (PTC) is the most frequent thyroid malignant neoplasia. Oncogene activation occurs in more than 70% of the cases. Indeed, about 40% of PTCs harbor mutations in BRAF gene, whereas RET rearrangements (RET/PTC oncogenes) are present in about 20% of cases. -

PIK3C2G (NM 004570) Human Mutant ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC402758 PIK3C2G (NM_004570) Human Mutant ORF Clone Product data: Product Type: Mutant ORF Clones Product Name: PIK3C2G (NM_004570) Human Mutant ORF Clone Mutation Description: P146L Affected Codon#: 146 Affected NT#: 437 Nucleotide Mutation: PIK3C2G Mutant (P146L), Myc-DDK-tagged ORF clone of Homo sapiens phosphoinositide-3- kinase, class 2, gamma polypeptide (PIK3C2G) as transfection-ready DNA Effect: Dibees, ype 2, ssoiion wih Symbol: PIK3C2G Synonyms: PI3K-C2-gamma; PI3K-C2GAMMA Vector: pCMV6-Entry (PS100001) Tag: Myc-DDK ACCN: NM_004570 ORF Size: 4335 bp ORF Nucleotide >RC402758 representing NM_004570 Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGGCATATTCTTGGCAAACGGATCCAAATCCTAATGAATCACACGAAAAGCAGTATGAACACCAAGAAT TTCTCTTTGTAAATCAACCCCATTCTTCTAGCCAAGTCAGTCTGGGTTTTGATCAGATAGTAGATGAGAT CAGTGGCAAAATTCCACACTACGAGAGTGAAATTGATGAAAACACCTTTTTTGTGCCCACTGCACCAAAA TGGGACTCAACAGGGCATTCATTAAATGAAGCACACCAAATATCCTTGAATGAATTCACTTCTAAAAGCC GTGAACTCTCCTGGCATCAAGTTAGCAAAGCACCAGCAATTGGTTTTAGTCCTTCTGTGTTACCAAAACC TCAAAATACGAATAAAGAATGCTCCTGGGGAAGCCCCATAGGAAAACATCATGGTGCTGATGATTCCAGA TTCAGTATTTTAGCTCTATCATTCACAAGTTTGGATAAAATTAATCTAGAGAAAGAATTAGAAAATGAAA ATCATAACTACCATATAGGATTTGAAAGTAGCATTCCTCCAACAAATTCATCCTTCTCAAGTGACTTCAT GCCGAAAGAAGAGAATAAAAGGAGTGGACATGTGAACATTGTGGAACCATCTTTGATGCTTTTGAAAGGC -

Phosphoinositide 3-Kinase-C2α Regulates Polycystin-2 Ciliary Entry
BASIC RESEARCH www.jasn.org Phosphoinositide 3-Kinase-C2a Regulates Polycystin-2 Ciliary Entry and Protects against Kidney Cyst Formation † Irene Franco,* Jean Piero Margaria,* Maria Chiara De Santis,* Andrea Ranghino, ‡ Daniel Monteyne, Marco Chiaravalli,§ Monika Pema,§ Carlo Cosimo Campa,* ‡| Edoardo Ratto,* Federico Gulluni,* David Perez-Morga, Stefan Somlo,¶ Giorgio R. Merlo,* Alessandra Boletta,§ and Emilio Hirsch* *Molecular Biotechnology Center, Department of Molecular Biotechnology and Health Sciences, University of Torino, Turin, Italy; †Renal Transplantation Center “A. Vercellone”, Division of Nephrology, Dialysis and Transplantation, Department of Medical Sciences, Città della Salute e della Scienza, Hospital and Research Center for Experimental Medicine (CeRMS) and Center for Molecular Biotechnology, University of Torino, Turin, Italy; ‡Laboratoire de Parasitologie Moléculaire, Institut de Biologie et de Médecine Moléculaires (IBMM), Université Libre de Bruxelles, Gosselies, Charleroi, Belgium; §Division of Genetics and Cell Biology, Dibit San Raffaele Scientific Institute, Milan, Italy; |Center for Microscopy and Molecular Imaging (CMMI), Université Libre de Bruxelles, Gosselies, Belgium; and ¶Section of Nephrology, Yale University School of Medicine, New Haven, Connecticut. ABSTRACT Signaling from the primary cilium regulates kidney tubule development and cyst formation. However, the mechanism controlling targeting of ciliary components necessary for cilium morphogenesis and signaling is largely unknown. Here, we studied the function of class II phosphoinositide 3-kinase-C2a (PI3K-C2a)inrenal tubule-derived inner medullary collecting duct 3 cells and show that PI3K-C2a resides at the recycling endo- some compartment in proximity to the primary cilium base. In this subcellular location, PI3K-C2a controlled the activation of Rab8, a key mediator of cargo protein targeting to the primary cilium. -

The DNA Methylation Landscape of Glioblastoma Disease Progression Shows Extensive Heterogeneity in Time and Space
bioRxiv preprint doi: https://doi.org/10.1101/173864; this version posted August 9, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. The DNA methylation landscape of glioblastoma disease progression shows extensive heterogeneity in time and space Johanna Klughammer1*, Barbara Kiesel2,3*, Thomas Roetzer3,4, Nikolaus Fortelny1, Amelie Kuchler1, Nathan C. Sheffield5, Paul Datlinger1, Nadine Peter3,4, Karl-Heinz Nenning6, Julia Furtner3,7, Martha Nowosielski8,9, Marco Augustin10, Mario Mischkulnig2,3, Thomas Ströbel3,4, Patrizia Moser11, Christian F. Freyschlag12, Jo- hannes Kerschbaumer12, Claudius Thomé12, Astrid E. Grams13, Günther Stockhammer8, Melitta Kitzwoegerer14, Stefan Oberndorfer15, Franz Marhold16, Serge Weis17, Johannes Trenkler18, Johanna Buchroithner19, Josef Pichler20, Johannes Haybaeck21,22, Stefanie Krassnig21, Kariem Madhy Ali23, Gord von Campe23, Franz Payer24, Camillo Sherif25, Julius Preiser26, Thomas Hauser27, Peter A. Winkler27, Waltraud Kleindienst28, Franz Würtz29, Tanisa Brandner-Kokalj29, Martin Stultschnig30, Stefan Schweiger31, Karin Dieckmann3,32, Matthias Preusser3,33, Georg Langs6, Bernhard Baumann10, Engelbert Knosp2,3, Georg Widhalm2,3, Christine Marosi3,33, Johannes A. Hainfellner3,4, Adelheid Woehrer3,4#§, Christoph Bock1,34,35# 1 CeMM Research Center for Molecular Medicine of the Austrian Academy of Sciences, Vienna, Austria. 2 Department of Neurosurgery, Medical University of Vienna, Vienna, Austria. 3 Comprehensive Cancer Center, Central Nervous System Tumor Unit, Medical University of Vienna, Austria. 4 Institute of Neurology, Medical University of Vienna, Vienna, Austria. 5 Center for Public Health Genomics, University of Virginia, Charlottesville VA, USA. 6 Department of Biomedical Imaging and Image-guided Therapy, Computational Imaging Research Lab, Medical University of Vi- enna, Vienna, Austria. -

Mutations in PIK3C2A Cause Syndromic Short Stature
University of Groningen Mutations in PIK3C2A cause syndromic short stature, skeletal abnormalities, and cataracts associated with ciliary dysfunction Tiosano, Dov; Baris, Hagit N; Chen, Anlu; Hitzert, Marrit M; Schueler, Markus; Gulluni, Federico; Wiesener, Antje; Bergua, Antonio; Mory, Adi; Copeland, Brett Published in: PLoS genetics DOI: 10.1371/journal.pgen.1008088 IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it. Please check the document version below. Document Version Publisher's PDF, also known as Version of record Publication date: 2019 Link to publication in University of Groningen/UMCG research database Citation for published version (APA): Tiosano, D., Baris, H. N., Chen, A., Hitzert, M. M., Schueler, M., Gulluni, F., ... Buchner, D. A. (2019). Mutations in PIK3C2A cause syndromic short stature, skeletal abnormalities, and cataracts associated with ciliary dysfunction. PLoS genetics, 15(4), [e1008088]. https://doi.org/10.1371/journal.pgen.1008088 Copyright Other than for strictly personal use, it is not permitted to download or to forward/distribute the text or part of it without the consent of the author(s) and/or copyright holder(s), unless the work is under an open content license (like Creative Commons). Take-down policy If you believe that this document breaches copyright please contact us providing details, and we will remove access to the work immediately and investigate your claim. Downloaded from the University of Groningen/UMCG research database (Pure): http://www.rug.nl/research/portal. For technical reasons the number of authors shown on this cover page is limited to 10 maximum. -
HCC and Cancer Mutated Genes Summarized in the Literature Gene Symbol Gene Name References*
HCC and cancer mutated genes summarized in the literature Gene symbol Gene name References* A2M Alpha-2-macroglobulin (4) ABL1 c-abl oncogene 1, receptor tyrosine kinase (4,5,22) ACBD7 Acyl-Coenzyme A binding domain containing 7 (23) ACTL6A Actin-like 6A (4,5) ACTL6B Actin-like 6B (4) ACVR1B Activin A receptor, type IB (21,22) ACVR2A Activin A receptor, type IIA (4,21) ADAM10 ADAM metallopeptidase domain 10 (5) ADAMTS9 ADAM metallopeptidase with thrombospondin type 1 motif, 9 (4) ADCY2 Adenylate cyclase 2 (brain) (26) AJUBA Ajuba LIM protein (21) AKAP9 A kinase (PRKA) anchor protein (yotiao) 9 (4) Akt AKT serine/threonine kinase (28) AKT1 v-akt murine thymoma viral oncogene homolog 1 (5,21,22) AKT2 v-akt murine thymoma viral oncogene homolog 2 (4) ALB Albumin (4) ALK Anaplastic lymphoma receptor tyrosine kinase (22) AMPH Amphiphysin (24) ANK3 Ankyrin 3, node of Ranvier (ankyrin G) (4) ANKRD12 Ankyrin repeat domain 12 (4) ANO1 Anoctamin 1, calcium activated chloride channel (4) APC Adenomatous polyposis coli (4,5,21,22,25,28) APOB Apolipoprotein B [including Ag(x) antigen] (4) AR Androgen receptor (5,21-23) ARAP1 ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 1 (4) ARHGAP35 Rho GTPase activating protein 35 (21) ARID1A AT rich interactive domain 1A (SWI-like) (4,5,21,22,24,25,27,28) ARID1B AT rich interactive domain 1B (SWI1-like) (4,5,22) ARID2 AT rich interactive domain 2 (ARID, RFX-like) (4,5,22,24,25,27,28) ARID4A AT rich interactive domain 4A (RBP1-like) (28) ARID5B AT rich interactive domain 5B (MRF1-like) (21) ASPM Asp (abnormal -

UC San Diego Electronic Theses and Dissertations
UC San Diego UC San Diego Electronic Theses and Dissertations Title Isolation and characterization of neuronal substrates of the ubiquitin proteasome system Permalink https://escholarship.org/uc/item/7jg5g3qg Author Keil, Jeffrey McCartney Publication Date 2011 Peer reviewed|Thesis/dissertation eScholarship.org Powered by the California Digital Library University of California UNIVERSITY OF CALIFORNIA, SAN DIEGO Isolation and Characterization of Neuronal Substrates of the Ubiquitin Proteasome System A dissertation submitted in partial satisfaction of the requirements for the degree Doctor of Philosophy in Biology by Jeffrey McCartney Keil Committee in charge: Professor Gentry Patrick, Chair Professor Michael Burkart Professor Randolph Hampton Professor Terunaga Nakagawa Professor Yimin Zou 2011 Copyright Jeffrey McCartney Keil, 2011 All rights reserved The dissertation of Jeffrey McCartney Keil is approved, and it is acceptable in quality and form for publication on microfilm and electronically: ________________________________________________________________________ ________________________________________________________________________ ________________________________________________________________________ ________________________________________________________________________ ________________________________________________________________________ Chair University of California, San Diego 2011 iii DEDICATION To my loving parents, Richard and Mary, who made this all possible. iv EPIGRAPH You could tell by the way he talked, though, -

Analysis of Inherited and Somatic Variants to Decipher Canine Complex Traits
Digital Comprehensive Summaries of Uppsala Dissertations from the Faculty of Medicine 1454 Analysis of inherited and somatic variants to decipher canine complex traits KATE MEGQUIER ACTA UNIVERSITATIS UPSALIENSIS ISSN 1651-6206 ISBN 978-91-513-0310-9 UPPSALA urn:nbn:se:uu:diva-347165 2018 Dissertation presented at Uppsala University to be publicly examined in B:22, BMC, Husargatan 3, Uppsala, Monday, 21 May 2018 at 13:15 for the degree of Doctor of Philosophy (Faculty of Medicine). The examination will be conducted in English. Faculty examiner: David Sargan (University of Cambridge). Abstract Megquier, K. 2018. Analysis of inherited and somatic variants to decipher canine complex traits. Digital Comprehensive Summaries of Uppsala Dissertations from the Faculty of Medicine 1454. 67 pp. Uppsala: Acta Universitatis Upsaliensis. ISBN 978-91-513-0310-9. This thesis presents several investigations of the dog as a model for complex diseases, focusing on cancers and the effect of genetic risk factors on clinical presentation. In Papers I and II, we performed genome-wide association studies (GWAS) to identify germline risk factors predisposing US golden retrievers to hemangiosarcoma (HSA) and B- cell lymphoma (BLSA). Paper I identified two loci predisposing to both HSA and BLSA, approximately 4 megabases (Mb) apart on chromosome 5. Carrying the risk haplotype at these loci was associated with separate changes in gene expression, both relating to T-cell activation and proliferation. Paper II followed up on the HSA GWAS by performing a meta-analysis with additional cases and controls. This confirmed three previously reported GWAS loci for HSA and revealed three new loci, the most significant on chromosome 18. -
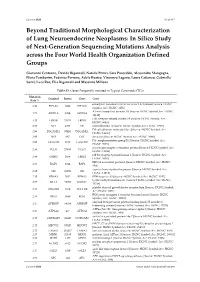
Beyond Traditional Morphological Characterization of Lung
Cancers 2020 S1 of S15 Beyond Traditional Morphological Characterization of Lung Neuroendocrine Neoplasms: In Silico Study of Next-Generation Sequencing Mutations Analysis across the Four World Health Organization Defined Groups Giovanni Centonze, Davide Biganzoli, Natalie Prinzi, Sara Pusceddu, Alessandro Mangogna, Elena Tamborini, Federica Perrone, Adele Busico, Vincenzo Lagano, Laura Cattaneo, Gabriella Sozzi, Luca Roz, Elia Biganzoli and Massimo Milione Table S1. Genes Frequently mutated in Typical Carcinoids (TCs). Mutation Original Entrez Gene Gene Rate % eukaryotic translation initiation factor 1A X-linked [Source: HGNC 4.84 EIF1AX 1964 EIF1AX Symbol; Acc: HGNC: 3250] AT-rich interaction domain 1A [Source: HGNC Symbol;Acc: HGNC: 4.71 ARID1A 8289 ARID1A 11110] LDL receptor related protein 1B [Source: HGNC Symbol; Acc: 4.35 LRP1B 53353 LRP1B HGNC: 6693] 3.53 NF1 4763 NF1 neurofibromin 1 [Source: HGNC Symbol;Acc: HGNC: 7765] DS cell adhesion molecule like 1 [Source: HGNC Symbol; Acc: 2.90 DSCAML1 57453 DSCAML1 HGNC: 14656] 2.90 DST 667 DST dystonin [Source: HGNC Symbol;Acc: HGNC: 1090] FA complementation group D2 [Source: HGNC Symbol; Acc: 2.90 FANCD2 2177 FANCD2 HGNC: 3585] piccolo presynaptic cytomatrix protein [Source: HGNC Symbol; Acc: 2.90 PCLO 27445 PCLO HGNC: 13406] erb-b2 receptor tyrosine kinase 2 [Source: HGNC Symbol; Acc: 2.44 ERBB2 2064 ERBB2 HGNC: 3430] BRCA1 associated protein 1 [Source: HGNC Symbol; Acc: HGNC: 2.35 BAP1 8314 BAP1 950] capicua transcriptional repressor [Source: HGNC Symbol; Acc: 2.35 CIC 23152 CIC HGNC: -

Duke University Dissertation Template
Gene-Environment Interactions in Cardiovascular Disease by Cavin Keith Ward-Caviness Graduate Program in Computational Biology and Bioinformatics Duke University Date:_______________________ Approved: ___________________________ Elizabeth R. Hauser, Supervisor ___________________________ William E. Kraus ___________________________ Sayan Mukherjee ___________________________ H. Frederik Nijhout Dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Graduate Program in Computational Biology and Bioinformatics in the Graduate School of Duke University 2014 i v ABSTRACT Gene-Environment Interactions in Cardiovascular Disease by Cavin Keith Ward-Caviness Graduate Program in Computational Biology and Bioinformatics Duke University Date:_______________________ Approved: ___________________________ Elizabeth R. Hauser, Supervisor ___________________________ William E. Kraus ___________________________ Sayan Mukherjee ___________________________ H. Frederik Nijhout An abstract of a dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Graduate Program in Computational Biology and Bioinformatics in the Graduate School of Duke University 2014 Copyright by Cavin Keith Ward-Caviness 2014 Abstract In this manuscript I seek to demonstrate the importance of gene-environment interactions in cardiovascular disease. This manuscript contains five studies each of which contributes to our understanding of the joint impact of genetic variation