(12) Patent Application Publication (10) Pub. No.: US 2007/0105105 A1 Clelland Et Al
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Mass Spectrometry-Based Proteomics Techniques and Their Application in Ovarian Cancer Research Agata Swiatly, Szymon Plewa, Jan Matysiak and Zenon J
Swiatly et al. Journal of Ovarian Research (2018) 11:88 https://doi.org/10.1186/s13048-018-0460-6 REVIEW Open Access Mass spectrometry-based proteomics techniques and their application in ovarian cancer research Agata Swiatly, Szymon Plewa, Jan Matysiak and Zenon J. Kokot* Abstract Ovarian cancer has emerged as one of the leading cause of gynecological malignancies. So far, the measurement of CA125 and HE4 concentrations in blood and transvaginal ultrasound examination are essential ovarian cancer diagnostic methods. However, their sensitivity and specificity are still not sufficient to detect disease at the early stage. Moreover, applied treatment may appear to be ineffective due to drug-resistance. Because of a high mortality rate of ovarian cancer, there is a pressing need to develop innovative strategies leading to a full understanding of complicated molecular pathways related to cancerogenesis. Recent studies have shown the great potential of clinical proteomics in the characterization of many diseases, including ovarian cancer. Therefore, in this review, we summarized achievements of proteomics in ovarian cancer management. Since the development of mass spectrometry has caused a breakthrough in systems biology, we decided to focus on studies based on this technique. According to PubMed engine, in the years 2008–2010 the number of studies concerning OC proteomics was increasing, and since 2010 it has reached a plateau. Proteomics as a rapidly evolving branch of science may be essential in novel biomarkers discovery, therapy decisions, progression predication, monitoring of drug response or resistance. Despite the fact that proteomics has many to offer, we also discussed some limitations occur in ovarian cancer studies. -

Original Article Dynamics of Lamins B and A/C and Nucleoporin Nup160 During Meiotic Maturation in Mouse Oocytes
Original Article Dynamics of Lamins B and A/C and Nucleoporin Nup160 during Meiotic Maturation in Mouse Oocytes (oocytes / meiosis / meiotic spindle / nuclear lamina / Nup107-160 / nuclear pore complex) V. NIKOLOVA, S. DELIMITREVA, I. CHAKAROVA, R. ZHIVKOVA, V. HADZHINESHEVA, M. MARKOVA Department of Biology, Medical Faculty, Medical University of Sofia, Bulgaria Abstract. This study was aimed at elucidating the plex reorganization of the cytoskeleton and nuclear en- fate of three important nuclear envelope components velope (Delimitreva et al., 2012). Although the early – lamins B and A/C and nucleoporin Nup160, during meiotic stages have been relatively well studied, the meiotic maturation of mouse oocytes. These proteins events of final steps of oocyte meiosis (from meiotic re- were localized by epifluorescence and confocal mi- sumption in late prophase I until metaphase II) are still croscopy using specific antibodies in oocytes at dif- poorly understood. The oocyte nucleus in late prophase ferent stages from prophase I (germinal vesicle) to I, traditionally called GV (germinal vesicle), becomes metaphase II. In immature germinal vesicle oocytes, competent to resume meiosis upon accumulation of all three proteins were detected at the nuclear pe- pericentriolar heterochromatin called karyosphere, sur- riphery. In metaphase I and metaphase II, lamin B rounded nucleolus or rimmed nucleolus (Can et al., co-localized with the meiotic spindle, lamin A/C was 2003; De la Fuente et al., 2004; Tan et al., 2009). Then, found in a diffuse halo surrounding the spindle and the nucleus disaggregates in the so-called germinal ve- to a lesser degree throughout the cytoplasm, and sicle breakdown (GVBD) stage. -
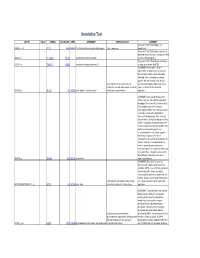
Supplementary Table 5. Functional Annotation of the Largest Gene Cluster(221 Element)
Annotation Tool AFFYID VALUE SYMBOL LOCUSLINK OMIM GENENAME GENEONTOLOGY SUMMARY [Proteome FUNCTION:] Expressed 203054_s_at TCTA 6988 600690 T-cell leukemia translocation altered gene tumor suppressor ubiquitously [Proteome FUNCTION:] May be involved in protein-protein interactions; contains five WD 44563_at FLJ10385 55135 hypothetical protein FLJ10385 domains (WD-40 repeats) [Proteome FUNCTION:] Weakly similarity to 212261_at TNRC15 26058 trinucleotide repeat containing 15 a region of rat nestin (Rn.9701) [SUMMARY:] Actin alpha 1 which is expressed in skeletal muscle is one of six different actin isoforms which have been identified. Actins are highly conserved proteins that are involved in cell motility, actin filament; motor activity; muscle structure and integrity. Alpha actins are a contraction; muscle development; structural major constituent of the contractile 203872_at ACTA1 58 102610 actin, alpha 1, skeletal muscle constituent of cytoskeleton apparatus. [SUMMARY:] Annexin VIII belong to the family of Ca (2+) dependent phospholipid binding proteins (annexins), and has a high 56% identity to annexin V (vascular anticoagulant-alpha). It was initially isolated as 2.2 kb vascular anticoagulant-beta transcript from human placenta, a Ca (2+) dependent phospholipid binding protein that inhibits coagulation and phospholipase A2 activity. However, the fact that annexin VIII is neither an extracellular protein nor associated with the cell surface suggests that it may not play a role in blood coagulation in vivo and its physiological role remains unknown. It is expressed at low levels in human placenta and shows restricted expression in lung endothelia, skin, liver, and kidney. The gene is also found to be selectively overexpressed in acute 203074_at ANXA8 244 602396 annexin A8 myelocytic leukemia. -

Antigen-Specific Memory CD4 T Cells Coordinated Changes in DNA
Downloaded from http://www.jimmunol.org/ by guest on September 24, 2021 is online at: average * The Journal of Immunology The Journal of Immunology published online 18 March 2013 from submission to initial decision 4 weeks from acceptance to publication http://www.jimmunol.org/content/early/2013/03/17/jimmun ol.1202267 Coordinated Changes in DNA Methylation in Antigen-Specific Memory CD4 T Cells Shin-ichi Hashimoto, Katsumi Ogoshi, Atsushi Sasaki, Jun Abe, Wei Qu, Yoichiro Nakatani, Budrul Ahsan, Kenshiro Oshima, Francis H. W. Shand, Akio Ametani, Yutaka Suzuki, Shuichi Kaneko, Takashi Wada, Masahira Hattori, Sumio Sugano, Shinichi Morishita and Kouji Matsushima J Immunol Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Author Choice option Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Freely available online through http://www.jimmunol.org/content/suppl/2013/03/18/jimmunol.120226 7.DC1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material Permissions Email Alerts Subscription Author Choice Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 24, 2021. Published March 18, 2013, doi:10.4049/jimmunol.1202267 The Journal of Immunology Coordinated Changes in DNA Methylation in Antigen-Specific Memory CD4 T Cells Shin-ichi Hashimoto,*,†,‡ Katsumi Ogoshi,* Atsushi Sasaki,† Jun Abe,* Wei Qu,† Yoichiro Nakatani,† Budrul Ahsan,x Kenshiro Oshima,† Francis H. -

Clinical Chemistry
June 1 2007, Volume 53, Issue 6,pp.999- 1180 Editorials Andrew McCaddon and Peter R. Hudson Methylation and Phosphorylation: A Tangled Relationship? Clin Chem 2007 53: 999-1000. Bob Palais Quantitative Heteroduplex Analysis Clin Chem 2007 53: 1001-1003. Eleftherios P. Diamandis Oncopeptidomics: A Useful Approach for Cancer Diagnosis? Clin Chem 2007 53: 1004-1006. Roger D. Klein The Pain Protective Haplotype: Introducing the Modern Genetic Test Clin Chem 2007 53: 1007-1009. Molecular Diagnostics and Genetics Jörn Lötsch, Inna Belfer, Anja Kirchhof, Bikash K. Mishra, Mitchell B. Max, Alexandra Doehring, Michael Costigan, Clifford J. Woolf, Gerd Geisslinger, and Irmgard Tegeder Reliable Screening for a Pain-Protective Haplotype in the GTP Cyclohydrolase 1 Gene (GCH1) Through the Use of 3 or Fewer Single Nucleotide Polymorphisms Clin Chem 2007 53: 1010-1015. Published online March 15, 2007; 10.1373/clinchem.2006.082883 Kerstin L. Edlefsen, Jonathan F. Tait, Mark H. Wener, and Michael Astion Utilization and Diagnostic Yield of Neurogenetic Testing at a Tertiary Care Facility Clin Chem 2007 53: 1016-1022. Published online April 19, 2007; 10.1373/clinchem.2006.083360 Sara Bremer, Helge Rootwelt, and Stein Bergan Real-Time PCR Determination of IMPDH1 and IMPDH2 Expression in Blood Cells Clin Chem 2007 53: 1023-1029. Published online April 26, 2007; 10.1373/clinchem.2006.081968 Elizabeth Herness Peters, Sandra Rojas-Caro, Mitchell G. Brigell, Robert J. Zahorchak, Shelley Ann des Etages, Patricia L. Ruppel, Charles R. Knight, Bradley Austermiller, Myrna C. Graham, Steve Wowk, Sean Banks, Lakshmi V. Madabusi, Patrick Turk, Donna Wilder, Carole Kempfer, Terry W. Osborn, and James C. -

Lineage-Specific Evolution of the Complex Nup160 Hybrid
GENETICS | INVESTIGATION Lineage-Specific Evolution of the Complex Nup160 Hybrid Incompatibility Between Drosophila melanogaster and Its Sister Species Shanwu Tang and Daven C. Presgraves1 Department of Biology, University of Rochester, New York 14627 ABSTRACT Two genes encoding protein components of the nuclear pore complex Nup160 and Nup96 cause lethality in F2-like hybrid genotypes between Drosophila simulans and Drosophila melanogaster. In particular, D. simulans Nup160 and Nup96 each cause inviability when hemizygous or homozygous in species hybrids that are also hemizygous (or homozygous) for the D. melanogaster X chromosome. The hybrid lethality of Nup160, however, is genetically complex, depending on one or more unknown additional factors in the autosomal background. Here we study the genetics and evolution of Nup160-mediated hybrid lethality in three ways. First, we test for variability in Nup160-mediated hybrid lethality within and among the three species of the D. simulans clade— D. simulans, D. sechellia,andD. mauritiana. We show that the hybrid lethality of Nup160 is fixed in D. simulans and D. sechellia but absent in D. mauritiana. Second, we explore how the hybrid lethality of Nup160 depends on other loci in the autosomal background. We find that D. simulans Nup160-mediated hybrid lethality does not depend on the presence of D. melanogaster Nup96,andwefind that D. simulans and D. mauritiana are functionally differentiated at Nup160 as well as at other autosomal factor(s). Finally, we use population genetics data to show that Nup160 has experienced histories of recurrent positive selection both before and after the split of the three D. simulans clade species 240,000 years ago. -

1 Supplementary Table S1. Histological And
Supplementary Table S1. Histological and immunocytochemestry of the bcMCF clones. Clones Tumor type AE1 CAM5.2 EMA Vimentin bcMCF-1 Invasive poorly - - - +++ bcMCF-4 differentiated spindle cell type bcMCF-2 Invasive poorly +/- +/- +/- ++ bcMCF-6 differentiated bcMCF-7 epithelial cell type bcMCF-3 Invasive poorly - +/- +/- ++ bcMCF-5 differentiated with mix features of spindle and epithelial type The mouse monoclonal antibodies anti- human cytokeratin of low molecular weight (AE1, Biogenex, San Ramon, CA), cytokeratin peptides 7 and 8 (CAM5.2, Ventana, Tucson, AZ), epithelial membrane antigen (EMA) clone E29 and vimentin, clone V9, both from DakoCytomation Inc. (Fort Collins, CO) were used. Negative (-), weak (+/-), moderate (++) or strong (+++). 1 Supplementary Table S2. Differentially expressed apoptosis genes (GO:00009165) in tumorigenic bcMCF cells. Fold change Symbol Gene name trMCF bcMC caMCF AHR aryl hydrocarbon receptor Ns* -2.1 Ns APP amyloid beta (A4) precursor protein Ns -1.7 -1.7 BAG1 BCL2-associated athanogene Ns -7.6 -9.2 BAG5 BCL2-associated athanogene 5 Ns -1.8 Ns BIRC4 baculoviral IAP repeat-containing 4 Ns -2.0 -2.7 BNIP3L BCL2/adenovirus E1B interacting protein 3-like Ns -2 -1.9 CASP14 caspase 14, apoptosis-related cysteine peptidase Ns -12 -12.3 CASP3 caspase 3, apoptosis-related cysteine peptidase Ns -2.4 Ns CASP6 caspase 6, apoptosis-related cysteine peptidase Ns -4.1 -4.1 CD14 CD14 antigen Ns -4.7 Ns DAPK1 death-associated protein kinase 1 Ns -4.1 -13.4 ELMO3 engulfment and cell motility 3 Ns -9.9 -10.8 ELMOD2 ELMO -

A Test of Double Interspecific Introgression of Nucleoporin Genes
INVESTIGATION A Test of Double Interspecific Introgression of Nucleoporin Genes in Drosophila Kyoichi Sawamura,*,1 Kazunori Maehara,† Yoko Keira,‡ Hiroyuki O. Ishikawa,‡ Takeshi Sasamura,§ Tomoko Yamakawa,§ and Kenji Matsuno§ *Faculty of Life and Environmental Sciences, and †Graduate School of Life and Environmental Sciences, University of Tsukuba, Tsukuba, Ibaraki 305-8572, ‡Department of Biology, Chiba University, Chiba, Chiba 263-8522, and § Department of Biological Sciences, Osaka University, Toyonaka, Osaka, Japan 560-0043 ABSTRACT In interspecific hybrids between Drosophila melanogaster and Drosophila simulans, the D. KEYWORDS simulans nucleoporin-encoding Nup96sim and Nup160sim can cause recessive lethality if the hybrid does Drosophila not also inherit the D. simulans X chromosome. In addition, Nup160sim leads to recessive female sterility in hybrid inviability the D. melanogaster genetic background. Here, we conducted carefully controlled crosses to better hybrid sterility understand the relationship between Nup96sim and Nup160sim. Nup96sim did not lead to female sterility nucleoporin in the D. melanogaster genetic background, and double introgression of Nup96sim and Nup160sim did not reproductive generally lead to lethality when one was heterozygous and the other homozygous (hemizygous). It appears isolation that introgression of additional autosomal D. simulans genes is necessary to cause lethality and that the speciation effect of the introgression is dominant to D. melanogaster alleles. Interestingly, the genetic background affected dominance of Nup96sim, and double introgression carrying homozygous Nup96sim and hemizy- gous Nup160sim resulted in lethality. Thus, Nup96sim and Nup160sim seem to be two components of the same incompatibility. A handful of hybrid incompatibility genes that are responsible for mutation of D. simulans. D. melanogaster/D. simulans hybrids carry- reproductive isolation between species have been identified (Johnson ing the D. -

The Genetic Program of Pancreatic Beta-Cell Replication in Vivo
Page 1 of 65 Diabetes The genetic program of pancreatic beta-cell replication in vivo Agnes Klochendler1, Inbal Caspi2, Noa Corem1, Maya Moran3, Oriel Friedlich1, Sharona Elgavish4, Yuval Nevo4, Aharon Helman1, Benjamin Glaser5, Amir Eden3, Shalev Itzkovitz2, Yuval Dor1,* 1Department of Developmental Biology and Cancer Research, The Institute for Medical Research Israel-Canada, The Hebrew University-Hadassah Medical School, Jerusalem 91120, Israel 2Department of Molecular Cell Biology, Weizmann Institute of Science, Rehovot, Israel. 3Department of Cell and Developmental Biology, The Silberman Institute of Life Sciences, The Hebrew University of Jerusalem, Jerusalem 91904, Israel 4Info-CORE, Bioinformatics Unit of the I-CORE Computation Center, The Hebrew University and Hadassah, The Institute for Medical Research Israel- Canada, The Hebrew University-Hadassah Medical School, Jerusalem 91120, Israel 5Endocrinology and Metabolism Service, Department of Internal Medicine, Hadassah-Hebrew University Medical Center, Jerusalem 91120, Israel *Correspondence: [email protected] Running title: The genetic program of pancreatic β-cell replication 1 Diabetes Publish Ahead of Print, published online March 18, 2016 Diabetes Page 2 of 65 Abstract The molecular program underlying infrequent replication of pancreatic beta- cells remains largely inaccessible. Using transgenic mice expressing GFP in cycling cells we sorted live, replicating beta-cells and determined their transcriptome. Replicating beta-cells upregulate hundreds of proliferation- related genes, along with many novel putative cell cycle components. Strikingly, genes involved in beta-cell functions, namely glucose sensing and insulin secretion were repressed. Further studies using single molecule RNA in situ hybridization revealed that in fact, replicating beta-cells double the amount of RNA for most genes, but this upregulation excludes genes involved in beta-cell function. -

Supporting Information for Proteomics DOI 10.1002/Pmic.200500652
Supporting Information for Proteomics DOI 10.1002/pmic.200500652 Jeffery Forbus, Heidi Spratt, John Wiktorowicz, Zheng Wu, Istvan Boldogh, Larry Denner, Alexander Kurosky, Robert C. Brasier, Bruce Luxon and Allan R. Brasier Functional analysis of the nuclear proteome of human A549 alveolar epithelial cells by HPLC-high resolution 2-D gel electrophoresis ª 2006 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim www.proteomics-journal.com Supplementary Material Forbus et al., Functional Analysis Of The Nuclear Proteome Of Human A549 Alveolar Epithelial Cells By HPLC- High Resolution 2D Gel Electrophoresis Figure 1. Nuclear Preparation. (A) Microscopic analysis of sucrose step gradient purified nuclei. Purified nuclei were diluted in PBS, plated on a microscope cover slip, and stained with DAPI (Methods). Top panel, high resolution phase contrast microscopy; bottom panel, DAPI staining. Arrows indicate intact nucleoli. (B) Western immunoblot analysis of cytoplasmic and sucrose cushion purified nuclei. Equivalent cell amounts were loaded corresponding to 1 X 106 cells and probed with the indicated antibody, shown at left. Figure 2. Annotated 2-DE template of A549 nuclear proteins. Shown is a master gel representation of the nuclear proteins subjected to HPLC prefractionation and identification by peptide mass fingerprinting. Horizontal dimension, IEF was conducted over a pH range from 3-10. Vertical dimension, fractionation by SDS-PAGE. Migration of molecular weight standards (in kDa) are shown at left. Identity of proteins is shown in Table I. Table I. Annotated proteins in 2-DE template. Shown are high probability identifications from peptide mass fingerprinting using masses measured by MALDI-TOF in a Bayesian algorithm (ProFound). -

Sui Et Al Supplementary Figures
Supplementary Figures Figure S1. Western blot of WI38 fibroblasts treated with fractions of PC3M-LN4 conditioned media eluted from a heparin-sepharose+Cu2+ column with a linear gradient of NaCl plus 20mM imidazole. Supplemental Figure S2. List of proteins present in Tsp-1 repressing fractions (1.0 and 1.1M) and inactive adjacent fractions 0.9M NaCl Keratin 9 1.0M NaCl Keratin 1 Keratin, CK1 1.1M NaCl Keratin, CK2 Keratin 9 Keratin 9 1.2M NaCl Keratin, CK 10 Keratin, CK10 Keratin 1 Keratin 1 Keratin 10 Keratin 2a Keratin, CK10 Keratin 9 Keratin, CK6a Lactotransferrin Keratin, CK 2 Keratin, CK10 Keratin, CK14 precursor Keratin, CK16 57 kDa protein Keratin, CK6e Keratin 1B Keratin, CK6C Keratin 10 Keratin, CK5 Serotransferrin Keratin, CK14 Keratin, CK2 Keratin, CK16 precursor Keratin, CK5 Keratin 1B Cytokeratin type II ALB protein GAPDH Keratin 6L Hornerin similar to KIAA1501 Keratin, CK13 Keratin, type I GAPDH 24- Histone H2A.m cytoskeletal 14 Histone H2A.m dehydrocholesterol Histone H2B.q Keratin 5c Keratin, CK15 reductase precursor Lactotransferrin Keratin, CK3 49 kDa protein Tropomodulin 1 precursor GAPDH Histone H2B.q Protease serine 2 Keratin, Hb4 ALB protein ALB protein isoform B Serotransferrin Hypothetical protein Keratin K6irs Hypothetical protein precursor FLJ20261 Lactotransferrin FLJ90556 Histone 1, H2aa similar to KIAA1501 precursor Splice Isoform 2 of Hypothetical protein similar to KRT8 Histone H4 WD-repeat protein LOC65250 keratin 25 irs1 Serotransferrin 22 DKFZp686J1375 ROK1 precursor Ciliary rootlet Desmoglein-1 Cadherin -

Mass Spectrometric Identification of Ancient Proteins As Potential Molecular Biomarkers for a 2000-Year-Old Osteogenic Sarcoma
Mass Spectrometric Identification of Ancient Proteins as Potential Molecular Biomarkers for a 2000-Year-Old Osteogenic Sarcoma Agnes Bona1, Zoltan Papai1, Gabor Maasz1,2,3,4, Gabor A. Toth5, Eva Jambor1,3, Janos Schmidt1,2,3, Csaba Toth6, Csilla Farkas7, Laszlo Mark1,2,3,4* 1 Department of Analytical Biochemistry, Institute of Biochemistry and Medical Chemistry, University of Pecs, Pecs, Hungary, 2 Janos Szentagothai Research Center, University of Pecs, Pecs, Hungary, 3 Imaging Center for Life and Material Sciences, University of Pecs, Pecs, Hungary, 4 PTE-MTA Human Reproduction Research Group, Pecs, Hungary, 5 Institute of Biology, University of West Hungary, Szombathely, Hungary, 6 Department of Pathology, University Teaching Hospital Markusovszky, Szombathely, Hungary, 7 Department of Archaeology, Vas County Museums’ Directorate, Szombathely, Hungary Abstract Osteosarcoma is the most common primary malignant tumor of bone usually occurring in young adolescent and children. This disease has a poor prognosis, because of the metastases in the period of tumor progression, which are usually developed previous to the clinical diagnosis. In this paper, a 2000-year-old ancient bone remain with osteogenic sarcoma was analyzed searching for tumor biomarkers which are closely related to this disease. After a specific extraction SDS-PAGE gel electrophoresis followed by tryptic digestion was performed. After the digestion the samples were measured using MALDI TOF/TOF MS. Healthy bone samples from same archaeological site were used as control samples. Our results show that in the pathological skeletal remain several well known tumor biomarkers are detected such as annexin A10, BCL-2-like protein, calgizzarin, rho GTPase-activating protein 7, HSP beta-6 protein, transferrin and vimentin compared to the control samples.